Circadian clocks are operative in virtually all light-sensitive organisms, including cyanobacteria, fungi, plants, protozoans and metazoans. These timing devices allow their possessors to adapt their physiological needs to the time of day in an anticipatory way. In mammals, circadian pacemakers regulate many systemic processes, such as sleep-wake cycles, body temperature, heartbeat, and many physiological outputs conducted by peripheral organs, such as liver, kidney and the digestive tract [1]. On the basis of surgical ablation and transplantation experiments, it was established that the suprachiasmatic nucleus (SCN) in the hypothalamus coordinates most of these daily rhythms [2], probably through both synaptic connections and humoral signals [3]. Interestingly, self-sustained and cell-autonomous molecular oscillators do not only exist in pacemaker cells such as SCN neurons, but are also operative in most peripheral, non-neuronal cell types [4]. These peripheral oscillators participate in the circadian control
Research Perspective
of animal physiology. During the past few years, analysis of animal transcriptomes with the DNA microarray technology showed that many aspects of physiology are directly controlled by the circadian clock through control of the expression of enzymes and regulators involved in these physiological processes [5,6]. Although the mechanisms involved in these regulations are not yet understood in detail, it is likely that transcription factors whose expression is controlled by the circadian clock are involved [7]. Based on these circadian transcriptome profiling studies it is commonly thought that circadian metabolism is mainly the consequence of circadian transcription and possible effects of circadian clock-controlled post-transcriptional regulatory mechanisms have been largely neglected.
Interestingly, most of the enzymes involved in liver metabolism are localized in the membrane of the endoplasmic reticulum (ER) of hepatocytes. The ER is a complex luminal network in which protein synthesis, maturation, folding, and transport take place. It has been previously shown that the ER of hepatocytes exhibits a circadian dilatation which is a sign of ER stress [8]. This ER stress triggers the unfolded protein response (UPR) which is a conserved adaptative response to cope with the accumulation of unfolded proteins in this organelle. When unfolded proteins accumulate in ER, three pathways are activated, IRE1α, PERK and ATF6, which lead to the nuclear translocation of the transcription factors XBP1, ATF4 and ATF6, respectively. These transcription factors activate in turn the expression of genes coding for proteins involved in peptide folding and degradation to limit the accumulation of unfolded proteins [9]. In this context, we have recently described the posttranslational regulation of liver enzymes through a circadian clock-coordinated 12-hours period rhythmic activation of the IRE1α pathway [10]. The observed rhythmic activation of the IRE1α pathway leads to the expression with a 12-hours period of the XBP1-regulated genes that are included in the 12-hours period genes described recently in mouse liver [11]. Persistent activation of the IRE1α pathway in circadian clock deficient Cry1/Cry2 ko mice induced the downregulation of ER membrane localized enzymes, including HMGCR and SCD1, leading to a perturbed lipid metabolism in the liver of this mice. The decreased expression of these enzymes could be caused by activation of the ER Associated Degradation (ERAD), a process involved in the elimination of unfolded proteins inside the ER[12] regulated by the IRE1α-XBP1 pathway [13], which has been shown to induce the degradation of HMGCR and SCD1. In addition, IRE1α is a ribonuclease that can also induce endonucleolytic decay of many ER-localized mRNA including Hmgcr mRNA [14,15]. These two functions could contribute in parallel to the regulation of lipid metabolism by ER stress. Elsewhere, the IRE1α-XBP1 pathway controls also lipid metabo-lism through direct transcriptional regulation of the genes Scd1, Dgat2 and Acc2 involved in lipogenesis. As a consequence, liver-specific deletion of the Xpb1 gene resulted in a dramatic reduction of plasma lipids [16]. Finally, it has been shown that ER stress induces the degradation of the apolipoprotein ApoB100 [17,18] and then blocks VLDL secretion [19], which might be responsible for the fat accumulation in the liver in tunicamycin-injected mice [20].
