Introduction
Disruption in metabolic homeostasis and over accumulation of metabolites, cholesterol, bile acids, triglycerides (fat), or glucose, play causative roles in the development of metabolic disorders, such as, atherosclerosis and related heart disease, fatty liver, obesity, and diabetes. The NAD+-dependent SIRT1 deacetylase plays a critical role in maintaining metabolic homeostasis which affects aging so that SIRT1 increases life spans in most organisms, including mammals [1-3]. Despite extensive studies on SIRT1 function and its beneficial metabolic effects, how the expression of SIRT1 is regulated under normal conditions and how SIRT1 levels are decreased in metabolic disease states remain unclear. In this review, we survey recent studies showing how SIRT1 expression is regulated at the post-transcriptional level, focusing on microRNAs (miRs) which have recently emerged as important cellular regulators [4-6]. We also review recent studies showing that the nuclear receptor FXR/SHP cascade pathway which controls expression of miR-34a and its target SIRT1 in normal conditions and is dysregulated in metabolic disease states.
SIRT1: a key regulator in cellular metabolism
Caloric restriction (CR) was shown to increase life span and promote survival in yeast, worms, flies, rodents and perhaps primates [1,2]. SIRT1 mediates the beneficial metabolic effects of CR in an NAD+-dependent manner by deacetylating and altering the activities of transcriptional factors which regulate metabolic genes [1,2,7]. SIRT1 deacetylates and activates transcript-tional ability of metabolic regulators, such as PGC-1α, p53, Foxo 1, NF-κB, LXR, and FXR that are involved in lipid and glucose metabolism, inflammation, mitochondrial biogenesis, and energy balance [1,2,8-12]. In addition, SIRT1 was shown to be recruited to the promoter of metabolic target genes and suppress their transcription [13,14]. It was reported that SIRT1 is associated with the promoter of PPARγ, a key adipogenic factor, and suppresses PPARγ transcription by recruiting the corepressors, NcoR1 and SMRT [14]. SIRT1 was reported to bind to the UCP 2 gene promoter and inhibit its transcription in pancreatic β-cells, resulting in increased ATP production and insulin secretion [13]. SIRT1 was also shown to improve insulin sensitivity by repressing transcription of protein tyrosine phosphatase 1B, a major negative regulator of insulin action, via histone deacetylation [15]. Beneficial metabolic functions of SIRT1 have been demonstrated in studies using small molecule activators and transgenic mice that are null for SIRT1 or overexpress SIRT1 [16-20]. The natural compound resveratrol and the synthetic compound SRT1720 are activators of SIRT1 and have been shown to ameliorate insulin resistance, increase mitochondrial content, improve metabolic profiles, and increase survival in mice fed a high-fat diet [16-18]. Transgenic mice expressing SIRT1 were shown to be resistant to body weight gain and ameliorated insulin resistance and glucose intolerance in these mice compared to wild-type control mice [20]. Further, transgenic mice expressing moderate amounts of SIRT1 were also shown to protect livers from diet-induced metabolic damage [12,21]. Consistent with these reports, in liver-specific SIRT1 null mice challenged with a high fat diet, fatty acid metabolism was altered and the development of fatty livers and inflammatory responses were promoted [19,22]. Loss of function studies also showed that SIRT1 decreases endothelial activation in hypercholesterolemic ApoE-/- mice without affecting endothelium-dependent vasodilatation [23]. All these recent studies demonstrate that SIRT1 is a key regulator of cellular metabolism and mediates beneficial metabolic effects.
