Introduction
Aging and longevity have always been an important issue in biological sciences. Several reports have indicated that calorie restriction leads to lifespan extension from yeast to humans [1-3]. Longevity effect is also seen when nutrient-sensing signaling pathways such as the target of rapamycin (TOR) pathway and insulin/IGF pathway are inhibited [4-7], and the molecular mechanisms that regulate these nutrient-dependent pathways can substantially promote longevity in various organisms and mammalian cells [8-13]. The budding yeast Saccharomyces cerevisiae provides valuable information on understanding aging process and identifying the molecular mechanisms that regulate longevity in eukaryotic organisms. Two types of aging, replicative and chronological, have been described in S. cerevisiae. Replicative aging is an aging model of mitotically active cells in which the lifespan of a mother cell is measured by the number of daughter cells produced before death [14, 15]. Chronological aging is an aging model of post-mitotic cells in which lifespan is defined by the survival time of cells in a non-dividing state [16]. In S. cerevisiae, a well-established mechanism for regulating longevity is the alteration of ribosomal DNA (rDNA) stability by the recombination between rDNA repeats and the accumulation of extrachromosomal rDNA circles (ERCs) [17]. Recently, we confirmed that the inhibition of TOR pathway stabilizes rDNA locus by enhancing the association of silent information regulator 2 (Sir2) with rDNA, thereby leading to replicative lifespan extension in S. cerevisiae [18]. In this research perspective, we review the implication of rDNA stabilizers in longevity regulation, discuss how Sir2 stabilizes rDNA under TOR inhibition and speculate on the link between sumoylation and Sir2-related aging pathway.
Sir2 and the epigenetic regulators of yeast rDNA locus
Sir2 proteins, or sirtuins, are a well-known, highly conserved, family of NAD+-dependent deacetylases that are involved in the regulation of lifespan from yeast to humans [19-22]. Many reports have shown that sirtuins are linked to multiple physiological processes including metabolic regulation, DNA repair, stress response, apoptosis, cell survival and longevity [23-31]. Sirtuins also mediate the increased spontaneous physical activity in flies on calorie restriction and regulate p53 function via deacetylation in human cells [32, 33]. In S. cerevisiae, Sir2 is a functional component of epigenetic complexes required for the establishment of silenced chromatin regions in rDNA locus, telomeres and silent mating-type loci, HML and HMR [34-38]. S. cerevisiae rDNA consists of a 9.1-kb unit that contains RNA polymerase (Pol) I-transcribed 35S ribosomal RNA (rRNA) and Pol III-transcribed 5S rRNA gene, separated by a non-transcribed spacer, and is repeated 100-200 times on chromosome XII [39]. Because of highly repetitive nature, rDNA array is intrinsically unstable and is an easy target for homologous recombination. A primary cause of aging in S. cerevisiae is known to be the homologous recombination between rDNA repeats, which leads to the formation of ERCs that accumulate to toxic levels in mother cells [17]. Sir2 promotes replicative lifespan by repressing the homologous recombination between rDNA repeats and the subsequent formation of ERCs [40].
A well-known epigenetic regulator of yeast rDNA locus is the regulator of nucleolar silencing and telophase exit (RENT) complex that is composed of Sir2, Net1 and Cdc14 [41]. Net1, the core subunit of this complex, localizes to rDNA, recruits Sir2 to rDNA and is required for transcriptional silencing at rDNA [41-43]. Interestingly, several other epigenetic regulatory proteins also seem to be linked to rDNA stability and longevity. Tof2 interacts with Net1 and Sir2, binds to rDNA and induces rDNA silencing [43]. Lrs4 and Csm1, two of three subunits of monopolin complex that co-orients sister chromatids during meiosis I, interact with Tof2, associate with rDNA, establish rDNA silencing and suppress unequal recombination at rDNA [43-45]. Heh1 and Nur1, chromosome linkage inner nuclear membrane proteins, are physically linked to Lrs4 and Csm1, tether rDNA to nuclear periphery and promote rDNA stability [46]. Yeast linker histone Hho1, histone acetyltransferase Ada2 and Esa1 are required for rDNA silencing [47-49]. Histone methyltransferase Set1 is required for rDNA silencing in a Sir2-independent manner [50, 51]. Chromatin remodeling proteins, such as Snf2, Fun30, Isw1 and Isw2, establish silencing at rDNA and maintain rDNA chromatin structure [52-56]. Collectively, these findings suggest that rDNA silencing factors, histone-modifying enzymes and chromatin remodeling factors promote rDNA stability and regulate ERC-mediated aging in yeast.