Interestingly, IRE1α activation has been recently linked to induction of autophagy through activation of the Jun-Kinase pathway [21]. In addition, a genomic screen in fly cells demonstrated that knocking down genes involved in protein folding inside the ER or in the UPR, including Xbp1, increases basal autophagy levels [22]. Autophagy is a survival pathway classically associated with adaptation to nutrient starvation [23] and, as UPR, autophagy presented a diurnal rhythm of activation in rodent liver [24,25]. This is of particular interest if we consider the fact that autophagy is linked to lipid metabolism through regulation of intracellular lipid stores [26]. As a consequence, mice with an adipose tissue-specific deletion of the Atg7 gene, an important regulator of autophagy, present an important defect in lipid storage [27,28]. IRE1α-dependent rhythmic regulation of autophagy could then participates to the circadian clock-coordinated lipid metabolism in mammals.
The disturbed metabolism observed in Cry1/Cry2 ko mice is probably responsible of the aberrant activation of the Sterol Responsive Element Binding Protein (SREBP) transcription factor, an ER membrane bond protein that, in low sterol conditions, translocates to the Golgi to be cleaved and released in order to migrate in the nucleus where it activates genes coding for enzymes involved in cholesterol and fatty acid metabolism [29]. It has been shown that the ER stress induced activation of SREBP1 and SREBP2 [30,31] correlates with the depletion of INSIG regulatory proteins probably through a decreased synthesis of the protein [32,33]. Interestingly, the circadian clock influences also the activation of the SREBP pathway through the control of Insig2 mRNA expression [34]. Both transcriptional and post-transcriptional circadian clock-coordinated events seem to be involved in the rhythmic activation of the SREBP pathway.
As summarised in Figure 1, in addition to their rhythmic activation, all these pathways have in common the fact that they are regulated by feeding-fasting events. However, this feeding rhythm, like most behaviour, is also controlled by the circadian clock. To discriminate the genes dependant or not on a functional local circadian oscillator, this local clock has been inactivated in mouse liver. This strategy reveals that the expression of approximately 90 % of the rhythmic genes is dependent on a functional circadian clock and only 10 % is dependent on systemic cues [35]. However, the influence of feeding on rhythmic gene expression has been evaluated by a recent study which discriminates between gene induced by feeding and fasting. As expected, food-induced and food-repressed genes present a rhythmic expression which is shifted in response to a change in the feeding schedule [36]. More interestingly, this shift in the feeding schedule is able to induce rhythmic expression of food-regulated genes in the liver of Cry1/Cry2 ko mice. These two studies raise the question of the differential influence of the molecular circadian oscillator and systemic cues on rhythmic gene expression: if these two signals can independently drive rhythmic gene expression, the circadian clock is able to fine-tune and modify feeding cues [34,36], whereas feeding cues can synchronize the molecular oscillator in peripheral organs [37].
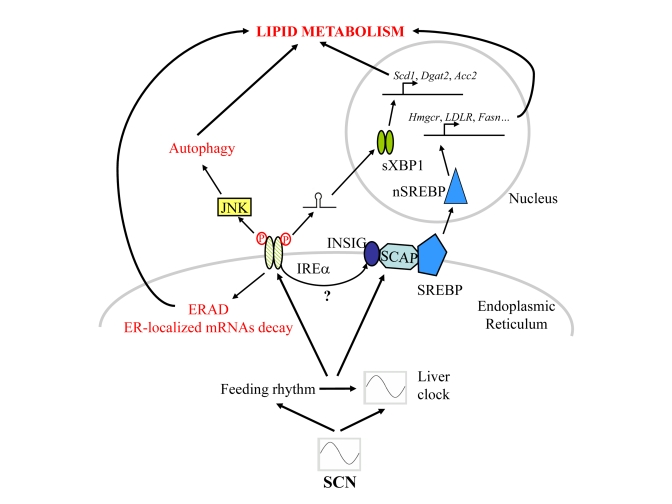
Figure 1. Schematic representation of the signalling pathways post-transcriptionally regulated by the circadian clock and/or rhythmic feeding cues in mouse liver.