MicroRNAs: emerging metabolic regulators
MicroRNAs (miRNAs) are small (approximately 22 nt) non-coding RNAs that control gene expression [4-6]. MiRs are transcribed from DNA by RNA polymerase II as hairpin precursors which are further processed to mature forms [4-6]. MiRs bind to the 3'-untranslated region (UTR) of target mRNAs and inhibit their expression by causing mRNA cleavage or inhibition of translation. Approximately 30% of all human genes are thought to be regulated by miRs [5,6] and indeed, miRs control gene expression in diverse biological processes including development, differentiation, cell prolifera-tion, and apoptosis. Recent studies have demonstrated crucial roles of miRNAs in the regulation of cellular metabolism [24-32]. MiRs are involved in lipid and glucose metabolism in major metabolic tissues, such as, liver, pancreas, adipose, and muscle as summarized in Table 1. Mir-122 is the most abundant miR in the liver and plays important roles in a wide variety of liver functions ranging from cholesterol metabolism, liver cancer, stress responses, viral infection, to circadian regulation of hepatic genes [24,28,29]. MiR-33 has been shown to contribute to the regulation of cholesterol homeostasis by targeting the cholesterol transporter genes, ABCA1 and ABCG1 [25,26]. Our group recently reported that miR-34a targets hepatic SIRT1 and, interestingly, expression of miR-34a was highly elevated and SIRT1 levels were decreased in fatty livers of diet-induced obese mice [30]. MiR-34a was also shown to suppress insulin secretion in pancreatic β-cells [33]. The roles of miR-375 in pancreatic islet functions, especially in insulin gene transcription, insulin secretion, and islet cell growth, are also well established [31,32]. Mir-27 and miR-378 were reported to control adipocyte differentiation and lipid synthesis, respectively [34,35]. MiR-223 was shown to regulate glucose uptake in cardiomyocytes and miR-696 to regulate mitochondria biogenesis and fatty acid oxidation in gastrocnemius muscle [36,37]. In line with their critical functions, miRs are often underexpressed or overexpressed in disease states [4,6,24,28,30,38-40]. Recent studies have shown that restoring miRs or downregulating miRs using antisense miR inhibitors, called antagomirs, has improved transcriptional and biological outcomes, demonstrating that miRs are promising therapeutic targets [4,24,38].
Down-regulation of SIRT1 by microRNAs
Consistent with its critical roles in diverse biological processes, the regulation of SIRT1 expression is fine tuned at multiple levels, including transcriptional, post-transcriptional, and post-translational levels. The general regulation of SIRT1 activity and expression has been thoroughly reviewed in excellent articles [1-3,41] and, therefore, this review focuses on the regulation of SIRT1 expression by miRs (Table 2). MiR-34a was first identified as a posttranscriptional regulator of SIRT1 in the regulation of apoptosis under cellular genotoxic stress in human colon cancer HCT116 cells [42]. MiR-34a binds to the 3' UTR of SIRT1 mRNA in a partial complementary manner and represses its translation but does not affect mRNA degradation [30,42]. Our group further reported that miR-34a targets hepatic SIRT1 in the regulation of cellular metabolism in human hepatoma HpeG2 cells and in mouse liver in vivo using adenoviral-mediated overexpression of miR-34a [30]. Remarkably, we observed that miR-34a levels are highly elevated and SIRT1 protein levels are substantially decreased in the fatty livers of both diet-induced obese mice and the leptin-deficient ob/ob mice [30]. These findings are in line with recent studies showing that miR-34a is the most elevated miR in livers exhibiting nonalcoholic steatohepatitis, a spectrum of nonalcoholic fatty liver diseases in humans [39]. Other miRs also target SIRT1. In response to nutritional availability, miR-132 was shown to downregulate SIRT1, resulting in activation of inflammatory pathways in adipose tissues [43]. MiR-199a was identified as a negative regulator of SIRT1 and HIF1a, a key mediator of hypoxia [44]. Low oxygen tension results in acute downregulation of miR-199a in cardiac myocytes and in porcine heart and this reduction is required for upregulation of its targets, HIF-1a and SIRT1 in response to decreased oxygen [44]. Interestingly, a recent study showed that SIRT1 protein levels are much higher in mouse embryonic stem cells (ESCs) than in differentiated tissues and that miRNAs, miR-181a and b, miR-9, miR-204, miR-199b, and miR-135, post-transcriptionally down-regulate SIRT1 during mouse ESC differentiation and maintain low levels of SIRT1 expression in differentiated tissues [45].
Table 1. MicroRNAs regulating cellular metabolism in major metabolic tissues.
| MicroRNA | Direct targets[putative] | Functions in Metabolism (references) | Tissues (cultured cells) |
| miR-33 | ABCA1, NPC1 | Cholesterol homeostasis (25, 26) | Liver (HepG2) |
| miR-34a | SIRT1 | lipid metabolism, promotes fatty liver (30) | |
| miR-370 | Cpt1a | Fatty acid and triglyceride biosynthesis (29) | |
| miR-122 | CAT-1 ADAM17 | Hepatic lipid metabolism (24, 29) Circadian gene expression (28) | |
| miR-34a | VAMP2 | B-cell exocytosis (33) | Pancreatic Islets (MIN6, INS-1) |
| miR-124a | Foxa2 | Intracellular signaling in pancreatic β-cell (27) | |
| miR-375 | MTPN | Regulates catecholamine release Inhibits insulin secretion (31, 32) | |
| miR-27a | [PPARγ, C/EBPα] | Inhibits adipocyte formation, Down-regulated during adipogenic differentiation (34) | (Adipocytes, 3T3-L1, ST2) |
| miR-378/378* | [Ribosomal proteins] | Upregulates adipocyte differentiation and lipid synthesis (35) | |
| miR-223 | Glut4 | Glucose uptake and insulin resistance (36) | Muscle Gastrocnemius (Cardiomyocyte, C2C12) |
| miR-696 | [PGC1α] | Muscle metabolism, mitochondria biogenesis and fatty acid oxidation (37) |
Table 2. MicroRNAs targeting SIRT12.