Sir2 is part of a pathway mediating the longevity effect of TORC1 inhibition
TOR kinase is a nutrient-responsive phosphatidylinositol kinase-related protein kinase structurally and functionally conserved from yeast to humans and plays critical roles in cell growth in response to nutrient availability by regulating transcription, translation, ribosome biogenesis and autophagy [13, 57-59]. In yeast, TOR kinase exists in two functionally distinct multiprotein complexes, TOR complex1 (TORC1) and TOR complex2 (TORC2), each of which signals via a different set of effector pathways [60]. An immunosuppressive and anticancer drug rapamycin specifically inhibits TORC1 and leads to a rapid decrease in ribosome biogenesis by regulating the transcription of all three kinds of RNA Pols [61, 62]. In past decade, it has been reported that TORC1 signaling is deeply involved in eukaryotic cell aging and aging-related diseases. The inhibition of TORC1 signaling by rapamycin can delay aging and prolong lifespan in S. cerevisiae, Caenorhabditis elegans and Drosophila melanogaster [63-66]. Recent studies have shown that rapamycin can significantly increase lifespan also in genetically heterogeneous mice [67, 68].
How the inhibition of TORC1 signaling extends lifespan is poorly understood. Meanwhile, whether Sir2 functions in rDNA stability and lifespan extension in yeast during TORC1 inhibition has been a matter of debate. Kaeberlein et al. reported that the inhibition of TORC1 signaling increases lifespan but has no effect on Sir2 activity [65], whereas Medvedik et al. indicated that TORC1 inhibition activates Sir2 by increasing the expression of PNC1 encoding a nicotinamidase that depletes cellular nicotinamide, a physiological inhibitor of Sir2, thereby suppressing the formation of ERCs and leading to the extension of replicative lifespan [66]. Corroborating the latter study, we found that the inhibition of TORC1 signaling increases the association of Sir2 with rDNA in Pnc1- and Net1-dependent manners, enhances the transcriptional silencing of Pol II-transcribed genes at rDNA and induces the deacetylation of histones at rDNA, thereby promoting rDNA stability and replicative lifespan extension [18]. These findings suggest that Sir2 contributes to lifespan extension by enhancing rDNA silencing and rDNA stability under TORC1 inhibition.
Recent studies have reported that Sir2 inhibits replicative senescence by additional non-rDNA mechanisms. Sir2 is required for the asymmetric inheritance of oxidatively damaged proteins during cytokinesis, resulting in an enhanced capacity to respond to oxidative stress in daughter cells [69-72]. In addition, an age-associated decrease in Sir2 protein abundance is accompanied by the increase in histone acetylation and the loss of histones at specific subtelomeric regions in replicatively old yeast cells, indicating that Sir2 regulates replicative longevity through the maintenance of telomeric chromatin [73]. Whether TORC1 signaling modulates these non-rDNA functions of Sir2 is not known yet. It will be interesting to check whether the Sir2-dependent asymmetric inheritance of oxidatively damaged proteins and the Sir2-dependent maintenance of telomeric chromatin are influenced by TORC1 signaling.