However, feeding and food-regulated signals, as for example food regulated hormones like insulin, glucagon or leptin, did not represent the only circadian clock-regulated cues that can influence lipid metabolism. For example, the pituitary-secreted growth hormone (GH) has been shown to influence lipid metabolism in mouse liver. Long term excess GH secretion produces high serum triglyceride levels through stimulated lipolysis [38], whereas inhibition of GH signaling induced perturbed lipid metabolism resulting in liver steatosis [39], probably caused by reduced activation of HNF3β [40]. Moreover, the ultradian secretion patterns of GH are directly responsible for the sexually dimorphic expres-sion of several hepatic enzymes involved in steroids and fatty acids metabolism [41]. Interestingly, this dimorphism is impaired in Cry1/Cry2 ko mice, with males exhibiting a feminized liver likely because of altered ultradian GH secretion in absence of a functional circadian clock [42].
During aging, the circadian system becomes much less responsive to entrainment by light [43,44], and displays loss of temporal precision and robustness [45-47]. Such alterations of the circadian clock likely drive attenuation of the diurnal rhythm in circulating leptin [48]. Pulsatile GH secretion is also dramatically impaired in elderly subjects [49-51], leading to modifications in GH-dependent liver metabolism that resemble those observed in clock-deficient animals [42,52]. Interesting-ly, the various UPR pathways also decline in the liver during aging [53], as well as autophagy [54]. In summary, many aspects of lipid metabolism that are regulated by the circadian clock exhibit profound changes when age increases, although the liver circadian oscillator appears preserved in aged rats [55]. These changes could thus at least partly originate from alterations of the network constituted of the central clock and other peripheral oscillators. In this respect, it is worth noting that mice bearing mutated alleles of the circadian genes Clock and Bmal1 display signs of premature aging [56,57]. The complexity of systemic cues influencing rhythmic gene expression has thus been rising during the last decade and defining the influence of these different signals on rhythmic gene expression will be thus an exciting challenge for the following years.
Conflicts of Interest
The authors of this manuscript have no conflict of interest to declare.
References
- 1. Green CB , Takahashi JS and Bass J. The meter of metabolism. Cell. 2008; 134: 728 -742. [PubMed] .
- 2. Ralph MR , Foster RG , Davis FC and Menaker M. Transplanted suprachiasmatic nucleus determines circadian period. Science. 1990; 247: 975 -978. [PubMed] .
- 3. Silver R , LeSauter J , Tresco PA and Lehman MN. A diffusible coupling signal from the transplanted suprachiasmatic nucleus controlling circadian locomotor rhythms. Nature. 1996; 382: 810 -813. [PubMed] .
- 4. Yamazaki S , Numano R , Abe M , Hida A , Takahashi R , Ueda M , Block GD , Sakaki Y , Menaker M and Tei H. Resetting central and peripheral circadian oscillators in transgenic rats. Science. 2000; 288: 682 -685. [PubMed] .
- 5. Akhtar RA , Reddy AB , Maywood ES , Clayton JD , King VM , Smith AG , Gant TW , Hastings MH and Kyriacou CP. Circadian cycling of the mouse liver transcriptome, as revealed by cdna microarray, is driven by the suprachiasmatic nucleus. Curr Biol. 2002; 12: 540 -550. [PubMed] .
- 6. Panda S , Antoch MP , Miller BH , Su AI , Schook AB , Straume M , Schultz PG , Kay SA , Takahashi JS and Hogenesch JB. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell. 2002; 109: 307 -320. [PubMed] .
- 7. Ueda HR , Chen W , Adachi A , Wakamatsu H , Hayashi S , Takasugi T , Nagano M , Nakahama K-I , Suzuki Y , Sugano S , Iino M , Shigeyoshi Y and Hashimoto S. A transcription factor response element for gene expression during circadian night. Nature. 2002; 418: 534 -539. [PubMed] .
- 8. Chedid A and Nair V: Diurnal rhythm in endoplasmic reticulum of rat liver. Electron microscopic study. Science. 1972; 175: 176 -179. [PubMed] .
- 9. Ron D and Walter P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Biol. 2007; 8: 519 -529. [PubMed] .
- 10. Cretenet G , Le Clech M and Gachon F. Circadian clock-coordinated 12 hr period rhythmic activation of the ire1α pathway controls lipid metabolism in mouse liver. Cell Metab. 2010; 11: 47 -57. [PubMed] .
- 11. Hughes ME , DiTacchio L , Hayes KR , Vollmers C , Pulivarthy S , Baggs JE , Panda S and Hogenesch JB. Harmonics of circadian gene transcription in mammals. PLoS Genet. 2009; 5: e1000442 [PubMed] .