| MicroRNA | Sequences of microRNAs | Size (nt) | Biological functions (references) |
| miR-34a | 5'-uggcagugucuuagcugguugu-3' | 22 | Hepatic lipid metabolism (30) Islet β-cell exocytosis (33) Cell apoptosis (42) |
| miR-132 | 5'-uaacagucuacagccauggucg-3' | 22 | Stress-induced chemokine production (43) |
| miR-199a | 5'-cccaguguucagacuaccuguuc-3' | 25 | Hypoxia preconditioning (44) |
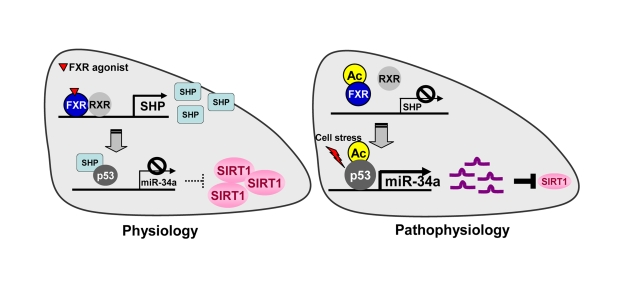
Figure 1. The FXR/SHP pathway controlling miR-34a and SIRT1 expression. Under normal
conditions, activation of FXR signaling induces the metabolic repressor SHP
in liver. SHP is then recruited to the miR-34a promoter and inhibits
binding of the key activator p53 to the DNA, resulting in decreased miR-34a
expression. Inhibition of miR-34a results in increased hepatic SIRT1 levels.
In contrast, under pathophysiological conditions such as fatty livers of
obese mice, the dysregulated FXR/SHP pathway due to highly elevated FXR
acetylation no longer inhibits transcription of miR-34a. The dysregulated
FXR/SHP pathway, along with acetylation of p53 due to cellular stress under
metabolic disease states, result in elevated miR-34a expression, which
contributes to decreased SIRT1 levels.
A novel FXR/SHP/miR-34a pathway controlling SIRT1 levels
The nuclear bile acid receptor, Farnesoid X Receptor (FXR), plays an important role in maintaining lipid and glucose levels by regulating expression of numerous metabolic genes mainly in the liver and intestine [46]. Consistent with its important metabolic functions, disruption of the FXR gene in transgenic mice was associated with metabolic diseases, including hypercholesterolemia, cholesterol gallstone disease, fatty liver, and type 2 diabetes [46-49]. Activation of FXR in diabetic obese mice improved metabolic outcomes by reducing serum glucose and lipid levels [50]. Although both FXR and SIRT1 have been shown to be critical for hepatic metabolism and activation of both proteins improves metabolic outcomes in diet-induced obese mice [17,18,46,47,50], it was unknown whether the expression and activity of these two proteins are coordinately regulated. In recent studies, we found that FXR positively regulates hepatic SIRT1 expression by inhibiting expression of miR-34a [30]. As shown in Figure 1, under normal conditions, miR-34a levels are down-regulated by a nuclear receptor cascade pathway involving FXR and orphan nuclear receptor and metabolic repressor, Small Heterodimer Partner (SHP) [51,52]. Upon induction by activated FXR, SHP is recruited to the miR-34a promo- ter and suppresses its transcription by inhibiting the promoter occupancy of p53, the key activator of the miR-34a gene [53]. Subsequently, inhibition of miR-34a contributes to increased expression of SIRT1. This FXR/SHP pathway was also shown to play a crucial role in the regulation of hepatic bile acid synthesis by inhibiting the rate-limiting bile acid synthetic enzyme CYP7A1 [51,52] and to suppress fatty liver formation by inhibiting the key lipogenic activator SREBP-1c [54]. Our group has identified molecular mechanisms by which SHP inhibits its target genes by coordinately recruiting chromatin modifying repressive cofactors, including HDACs, G9a metyltransferase, and Brm-containing Swi/Snf remodeling complex [55-57]. Consistent with these previous findings, we observed recruitment of HDACs to the miR-34a promoter in mouse liver after treatment with the synthetic FXR agonist, GW4064 (not shown). In contrast, in fatty livers of obese mice, the FXR/SHP pathway is dysregulated such that miR-34a levels are highly elevated, which contributes to reduced SIRT1 levels [30]. Interestingly, activation of FXR signaling in obese mice by daily treatment with GW4064 for 5 days or by hepatic expression of FXR using adenoviral delivery decreased miR-34a levels and restored SIRT1 levels [30]. Consistent with a critical role for FXR in positively controlling SIRT1 through the inhibition of miR-34a, miR-34a levels were indeed elevated and SIRT1 protein levels are substantially decreased in FXR null mice [30]. Our findings suggest an intriguing link among FXR activation, decreased miR-34a levels, increased SIRT1 levels, and beneficial metabolic outcomes.