Sumoylation: a potential mechanism for the regulation of Sir2 function in rDNA maintenance
Small ubiquitin-like modifier (SUMO) modification is an important posttranslational mechanism for regulating protein function, especially in the nucleus and nucleolus. Sumoylation is involved in various nuclear functions such as transcription, DNA repair and nuclear domain organization [74, 75]. In yeast, SUMO is prominently enriched in the nucleolus and affects rDNA segregation and maintenance [76, 77]. Topoisomerase Top1, which is sumoylated by Siz1 and Siz2, facilitates rDNA transcription and replication, is required for rDNA silencing and maintains rDNA integrity [77, 78]. SUMO ligase Mms21, a subunit of the Smc5/Smc6 complex, binds to rDNA and maintains rDNA stability [79, 80]. Moreover, cohesin and condensin subunits, which play important roles in rDNA stability and structures, are sumoylated by Mms21, and the association of these subunits with rDNA is regulated by Mms21 [77]. Additionally, sumoylation of Rad52 suppresses rDNA recombination and affects the efficiency of recombinational DNA repair [79, 81]. These findings raise an interesting possibility that sumoylation may contribute to maintaining stable rDNA structure and thus promoting longevity.
Recent studies have shown that sumoylation is involved in the regulation of Sir2 function in the maintenance of chromatin structure at rDNA. Esc2, a member of a conserved family of proteins that contain SUMO-like domains, interacts with Sir2 through a SUMO-binding motif and is required for the maintenance of silent chromatin structure at rDNA [82]. A SUMO-binding motif of Esc2 is necessary and sufficient for interacting with both Sir2 and SUMO, and is required for the function of Esc2 in transcriptional silencing. These results raise a possibility that Sir2 is a sumoylated protein and Esc2 binds to sumoylated Sir2 via its SUMO-binding motif. A previous study has reported that the human sirtuin SIRT1 is sumoylated and this modification increases its deacetylase activity [83]. Although the sumoylation of yeast Sir2 has not been established yet, it is plausible that the activity of Sir2 may be regulated by its sumoylation. If Sir2 is not sumoylated, it is presumable that some sumoylated proteins may mediate its interaction with other proteins such as Esc2 and regulate its activity.
Conclusion
Sirtuins are a highly conserved family of proteins that have been implicated in regulating diverse functions including longevity in a variety of species. Many studies suggest that Sir2 regulates rDNA recombination, DNA repair, chromosomal stability and longevity. In this research perspective, we briefly reviewed how Sir2 and other epigenetic regulators function as the stabilizer of rDNA and regulate longevity. Sir2 is part of a pathway mediating the longevity effect of TORC1 inhibition. Based on recent findings, we also propose that sumoylation may be involved in the regulation of Sir2 function in rDNA maintenance and thus longevity (Figure 1). Given that the human sirtuin SIRT1 is sumoylated and this modification increases its deacetylase activity [83], it will be interesting to investigate whether Sir2 and other sirtuins are sumoylated and, if so, whether their sumoylation is influenced by TORC1 signaling and how their functions are affected by sumoylation. These studies will provide deeper understanding of the mechanisms by which Sir2 and TORC1 signaling regulate longevity.
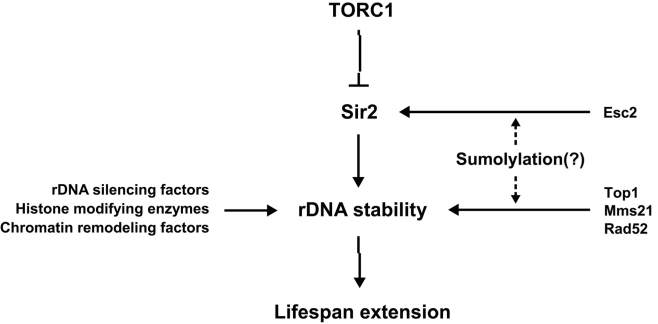
Figure 1. A model for the link between sumoylation and Sir2-related aging pathway Various factors stabilize rDNA to affect lifespan extension in S. cerevisiae. TORC1 inhibition, Sir2 and several epigenetic regulators of rDNA promote rDNA stability and longevity. Sumoylation may regulate the activity of Sir2 and contribute to maintaining stable rDNA structure, leading to lifespan extension.