- 12. Meusser B , Hirsch C , Jarosch E and Sommer T. Erad: The long road to destruction. Nat Cell Biol. 2005; 7: 766 -772. [PubMed] .
- 13. Yoshida H , Matsui T , Hosokawa N , Kaufman RJ , Nagata K and Mori K. A time-dependent phase shift in the mammalian unfolded protein response. Dev Cell. 2003; 4: 265 -271. [PubMed] .
- 14. Han D , Lerner AG , Vande Walle L , Upton J-P , Xu W , Hagen A , Backes BJ , Oakes SA and Papa FR. Ire1α kinase activation modes control alternate endoribonuclease outputs to determine divergent cell fates. Cell. 2009; 138: 562 -575. [PubMed] .
- 15. Hollien J , Lin JH , Li H , Stevens N , Walter P and Weissman JS. Regulated ire1-dependent decay of messenger rnas in mammalian cells. J Cell Biol. 2009; 186: 323 -331. [PubMed] .
- 16. Lee A-H , Scapa EF , Cohen DE and Glimcher LH. Regulation of hepatic lipogenesis by the transcription factor xbp1. Science. 2008; 320: 1492 -1496. [PubMed] .
- 17. Liao W and Chan L. Tunicamycin induces ubiquitination and degradation of apolipoprotein b in hepg2 cells. Biochem J. 2001; 353: 493 -501. [PubMed] .
- 18. Qiu W , Kohen-Avramoglu R , Mhapsekar S , Tsai J , Austin RC and Adeli K. Glucosamine-induced endoplasmic reticulum stress promotes apob100 degradation: Evidence for grp78-mediated targeting to proteasomal degradation. Arterioscler Thromb Vasc Biol. 2005; 25: 571 -577. [PubMed] .
- 19. Ota T , Gayet C and Ginsberg HN. Inhibition of apolipoprotein b100 secretion by lipid-induced hepatic endoplasmic reticulum stress in rodents. J Clin Invest. 2008; 118: 316 -332. [PubMed] .
- 20. Rutkowski DT , Wu J , Back S-H , Callaghan MU , Ferris SP , Iqbal J , Clark R , Miao H , Hassler JR , Fornek J , Katze MG , Hussain MM , Song B , Swathirajan J , Wang J , Yau GDY and Kaufman RJ. Upr pathways combine to prevent hepatic steatosis caused by er stress-mediated suppression of transcriptional master regulators. Dev Cell. 2008; 15: 829 -840. [PubMed] .
- 21. Ogata M , Hino S-i , Saito A , Morikawa K , Kondo S , Kanemoto S , Murakami T , Taniguchi M , Tanii I , Yoshinaga K , Shiosaka S , Hammarback JA , Urano F and Imaizumi K. Autophagy is activated for cell survival after endoplasmic reticulum stress. Mol Cell Biol. 2006; 26: 9220 -9231. [PubMed] .
- 22. Arsham AM and Neufeld TP. A genetic screen in drosophila reveals novel cytoprotective functions of the autophagy-lysosome pathway. PLoS ONE. 2009; 4: e6068 [PubMed] .
- 23. Levine B and Kroemer G. Autophagy in the pathogenesis of disease. Cell. 2008; 132: 27 -42. [PubMed] .
- 24. Pfeifer U and Scheller H. A morphometric study of cellular autophagy including diurnal variations in kidney tubules of normal rats. J Cell Biol. 1975; 64: 608 -621. [PubMed] .
- 25. Pfeifer U and Strauss P. Autophagic vacuoles in heart muscle and liver. A comparative morphometric study including circadian variations in meal-fed rats. J Mol Cell Cardiol. 1981; 13: 37 -49. [PubMed] .
- 26. Singh R , Kaushik S , Wang Y , Xiang Y , Novak I , Komatsu M , Tanaka K , Cuervo AM and Czaja MJ. Autophagy regulates lipid metabolism. Nature. 2009; 458: 1131 -1135. [PubMed] .