A positively interacting FXR/SIRT1 regulatory loop
In the FXR/SHP/miR-34a pathway, FXR positively regulates hepatic SIRT1 levels by inhibiting transcription of the miR-34a gene. These findings, along with previous studies showing the p53/miR-34a/SIRT1 feedback loop [42,58], suggest intriguing regulatory loops controlling SIRT1 expression (Figure 2). In the short regulatory loop, SIRT1 positively auto-regulates its own expression by deacetylating p53 and histones at the miR-34a promoter, resulting in suppression of miR-34a [9,30,42,53,58]. In the long regulatory loop, SIRT1-mediated deacetylation of FXR increases FXR's transactivation ability by increasing binding of the FXR/RXR heterodimer to DNA resulting in induction of SHP and repression of miR-34a expression [11,30]. We observed that FXR acetylation is dynamically controlled by p300 acetylase and SIRT1 deacetylase under normal conditions, and remarkably, FXR acetylation levels are highly elevated in fatty livers of obese mice [11]. Interestingly, treatment daily with the SIRT1 activator resveratrol for 1 week or adenoviral-mediated hepatic expression of SIRT1 substantially reduced FXR acetylation with beneficial metabolic effects [11]. These results are consistent with the idea that the transactivation activity of FXR is low in obese mice due to highly elevated FXR acetylation, which contributes to increased expression of miR-34a. Subsequently, elevated miR- 34a suppresses expression of SIRT1, which then further decreases FXR activity, resulting in a vicious FXR/miR-34a/SIRT1 regulatory loop in metabolic disease states. In addition to deacetylation of FXR, SIRT1 has been implicated as a positive regulator of the expression and activity of FXR. During fasting, PGC-1α was shown to increase expression of the FXR gene and function as a coactivator of FXR [59]. Since SIRT1 deacetylates and increases PGC-1α activity [8], SIRT1 should increase FXR expression and activity by enhancing PGC-1α activity. All these recent studies strongly suggest that the expression and activity of these two proteins are mutually and coordinately regulated.
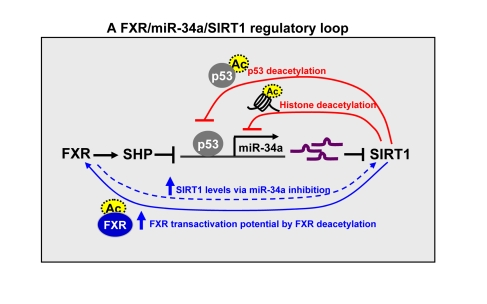
Figure 2. A FXR/SIRT1
positive-feedback regulatory loop. The expression and activity of FXR
and SIRT1 are mutually and coordinately regulated. SIRT1 positively
auto-regulates its own expression by inhibiting miR-34a via deacetylation
(as indicated by dotted circles) of p53 and histones at the miR-34a
promoter (short loop) and by increasing transactivation potential of FXR
via deacetylating the FXR (long loop). SIRT1 also increases FXR expression
and activity via deacetylation of PGC-1α.
FXR in turn positively regulates hepatic SIRT1 expression by inhibiting
miR-34a which targets SIRT1.