Acknowledgments
This work was supported by the Basic Science Research Program through the Nation Research Foundation of Korea [2010-0013678] and the SRC/ERC Program of MOST/KOSEF, Republic of Korea. C.W.H. was supported by the BK21 ResearchFellowship from the Ministry of Education, Science and Technology, Republic of Korea.
Conflicts of Interest
The authors of this manuscript have no conflict of interests to declare.
References
- 1. Kennedy BK, Steffen KK, Kaeberlein M. Ruminations on dietary restriction and aging. Cell Mol Life Sci. 2007; 64: 1323 -1328. [PubMed] .
- 2. Fontana L, Partridge L, Longo VD. Extending healthy life span--from yeast to humans. Science. 2010; 328: 321 -326. [PubMed] .
- 3. Bishop NA and Guarente L. Genetic links between diet and lifespan: shared mechanisms from yeast to humans. Nat Rev Genet. 2007; 8: 835 -844. [PubMed] .
- 4. Kenyon C. The plasticity of aging: insights from long-lived mutants. Cell. 2005; 120: 449 -460. [PubMed] .
- 5. Clancy DJ, Gems D, Hafen E, Leevers SJ, Partridge L. Dietary restriction in long-lived dwarf flies. Science. 2002; 296: 319 [PubMed] .
- 6. Flurkey K, Papaconstantinou J, Miller RA, Harrison DE. Lifespan extension and delayed immune and collagen aging in mutant mice with defects in growth hormone production. Proc Natl Acad Sci U S A. 2001; 98: 6736 -6741. [PubMed] .
- 7. Longo VD. Linking sirtuins, IGF-I signaling, and starvation. Exp Gerontol. 2009; 44: 70 -74. [PubMed] .
- 8. Pan Y and Shadel GS. Extension of chronological life span by reduced TOR signaling requires down-regulation of Sch9p and involves increased mitochondrial OXPHOS complex density. Aging (Albany NY). 2009; 1: 131 -145. [PubMed] .
- 9. Heeren G, Rinnerthaler M, Laun P, von Seyerl P, Kossler S, Klinger H, Hager M, Bogengruber E, Jarolim S, Simon-Nobbe B, et al. The mitochondrial ribosomal protein of the large subunit, Afo1p, determines cellular longevity through mitochon-drial back-signaling via TOR1. Aging (Albany NY). 2009; 1: 622 -636. [PubMed] .
- 10. Korotchkina LG, Leontieva OV, Bukreeva EI, Demidenko ZN, Gudkov AV, Blagosklonny MV. The choice between p53-induced senescence and quiescence is determined in part by the mTOR pathway. Aging (Albany NY). 2010; 2: 344 -352. [PubMed] .
- 11. Lee JH, Bodmer R, Bier E, Karin M. Sestrins at the crossroad between stress and aging. Aging (Albany NY). 2010; 2: 369 -374. [PubMed] .
- 12. Goldberg AA, Richard VR, Kyryakov P, Bourque SD, Beach A, Burstein MT, Glebov A, Koupaki O, Boukh-Viner T, Gregg C, et al. Chemical genetic screen identifies lithocholic acid as an anti-aging compound that extends yeast chronological life span in a TOR-independent manner, by modulating housekeeping longevity assurance processes. Aging (Albany NY). 2010; 2: 393 -414. [PubMed] .
- 13. Hands SL, Proud CG, Wyttenbach A. mTOR's role in ageing: protein synthesis or autophagy? Aging (Albany NY). 2009; 1: 586 -597. [PubMed] .
- 14. Kaeberlein M. Lessons on longevity from budding yeast. Nature. 2010; 464: 513 -519. [PubMed] .
- 15. Steinkraus KA, Kaeberlein M, Kennedy BK. Replicative aging in yeast: the means to the end. Annu Rev Cell Dev Biol. 2008; 24: 29 -54. [PubMed] .
- 16. Kennedy BK, Smith ED, Kaeberlein M. The enigmatic role of Sir2 in aging. Cell. 2005; 123: 548 -550. [PubMed] .