- 27. Singh R , Xiang Y , Wang Y , Baikati K , Cuervo AM , Luu YK , Tang Y , Pessin JE , Schwartz GJ and Czaja MJ. Autophagy regulates adipose mass and differentiation in mice. J Clin Invest. 2009; 119: 3329 -3339. [PubMed] .
- 28. Zhang Y , Goldman S , Baerga R , Zhao Y , Komatsu M and Jin S. Adipose-specific deletion of autophagy-related gene 7 (atg7) in mice reveals a role in adipogenesis. Proc Natl Acad Sci U S A. 2009; 106: 19860 -19865. [PubMed] .
- 29. Espenshade PJ and Hughes AL. Regulation of sterol synthesis in eukaryotes. Annu Rev Genet. 2007; 41: 401 -427. [PubMed] .
- 30. Colgan SM , Tang D , Werstuck GH and Austin RC. Endoplasmic reticulum stress causes the activation of sterol regulatory element binding protein-2. Int J Biochem Cell Biol. 2007; 39: 1843 -1851. [PubMed] .
- 31. Wang H , Kouri G and Wollheim CB. Er stress and srebp-1 activation are implicated in β-cell glucolipotoxicity. J Cell Sci. 2005; 118: 3905 -3915. [PubMed] .
- 32. Bobrovnikova-Marjon E , Hatzivassiliou G , Grigoriadou C , Romero M , Cavener DR , Thompson CB and Diehl JA. Perk-dependent regulation of lipogenesis during mouse mammary gland development and adipocyte differentiation. Proc Natl Acad Sci U S A. 2008; 105: 16314 -16319. [PubMed] .
- 33. Lee JN and Ye J. Proteolytic activation of sterol regulatory element-binding protein induced by cellular stress through depletion of insig-1. J Biol Chem. 2004; 279: 45257 -45265. [PubMed] .
- 34. Le Martelot G , Claudel T , Gatfield D , Schaad O , Kornmann B , Sasso GL , Moschetta A and Schibler U. Rev-erbα participates in circadian srebp signaling and bile acid homeostasis. PLoS Biol. 2009; 7: e1000181 [PubMed] .
- 35. Kornmann B , Schaad O , Bujard H , Takahashi JS and Schibler U. System-driven and oscillator-dependent circadian transcription in mice with a conditionally active liver clock. PLoS Biol. 2007; 5: 179 -189. .
- 36. Vollmers C , Gill S , DiTacchio L , Pulivarthy SR , Le HD and Panda S. Time of feeding and the intrinsic circadian clock drive rhythms in hepatic gene expression. Proc Natl Acad Sci U S A. 2009; 106: 21453 -21458. [PubMed] .
- 37. Damiola F , Le Minh N , Preitner N , Kornmann B , Fleury-Olela F and Schibler U. Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev. 2000; 14: 2950 -2961. [PubMed] .
- 38. Frick F , Bohlooly-Y M , Linden D , Olsson B , Tornell J , Eden S and Oscarsson J. Long-term growth hormone excess induces marked alterations in lipoprotein metabolism in mice. Am J Physiol Endocrinol Metab. 2001; 281: E1230 -1239. [PubMed] .
- 39. Fan Y , Menon RK , Cohen P , Hwang D , Clemens T , DiGirolamo DJ , Kopchick JJ , Le Roith D , Trucco M and Sperling MA. Liver-specific deletion of the growth hormone receptor reveals essential role of growth hormone signaling in hepatic lipid metabolism. J Biol Chem. 2009; 284: 19937 -19944. [PubMed] .
- 40. Rowland JE , Lichanska AM , Kerr LM , White M , d'Aniello EM , Maher SL , Brown R , Teasdale RD , Noakes PG and Waters MJ. In vivo analysis of growth hormone receptor signaling domains and their associated transcripts. Mol Cell Biol. 2005; 25: 66 -77. [PubMed] .
- 41. Waxman DJ and Holloway MG. Sex differences in the expression of hepatic drug metabolizing enzymes. Mol Pharmacol. 2009; 76: 215 -228. [PubMed] .