Concluding remarks
Because of SIRT1's anti-aging properties and its beneficial effects on a wide range of age-related disease [1-3,21], it has been intensively studied. SIRT1 levels were reported to be decreased in liver, muscle, and adipose tissues of diet-induced obese mice in vivo as well as in cultured cell models of insulin resistance [15,30,60], but the underlying mechanisms remain unclear. The discovery of the FXR/miR-34a pathway controlling SIRT1 levels provides a partial explanation since elevated miR-34a levels in obese mice contribute to decreased SIRT1 levels [30]. Based on these findings, together with the development of effective inhibitors of miRs, the antagomirs [4,24,38], it will be interesting to see whether the reduction of elevated miR-34a in fatty livers of obesity improves transcriptional profiles of metabolic genes and metabolic outcomes. Also, it will be important to understand how the FXR/SIRT1 regulatory network is dysregulated in metabolic disease states which likely involves altered cellular kinase signaling pathways that post-transcriptionally affect SIRT1 and FXR levels and activities. Development of drugs that target the FXR/miR-34a pathway and other miRs controlling SIRT1 expression may lead to novel therapeutic options for treating age-related metabolic disease including fatty liver, obesity and type II diabetes.
References
- 1. Haigis MC and Sinclair DA. Mammalian sirtuins: biological insights and disease relevance. Annu Rev Pathol. 2010; 5: 253 -295. [PubMed] .
- 2. Donmez G and Guarente L. Aging and disease: connections to sirtuins. Aging Cell. 2010; 9: 285 -290. [PubMed] .
- 3. Brooks CL and Gu W. Anti-aging protein SIRT1: a role in cervical cancer. Aging. 2009; 1: 278 -280. [PubMed] .
- 4. Krutzfeldt J , Rajewsky N , Braich R , Rajeev KG , Tuschl T , Manoharan M and Stoffel M. Silencing of microRNAs in vivo with 'antagomirs'. Nature. 2005; 438: 685 -689. [PubMed] .
- 5. Neilson JR and Sharp PA. Small RNA regulators of gene expression. Cell. 2008; 134: 899 -902. [PubMed] .
- 6. Hobert O Gene regulation by transcription factors and microRNAs. Science. 2008; 319: 1785 -1786. [PubMed] .
- 7. Canto C and Auwerx J. Caloric restriction, SIRT1 and longevity. Trends Endocrinol Metab. 2009; 20: 325 -231. [PubMed] .
- 8. Rodgers JT , Lerin C , Haas W , Gygi SP , Spiegelman BM and Puigserver P. Nutrient control of glucose homeostasis through a complex of PGC-1alpha and SIRT1. Nature. 2005; 434: 113 -118. [PubMed] .
- 9. Luo J , Nikolaev AY , Imai S , Chen D , Su F , Shiloh A , Guarente L and Gu W. Negative control of p53 by Sir2alpha promotes cell survival under stress. Cell. 2001; 107: 137 -148. [PubMed] .
- 10. Li X , Zhang S , Blander G , Tse JG , Krieger M and Guarente L. SIRT1 deacetylates and positively regulates the nuclear receptor LXR. Mol Cell. 2007; 28: 91 -106. [PubMed] .
- 11. Kemper JK , Xiao Z , Ponugoti B , Miao J , Fang S , Kanamaluru D , Tsang S , Wu S , Chiang CM and Veenstra TD. FXR acetylation is normally dynamically regulated by p300 and SIRT1 but constitutively elevated in metabolic disease states. Cell Metabolism. 2009; 10: 392 -404. [PubMed] .
- 12. Pfluger PT , Herranz D , Velasco-Miguel S , Serrano M and Tschop MH. Sirt1 protects against high-fat diet-induced metabolic damage. Proc Natl Acad Sci U S A. 2008; 105: 9793 -9798. [PubMed] .
- 13. Bordone L , Motta MC , Picard F , Robinson A , Jhala US , Apfeld J , McDonagh T , Lemieux M , McBurney M , Szilvasi A , Easlon EJ , Lin SJ and Guarente L. Sirt1 regulates insulin secretion by repressing UCP2 in pancreatic beta cells. PLoS Biol. 2006; 4: e31 [PubMed] .
- 14. Picard F , Kurtev M , Chung N , Topark-Ngarm A , Senawong T , Machado De Oliveira R , Leid M , McBurney MW and Guarente L. Sirt1 promotes fat mobilization in white adipocytes by repressing PPAR-gamma. Nature. 2004; 429: 771 -776. [PubMed] .
- 15. Sun C , Zhang F , Ge X , Yan T , Chen X , Shi X and Zhai Q. SIRT1 improves insulin sensitivity under insulin-resistant conditions by repressing PTP1B. Cell Metab. 2007; 6: 307 -319. [PubMed] .