- 17. Sinclair DA and Guarente L. Extrachromosomal rDNA circles--a cause of aging in yeast. Cell. 1997; 91: 1033 -1042. [PubMed] .
- 18. Ha CW and Huh WK. Rapamycin increases rDNA stability by enhancing association of Sir2 with rDNA in Saccharomyces cerevisiae. Nucleic Acids Res. 2011; 39: 1336 -1350. [PubMed] .
- 19. Finkel T, Deng CX, Mostoslavsky R. Recent progress in the biology and physiology of sirtuins. Nature. 2009; 460: 587 -591. [PubMed] .
- 20. Imai S, Armstrong CM, Kaeberlein M, Guarente L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature. 2000; 403: 795 -800. [PubMed] .
- 21. Landry J, Sutton A, Tafrov ST, Heller RC, Stebbins J, Pillus L, Sternglanz R. The silencing protein SIR2 and its homologs are NAD-dependent protein deacetylases. Proc Natl Acad Sci U S A. 2000; 97: 5807 -5811. [PubMed] .
- 22. Smith JS, Brachmann CB, Celic I, Kenna MA, Muhammad S, Starai VJ, Avalos JL, Escalante-Semerena JC, Grubmeyer C, Wolberger C, et al. A phylogenetically conserved NAD+-dependent protein deacetylase activity in the Sir2 protein family. Proc Natl Acad Sci U S A. 2000; 97: 6658 -6663. [PubMed] .
- 23. Blander G and Guarente L. The Sir2 family of protein deacetylases. Annu Rev Biochem. 2004; 73: 417 -435. [PubMed] .
- 24. Michan S and Sinclair D. Sirtuins in mammals: insights into their biological function. Biochem J. 2007; 404: 1 -13. [PubMed] .
- 25. Guarente L. Sirtuins as potential targets for metabolic syndrome. Nature. 2006; 444: 868 -874. [PubMed] .
- 26. Imai S and Guarente L. Ten years of NAD-dependent SIR2 family deacetylases: implications for metabolic diseases. Trends Pharmacol Sci. 2010; 31: 212 -220. [PubMed] .
- 27. Haigis MC and Sinclair DA. Mammalian sirtuins: biological insights and disease relevance. Annu Rev Pathol. 2010; 5: 253 -295. [PubMed] .
- 28. Bauer J, Antosh M, Chang C, Schorl C, Kolli S, Neretti N, Helfand SL. Comparative transcriptional profiling identifies takeout as a gene that regulates life span. Aging (Albany NY). 2010; 2: 298 -310. [PubMed] .
- 29. Bauer JH, Morris SN, Chang C, Flatt T, Wood JG, Helfand SL. dSir2 and Dmp53 interact to mediate aspects of CR-dependent lifespan extension in D. melanogaster. Aging (Albany NY). 2009; 1: 38 -48. [PubMed] .
- 30. Donehower LA. Longevity regulation in flies: a role for p53. Aging (Albany NY). 2009; 1: 6 -8. [PubMed] .
- 31. Frankel S, Ziafazeli T, Rogina B. dSir2 and longevity in Drosophila. Exp Gerontol. 2010; .
- 32. Parashar V and Rogina B. dSir2 mediates the increased spontaneous physical activity in flies on calorie restriction. Aging (Albany NY). 2009; 1: 529 -541. [PubMed] .
- 33. Vaziri H, Dessain SK, Ng Eaton E, Imai SI, Frye RA, Pandita TK, Guarente L, Weinberg RA. hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell. 2001; 107: 149 -159. [PubMed] .
- 34. Aparicio OM, Billington BL, Gottschling DE. Modifiers of position effect are shared between telomeric and silent mating-type loci in S. cerevisiae. Cell. 1991; 66: 1279 -1287. [PubMed] .
- 35. Rusche LN, Kirchmaier AL, Rine J. The establishment, inheritance, and function of silenced chromatin in Saccharomyces cerevisiae. Annu Rev Biochem. 2003; 72: 481 -516. [PubMed] .