- 42. Bur IM , Cohen-Solal AM , Carmignac D , Abecassis P-Y , Chauvet N , Martin AO , van der Horst GTJ , Robinson ICAF , Maurel P , Mollard P and Bonnefont X. The circadian clock components cry1 and cry2 are necessary to sustain sex dimorphism in mouse liver metabolism. J Biol Chem. 2009; 284: 9066 -9073. [PubMed] .
- 43. Sutin EL , Dement WC , Heller HC and Kilduff TS. Light-induced gene expression in the suprachiasmatic nucleus of young and aging rats. Neurobiol Aging. 1993; 14: 441 -446. [PubMed] .
- 44. Zhang Y , Kornhauser JM , Zee PC , Mayo KE , Takahashi JS and Turek FW. Effects of aging on light-induced phase-shifting of circadian behavioral rhythms, fos expression and creb phosphorylation in the hamster suprachiasmatic nucleus. Neuroscience. 1996; 70: 951 -961. [PubMed] .
- 45. Peng M-t , Jiang M-J and Hsu H-K. Changes in running-wheel activity, eating and drinking and their day/night distributions throughout the life span of the rat. J Gerontol. 1980; 35: 339 -347. [PubMed] .
- 46. Scarbrough K , Losee-Olson S , Wallen EP and Turek FW. Aging and photoperiod affect entrainment and quantitative aspects of locomotor behavior in syrian hamsters. Am J Physiol Regul Integr Comp Physiol. 1997; 272: R1219 -1225. .
- 47. Valentinuzzi VS , Scarbrough K , Takahashi JS and Turek FW. Effects of aging on the circadian rhythm of wheel-running activity in c57bl/6 mice. Am J Physiol Regul Integr Comp Physiol. 1997; 273: R1957 -1964. .
- 48. Downs JL and Urbanski HF. Aging-related sex-dependent loss of the circulating leptin 24-h rhythm in the rhesus monkey. J Endocrinol. 2006; 190: 117 -127. [PubMed] .
- 49. Giustina A and Veldhuis JD. Pathophysiology of the neuroregulation of growth hormone secretion in experimental animals and the human. Endocr Rev. 1998; 19: 717 -797. [PubMed] .
- 50. Kappeler L , Zizzari P , Alliot J , Epelbaum J and Bluet-Pajot M-T. Delayed age-associated decrease in growth hormone pulsatile secretion and increased orexigenic peptide expression in the lou c/jall rat. Neuroendocrinology. 2004; 80: 273 -283. [PubMed] .
- 51. Woller MJ , Everson-Binotto G , Nichols E , Acheson A , Keen KL , Bowers CY and Terasawa E. Aging-related changes in release of growth hormone and luteinizing hormone in female rhesus monkeys. J Clin Endocrinol Metab. 2002; 87: 5160 -5167. [PubMed] .
- 52. Dhir RN and Shapiro BH. Interpulse growth hormone secretion in the episodic plasma profile causes the sex reversal of cytochrome p450s in senescent male rats. Proc Natl Acad Sci U S A. 2003; 100: 15224 -15228. [PubMed] .
- 53. Naidoo N Er and aging-protein folding and the er stress response. Ageing Res Rev. 2009; 8: 150 -159. [PubMed] .
- 54. Cuervo AM , Bergamini E , Brunk UT , Dröge W , Ffrench M and Terman A. Autophagy and aging: The importance of maintaining "Clean" Cells. Autophagy. 2005; 1: 131 -140. [PubMed] .
- 55. Yamazaki S , Straume M , Tei H , Sakaki Y , Menaker M and Block GD. Effects of aging on central and peripheral mammalian clocks. Proc Natl Acad Sci U S A. 2002; 99: 10801 -10806. [PubMed] .
- 56. Antoch MP , Gorbacheva VY , Vykhovanets O , Toshkov IA , Kondratov RV , Kondratova AA , Lee C and Nikitin AY. Disruption of the circadian clock due to the clock mutation has discrete effects on aging and carcinogenesis. Cell Cycle. 2008; 7: 1197 -1204. [PubMed] .
- 57. Kondratov RV , Kondratova AA , Gorbacheva VY , Vykhovanets OV and Antoch MP. Early aging and age-related pathologies in mice deficient in bmal1, the core componentof the circadian clock. Genes Dev. 2006; 20: 1868 -1873. [PubMed] .