- 16. Feige JN , Lagouge M , Canto C , Strehle A , Houten SM , Milne JC , Lambert PD , Mataki C , Elliott PJ and Auwerx J. Specific SIRT1 activation mimics low energy levels and protects against diet-induced metabolic disorders by enhancing fat oxidation. Cell Metab. 2008; 8: 347 -358. [PubMed] .
- 17. Lagouge M , Argmann C , Gerhart-Hines Z , Meziane H , Lerin C , Daussin F , Messadeq N , Milne J , Lambert P , Elliott P , Geny B , Laakso M and Puigserver P. Resveratrol improves mitochon-drial function and protects against metabolic disease by activating SIRT1 and PGC-1alpha. Cell. 2006; 127: 1109 -1122. [PubMed] .
- 18. Baur JA , Pearson KJ , Price NL , Jamieson HA , Lerin C , Kalra A , Prabhu VV , Allard JS , Lopez-Lluch G , Lewis K , Pistell PJ , Poosala S and Becker KG. Resveratrol improves health and survival of mice on a high-calorie diet. Nature. 2006; 444: 337 -342. [PubMed] .
- 19. Purushotham A , Schug TT , Xu Q , Surapureddi S , Guo X and Li X. Hepatocyte-specific deletion of SIRT1 alters fatty acid metabolism and results in hepatic steatosis and inflammation. Cell Metab. 2009; 9: 327 -338. [PubMed] .
- 20. Bordone L , Cohen D , Robinson A , Motta MC , van Veen E , Czopik A , Steele AD , Crowe H , Marmor S , Luo J , Gu W and Guarente L. SIRT1 transgenic mice show phenotypes resembling calorie restriction. Aging Cell. 2007; 6: 759 -767. [PubMed] .
- 21. Herranz D and Serrano M. Impact of Sirt1 on mammalian aging. Aging. 2010; 2: 315 -316. [PubMed] .
- 22. Purushotham A , Schug TT and Li X. SIRT1 performs a balancing act on the tight-rope toward longevity. Aging. 2009; 1: 669 -673. [PubMed] .
- 23. Stein S , Schafer N , Breitenstein A , Besler C , Winnik S , Lohmann C , Heinrich K , Brokopp CE , Handschin C , Landmesser U , Tanner FC , Luscher TF and Matter CM. SIRT1 reduces endothelial activation without affecting vascular function in ApoE-/- mice. Aging. 2010; 2: 353 -360. [PubMed] .
- 24. Esau C , Davis S , Murray SF , Yu XX , Pandey SK , Pear M , Watts L , Booten SL , Graham M , McKay R , Subramaniam A , Propp S and Lollo BA. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab. 2006; 3: 87 -98. [PubMed] .
- 25. Najafi-Shoushtari SH , Kristo F , Li Y , Shioda T , Cohen DE , Gerszten RE and Naar AM. MicroRNA-33 and the SREBP host genes cooperate to control cholesterol homeostasis. Science. 2010; 328: 1566 -1569. [PubMed] .
- 26. Rayner KJ , Suarez Y , Davalos A , Parathath S , Fitzgerald ML , Tamehiro N , Fisher EA , Moore KJ and Fernandez-Hernando C. MiR-33 contributes to the regulation of cholesterol homeostasis. Science. 2010; 328: 1570 -1573. [PubMed] .
- 27. Baroukh N , Ravier MA , Loder MK , Hill EV , Bounacer A , Scharfmann R , Rutter GA and Van Obberghen E. MicroRNA-124a regulates Foxa2 expression and intracellular signaling in pancreatic beta-cell lines. J Biol Chem. 2007; 282: 19575 -19588. [PubMed] .
- 28. Gatfield D , Le Martelot G , Vejnar CE , Gerlach D , Schaad O , Fleury-Olela F , Ruskeepaa AL , Oresic M , Esau CC , Zdobnov EM and Schibler U. Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev. 2009; 23: 1313 -1326. [PubMed] .
- 29. Iliopoulos D , Drosatos K , Hiyama Y , Goldberg IJ and Zannis VI. MicroRNA-370 controls the expression of MicroRNA-122 and Cpt1alpha and affects lipid metabolism. J Lipid Res. 2010; 51: 1513 -1523. [PubMed] .