- 36. Guarente L. Diverse and dynamic functions of the Sir silencing complex. Nat Genet. 1999; 23: 281 -285. [PubMed] .
- 37. Bryk M, Banerjee M, Murphy M, Knudsen KE, Garfinkel DJ, Curcio MJ. Transcriptional silencing of Ty1 elements in the RDN1 locus of yeast. Genes Dev. 1997; 11: 255 -269. [PubMed] .
- 38. Smith JS and Boeke JD. An unusual form of transcriptional silencing in yeast ribosomal DNA. Genes Dev. 1997; 11: 241 -254. [PubMed] .
- 39. Johnston M, Hillier L, Riles L, Albermann K, Andre B, Ansorge W, Benes V, Bruckner M, Delius H, Dubois E, et al. The nucleotide sequence of Saccharomyces cerevisiae chromosome XII. Nature. 1997; 387: 87 -90. [PubMed] .
- 40. Kaeberlein M, McVey M, Guarente L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 1999; 13: 2570 -2580. [PubMed] .
- 41. Huang J and Moazed D. Association of the RENT complex with nontranscribed and coding regions of rDNA and a regional requirement for the replication fork block protein Fob1 in rDNA silencing. Genes Dev. 2003; 17: 2162 -2176. [PubMed] .
- 42. Straight AF, Shou W, Dowd GJ, Turck CW, Deshaies RJ, Johnson AD, Moazed D. Net1, a Sir2-associated nucleolar protein required for rDNA silencing and nucleolar integrity. Cell. 1999; 97: 245 -256. [PubMed] .
- 43. Huang J, Brito IL, Villen J, Gygi SP, Amon A, Moazed D. Inhibition of homologous recombination by a cohesin-associated clamp complex recruited to the rDNA recombination enhancer. Genes Dev. 2006; 20: 2887 -2901. [PubMed] .
- 44. Toth A, Rabitsch KP, Galova M, Schleiffer A, Buonomo SB, Nasmyth K. Functional genomics identifies monopolin: a kinetochore protein required for segregation of homologs during meiosis i. Cell. 2000; 103: 1155 -1168. [PubMed] .
- 45. Rabitsch KP, Petronczki M, Javerzat JP, Genier S, Chwalla B, Schleiffer A, Tanaka TU, Nasmyth K. Kinetochore recruitment of two nucleolar proteins is required for homolog segregation in meiosis I. Dev Cell. 2003; 4: 535 -548. [PubMed] .
- 46. Mekhail K, Seebacher J, Gygi SP, Moazed D. Role for perinuclear chromosome tethering in maintenance of genome stability. Nature. 2008; 456: 667 -670. [PubMed] .
- 47. Levy A, Eyal M, Hershkovits G, Salmon-Divon M, Klutstein M, Katcoff DJ. Yeast linker histone Hho1p is required for efficient RNA polymerase I processivity and transcriptional silencing at the ribosomal DNA. Proc Natl Acad Sci U S A. 2008; 105: 11703 -11708. [PubMed] .
- 48. Jacobson S and Pillus L. The SAGA subunit Ada2 functions in transcriptional silencing. Mol Cell Biol. 2009; 29: 6033 -6045. [PubMed] .
- 49. Clarke AS, Samal E, Pillus L. Distinct roles for the essential MYST family HAT Esa1p in transcriptional silencing. Mol Biol Cell. 2006; 17: 1744 -1757. [PubMed] .
- 50. Bryk M, Briggs SD, Strahl BD, Curcio MJ, Allis CD, Winston F. Evidence that Set1, a factor required for methylation of histone H3, regulates rDNA silencing in S. cerevisiae by a Sir2-independent mechanism. Curr Biol. 2002; 12: 165 -170. [PubMed] .
- 51. Briggs SD, Bryk M, Strahl BD, Cheung WL, Davie JK, Dent SY, Winston F, Allis CD. Histone H3 lysine 4 methylation is mediated by Set1 and required for cell growth and rDNA silencing in Saccharomyces cerevisiae. Genes Dev. 2001; 15: 3286 -3295. [PubMed] .