- 30. Lee J , Padhye A , Sharma A , Song G , Miao J , Mo YY , Wang L and Kemper JK. A pathway involving farnesoid X receptor and small heterodimer partner positively regulates hepatic sirtuin 1 levels via microRNA-34a inhibition. J Biol Chem. 2010; 285: 12604 -12611. [PubMed] .
- 31. Poy MN , Eliasson L , Krutzfeldt J , Kuwajima S , Ma X , Macdonald PE , Pfeffer S , Tuschl T , Rajewsky N , Rorsman P and Stoffel M. A pancreatic islet-specific microRNA regulates insulin secretion. Nature. 2004; 432: 226 -230. [PubMed] .
- 32. Poy MN , Hausser J , Trajkovski M , Braun M , Collins S , Rorsman P , Zavolan M and Stoffel M. miR-375 maintains normal pancreatic alpha- and beta-cell mass. Proc Natl Acad Sci U S A. 2009; 106: 5813 -5818. [PubMed] .
- 33. Lovis P , Roggli E , Laybutt DR , Gattesco S , Yang JY , Widmann C , Abderrahmani A and Regazzi R. Alterations in microRNA expression contribute to fatty acid-induced pancreatic beta-cell dysfunction. Diabetes. 2008; 57: 2728 -2736. [PubMed] .
- 34. Lin Q , Gao Z , Alarcon RM , Ye J and Yun Z. A role of miR-27 in the regulation of adipogenesis. Febs J. 2009; 276: 2348 -2358. [PubMed] .
- 35. Gerin I , Bommer GT , McCoin CS , Sousa KM , Krishnan V and MacDougald OA. Roles for miRNA-378/378* in adipocyte gene expression and lipogenesis. Am J Physiol Endocrinol Metab. 2010; 299: E198 -206. [PubMed] .
- 36. Lu H , Buchan RJ and Cook SA. MicroRNA-223 regulates Glut4 expression and cardiomyocyte glucose metabolism. Cardiovasc Res. 2010; 86: 410 -420. [PubMed] .
- 37. Aoi W , Naito Y , Mizushima K , Takanami Y , Kawai Y , Ichikawa H and Yoshikawa T. The microRNA miR-696 regulates PGC-1{alpha} in mouse skeletal muscle in response to physical activity. Am J Physiol Endocrinol Metab. 2010; 298: E799 -806. [PubMed] .
- 38. Kota J , Chivukula RR , O'Donnell KA , Wentzel EA , Montgomery CL , Hwang HW , Chang TC , Vivekanandan P , Torbenson M , Clark KR , Mendell JR and Mendell JT. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009; 137: 1005 -1017. [PubMed] .
- 39. Cheung O , Puri P , Eicken C , Contos MJ , Mirshahi F , Maher JW , Kellum JM , Min H , Luketic VA and Sanyal AJ. Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression. Hepatology. 2008; 48: 1810 -1820. [PubMed] .
- 40. Nakanishi N , Nakagawa Y , Tokushige N , Aoki N , Matsuzaka T , Ishii K , Yahagi N , Kobayashi K , Yatoh S , Takahashi A , Suzuki H , Urayama O and Yamada N. The up-regulation of microRNA-335 is associated with lipid metabolism in liver and white adipose tissue of genetically obese mice. Biochem Biophys Res Commun. 2009; 385: 492 -496. [PubMed] .
- 41. Kwon HS and Ott M. The ups and downs of SIRT1. Trends Biochem Sci. 2008; 33: 517 -525. [PubMed] .
- 42. Yamakuchi M , Ferlito M and Lowenstein CJ. miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci U S A. 2008; 105: 13421 -13426. [PubMed] .
- 43. Strum JC , Johnson JH , Ward J , Xie H , Feild J , Hester A , Alford A and Waters KM. MicroRNA 132 regulates nutritional stress-induced chemokine production through repression of SirT1. Mol Endocrinol. 2009; 23: 1876 -1884. [PubMed] .
- 44. Rane S , He M , Sayed D , Vashistha H , Malhotra A , Sadoshima J , Vatner DE , Vatner SF and Abdellatif M. Downregulation of miR-199a derepresses hypoxia-inducible factor-1alpha and Sirtuin 1 and recapitulates hypoxia preconditioning in cardiac myocytes. Circ Res. 2009; 104: 879 -886. [PubMed] .
- 45. Saunders LR , Sharma AD , Tawney J , Nakagawa M , Okita K , Yamanaka S , Willenbring H and Verdin E. miRNAs regulate SIRT1 expression during mouse embryonic stem cell differentiation and in adult mouse tissues. Aging. 2010; Jul, 415 -432. .
- 46. Cariou B and Staels B. FXR: a promising target for the metabolic syndrome. Trends Pharmacol Sci. 2007; 28: 236 -243. [PubMed] .
- 47. Sinal C , Tohkin M , Miyata M , Ward J , Lambert G and Gonzalez FJ. Targeted disruption of the nuclear receptor FXR/BAR impairs bile acid and lipid homeostasis. Cell. 2000; 102: 731 -744. [PubMed] .
- 48. Moschetta A , Bookout AL and Mangelsdorf DJ. Prevention of cholesterol gallstone disease by FXR agonists in a mouse model. Nat Med. 2004; 10: 1352 -1358. [PubMed] .
- 49. Ma K , Saha PK , Chan L and Moore DD. Farnesoid X receptor is essential for normal glucose homeostasis. J Clin Invest. 2006; 116: 1102 -9. [PubMed] .
- 50. Zhang Y , Lee FY , Barrera G , Lee H , Vales C , Gonzalez FJ , Willson TM and Edwards PA. Activation of the nuclear receptor FXR improves hyperglycemia and hyperlipidemia in diabetic mice. Proc Natl Acad Sci U S A. 2006; 103: 1006 -1011. [PubMed] .
- 51. Goodwin B , Jones SA , Price RR , Watson MA , McKee DD , Moore LB , Galardi C , Wilson JG , Lewis MC , Roth ME , Maloney PR , Wilson TM and Kliewer SA. A regulatory cascade of the nuclear receptors FXR, SHP-1, and LRH-1 represses bile acid biosynthesis. Molecular Cell. 2000; 6: 517 -526. [PubMed] .
- 52. Lu TT , Makishima M , Repa JJ , Schoonjans K , Kerr TA , Auwerx J and Mangelsdorf DJ. Molecular basis for feedback regulation of bile acid synthesis by nuclear receptors. Mol Cell. 2000; 6: 507 -515. [PubMed] .
- 53. Chang TC , Wentzel EA , Kent OA , Ramachandran K , Mullendore M , Lee KH , Feldmann G , Yamakuchi M , Ferlito M , Lowenstein CJ , Arking DE , Beer MA and Maitra A. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007; 26: 745 -752. [PubMed] .
- 54. Watanabe M , Houten SM , Wang L , Moschetta A , Mangelsdorf DJ , Heyman RA , Moore DD and Auwerx J. Bile acids lower triglyceride levels via a pathway involving FXR, SHP, and SREBP-1c. Journal of Clinical Investigation. 2004; 113: 1408 -1418. [PubMed] .
- 55. Fang S , Miao J , Xiang L , Ponugoti B , Treuter E and Kemper JK. Coordinated recruitment of histone methyltransferase G9a and other chromatin-modifying enzymes in SHP-mediated regulation of hepatic bile acid metabolism. Mol Cell Biol. 2007; 27: 1407 -1424. [PubMed] .
- 56. Kemper J , Kim H , Miao J , Bhalla S and Bae Y. Role of a mSin3A-Swi/Snf chromatin remodeling complex in the feedback repression of bile acid biosynthesis by SHP. Mol Cell Biol. 2004; 24: 7707 -7719. [PubMed] .
- 57. Miao J , Fang S , Lee J , Comstock C , Knudsen KE and Kemper JK. Functional specificities of Brm and Brg-1 Swi/Snf ATPases in the feedback regulation of hepatic bile acid biosynthesis. Mol Cell Biol. 2009; 29: 6170 -6181. [PubMed] .
- 58. Yamakuchi M and Lowenstein CJ. MiR-34, SIRT1 and p53: the feedback loop. Cell Cycle. 2009; 8: 712 -715. [PubMed] .
- 59. Zhang Y , Castellani LW , Sinal CJ , Gonzalez FJ and Edwards PA. Peroxisome proliferator-activated receptor-gamma coactivator 1alpha (PGC-1alpha) regulates triglyceride metabolism by activation of the nuclear receptor FXR. Genes Dev. 2004; 18: 157 -169. [PubMed] .
- 60. Coste A , Louet JF , Lagouge M , Lerin C , Antal MC , Meziane H , Schoonjans K , Puigserver P , O'Malley BW and Auwerx J. The genetic ablation of SRC-3 protects against obesity and improves insulin sensitivity by reducing the acetylation of PGC-1{alpha}. Proc Natl Acad Sci U S A. 2008; 105: 17187 -17192. [PubMed] .