- 52. Dror V and Winston F. The Swi/Snf chromatin remodeling complex is required for ribosomal DNA and telomeric silencing in Saccharomyces cerevisiae. Mol Cell Biol. 2004; 24: 8227 -8235. [PubMed] .
- 53. Mueller JE and Bryk M. Isw1 acts independently of the Isw1a and Isw1b complexes in regulating transcriptional silencing at the ribosomal DNA locus in Saccharomyces cerevisiae. J Mol Biol. 2007; 371: 1 -10. [PubMed] .
- 54. Mueller JE, Li C, Bryk M. Isw2 regulates gene silencing at the ribosomal DNA locus in Saccharomyces cerevisiae. Biochem Biophys Res Commun. 2007; 361: 1017 -1021. [PubMed] .
- 55. Saha A, Wittmeyer J, Cairns BR. Chromatin remodelling: the industrial revolution of DNA around histones. Nat Rev Mol Cell Biol. 2006; 7: 437 -447. [PubMed] .
- 56. Neves-Costa A, Will WR, Vetter AT, Miller JR, Varga-Weisz P. The SNF2-family member Fun30 promotes gene silencing in heterochromatic loci. PLoS One. 2009; 4: e8111 [PubMed] .
- 57. Schneper L, Duvel K, Broach JR. Sense and sensibility: nutritional response and signal integration in yeast. Curr Opin Microbiol. 2004; 7: 624 -630. [PubMed] .
- 58. Dennis PB, Fumagalli S, Thomas G. Target of rapamycin (TOR): balancing the opposing forces of protein synthesis and degradation. Curr Opin Genet Dev. 1999; 9: 49 -54. [PubMed] .
- 59. Schmelzle T and Hall MN. TOR, a central controller of cell growth. Cell. 2000; 103: 253 -262. [PubMed] .
- 60. Loewith R, Jacinto E, Wullschleger S, Lorberg A, Crespo JL, Bonenfant D, Oppliger W, Jenoe P, Hall MN. Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Mol Cell. 2002; 10: 457 -468. [PubMed] .
- 61. Mayer C and Grummt I. Ribosome biogenesis and cell growth: mTOR coordinates transcription by all three classes of nuclear RNA polymerases. Oncogene. 2006; 25: 6384 -6391. [PubMed] .
- 62. Powers T and Walter P. Regulation of ribosome biogenesis by the rapamycin-sensitive TOR-signaling pathway in Saccharomyces cerevisiae. Mol Biol Cell. 1999; 10: 987 -1000. [PubMed] .
- 63. Vellai T, Takacs-Vellai K, Zhang Y, Kovacs AL, Orosz L, Muller F. Genetics: influence of TOR kinase on lifespan in C. elegans. Nature. 2003; 426: 620 [PubMed] .
- 64. Kapahi P, Zid BM, Harper T, Koslover D, Sapin V, Benzer S. Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr Biol. 2004; 14: 885 -890. [PubMed] .
- 65. Kaeberlein M, Powers RW 3rd, Steffen KK, Westman EA, Hu D, Dang N, Kerr EO, Kirkland KT, Fields S, Kennedy BK. Regulation of yeast replicative life span by TOR and Sch9 in response to nutrients. Science. 2005; 310: 1193 -1196. [PubMed] .
- 66. Medvedik O, Lamming DW, Kim KD, Sinclair DA. MSN2 and MSN4 link calorie restriction and TOR to sirtuin-mediated lifespan extension in Saccharomyces cerevisiae. PLoS Biol. 2007; 5: e261 [PubMed] .
- 67. Harrison DE, Strong R, Sharp ZD, Nelson JF, Astle CM, Flurkey K, Nadon NL, Wilkinson JE, Frenkel K, Carter CS, et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature. 2009; 460: 392 -395. [PubMed] .
- 68. Miller RA, Harrison DE, Astle CM, Baur JA, Boyd AR, de Cabo R, Fernandez E, Flurkey K, Javors MA, Nelson JF, et al. Rapamycin, But Not Resveratrol or Simvastatin, Extends Life Span of Genetically Heterogeneous Mice. J Gerontol A Biol Sci Med Sci. 2010; .
- 69. Aguilaniu H, Gustafsson L, Rigoulet M, Nystrom T. Asymmetric inheritance of oxidatively damaged proteins during cytokinesis. Science. 2003; 299: 1751 -1753. [PubMed] .
- 70. Erjavec N and Nystrom T. Sir2p-dependent protein segregation gives rise to a superior reactive oxygen species management in the progeny of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 2007; 104: 10877 -10881. [PubMed] .
- 71. Erjavec N, Cvijovic M, Klipp E, Nystrom T. Selective benefits of damage partitioning in unicellular systems and its effects on aging. Proc Natl Acad Sci U S A. 2008; 105: 18764 -18769. [PubMed] .
- 72. Hekimi S and Guarente L. Genetics and the specificity of the aging process. Science. 2003; 299: 1351 -1354. [PubMed] .
- 73. Dang W, Steffen KK, Perry R, Dorsey JA, Johnson FB, Shilatifard A, Kaeberlein M, Kennedy BK, Berger SL. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature. 2009; 459: 802 -807. [PubMed] .
- 74. Johnson ES. Protein modification by SUMO. Annu Rev Biochem. 2004; 73: 355 -382. [PubMed] .
- 75. Geiss-Friedlander R and Melchior F. Concepts in sumoylation: a decade on. Nat Rev Mol Cell Biol. 2007; 8: 947 -956. [PubMed] .
- 76. Takahashi Y and Strunnikov A. In vivo modeling of polysumoylation uncovers targeting of Topoisomerase II to the nucleolus via optimal level of SUMO modification. Chromosoma. 2008; 117: 189 -198. [PubMed] .
- 77. Takahashi Y, Dulev S, Liu X, Hiller NJ, Zhao X, Strunnikov A. Cooperation of sumoylated chromosomal proteins in rDNA maintenance. PLoS Genet. 2008; 4: e1000215 [PubMed] .
- 78. Smith JS, Caputo E, Boeke JD. A genetic screen for ribosomal DNA silencing defects identifies multiple DNA replication and chromatin-modulating factors. Mol Cell Biol. 1999; 19: 3184 -3197. [PubMed] .
- 79. Torres-Rosell J, Machin F, Farmer S, Jarmuz A, Eydmann T, Dalgaard JZ, Aragon L. SMC5 and SMC6 genes are required for the segregation of repetitive chromosome regions. Nat Cell Biol. 2005; 7: 412 -419. [PubMed] .
- 80. Zhao X and Blobel G. A SUMO ligase is part of a nuclear multiprotein complex that affects DNA repair and chromosomal organization. Proc Natl Acad Sci U S A. 2005; 102: 4777 -4782. [PubMed] .
- 81. Altmannova V, Eckert-Boulet N, Arneric M, Kolesar P, Chaloupkova R, Damborsky J, Sung P, Zhao X, Lisby M, Krejci L. Rad52 SUMOylation affects the efficiency of the DNA repair. Nucleic Acids Res. 2010; 38: 4708 -4721. [PubMed] .
- 82. Yu Q, Kuzmiak H, Olsen L, Kulkarni A, Fink E, Zou Y, Bi X. Saccharomyces cerevisiae Esc2p interacts with Sir2p through a small ubiquitin-like modifier (SUMO)-binding motif and regulates transcriptionally silent chromatin in a locus-dependent manner. J Biol Chem. 2010; 285: 7525 -7536. [PubMed] .
- 83. Yang Y, Fu W, Chen J, Olashaw N, Zhang X, Nicosia SV, Bhalla K, Bai W. SIRT1 sumoylation regulates its deacetylase activity and cellular response to genotoxic stress. Nat Cell Biol. 2007; 9: 1253 -1262. [PubMed] .


