Significance
Despite promising results from candidate gene studies, a systematic and comprehensive examination of genetic determinants of gait speed in a large sample of older adults has been lacking. Furthermore, previous study samples have been too small to detect the expected modest genetic effects especially in such complex and polygenic encoded traits. To address these limitations, we conducted a meta-analysis of GWAS of gait speed in 31,478 older adults and validate our candidate signal in a cohort of 2588 older adults. Close to 600 candidate genetic variants have been linked to gait speed. Such efforts have provided us with an increased knowledge of the biological systems which impact on gait speed; this may contribute to improved treatment strategies and drug development to promote aging with grace.
Introduction
Gait speed has been described as the “sixth vital sign” because it is a core indicator of health and function in aging and disease [1]. Decline in gait speed is ubiquitous with aging in both men and women [2]. Gait speed is used to establish thresholds in community based activities, such as crossing a street [3,4] or ambulating [5–7]. Slow gait speed is a consistent risk factor for disability, cognitive impairment, institutionalization, falls, hospitalization and mortality [8–10]. Improvement in gait speed is associated with better function and survival.
Many genetic and non-genetic factors (environment and disease) are likely to affect quantitative complex traits such as gait speed. There are individual differences in rates of decline in physical function, and genetic epidemiological studies provide a method for decomposing that variance into genetic and environmental sources. Twin studies suggest that genetic factors account for 15-51% of the variance of gait speed in older adults [11,12]. Moreover, the contribution of genetic factors may increase with age [2,11,13–15]. Offspring of parents with exceptional longevity have better physical function and gait speed in age-specific comparisons to other individuals of comparable age and other characteristics [16,17]. Effective gait requires the integration of many physiological systems, including the central and peripheral nervous system that create and execute the motor program, the musculoskeletal system that moves and supports the body, and the cardio-pulmonary function that provides perfusion of adequate nutrients and oxygen to all of the integrated parts. All these physiological systems can be affected by genetic variation. Given the many pathways that may contribute to gait impairment, effect sizes of individual genetic variants are expected to be limited.
Previous candidate gene studies have implicated several loci as relevant to gait speed. Single nucleotide polymorphisms (SNP) in the Angiotensin-Converting Enzyme (ACE) gene have been linked to better mobility response to exercise. The R577X polymorphism in the alpha-actinin-3 encoding gene (ACTN3) was associated with elite athletic performance, and muscle strength and power in the general population, especially in women [18]. There is evidence that ACE I/D and ACTN3 R577X polymorphisms, individually or in combination, have a significant influence on mobility and gait speed phenotypes in older women [19,20]. Catechol-O-methyltransferase (COMT) polymorphisms have been associated with cognitive functions and gait speed [21]. The Met (158) Val polymorphism in COMT was linked to faster gait speed in older adults [21]. In addition, apolipoprotein E (APOE) genetic variation has been shown to influence the risk of gait speed decline [22–24]. Despite these promising results from candidate gene studies, a systematic and comprehensive examination of genetic determinants of gait speed in a large sample of older adults has been lacking. Furthermore, previous study samples have been too small to detect the expected modest genetic effects [25] especially in such complex and polygenic encoded traits [26].
To address these limitations, we conducted a meta-analysis of GWAS studies of gait speed in 31,478 older adults from the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) consortium. We then tested our findings in a validation cohort of 2588 older adults participating in four independent studies.
Results
Gait speed is considered a marker of health and fitness in aging. Slow gait in older adults is associated with increased risk of multiple adverse events including loss of independence, increased risk of disability, falls [27,28], progression of age-related disease including dementia [29] and death [9]. Slowing of gait is multifactorial with major contributions from potentially modifiable risk factors such as physical inactivity, cognitive impairment, muscle weakness, pain, poor vision, falls and obesity [30]. Gait speed was timed over fixed distance, and reported in m/sec units.
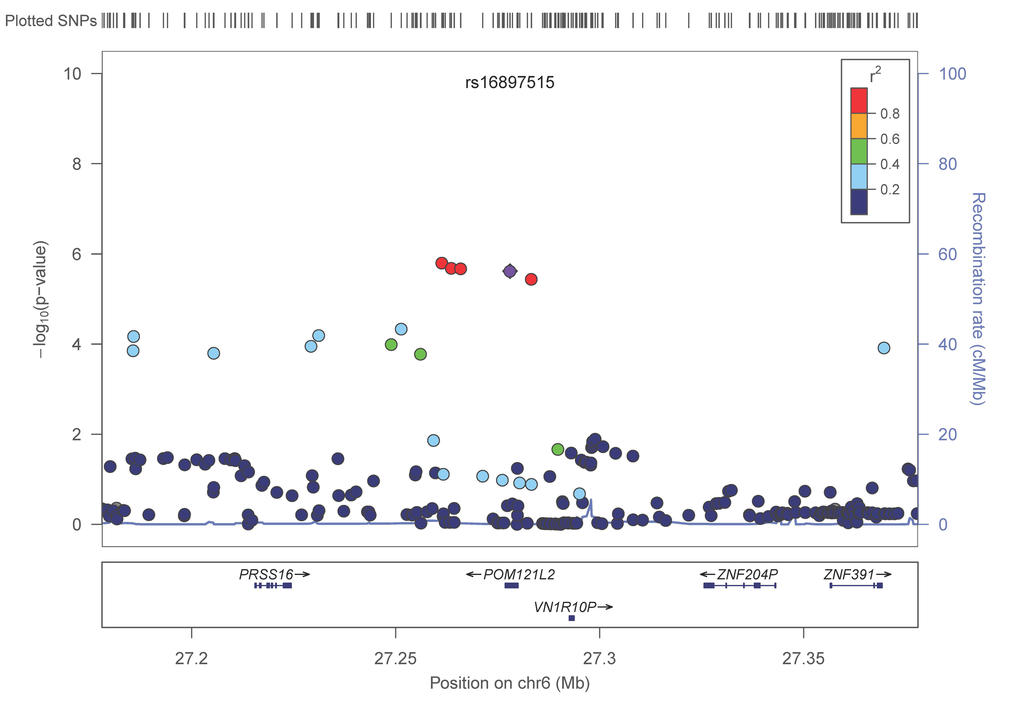
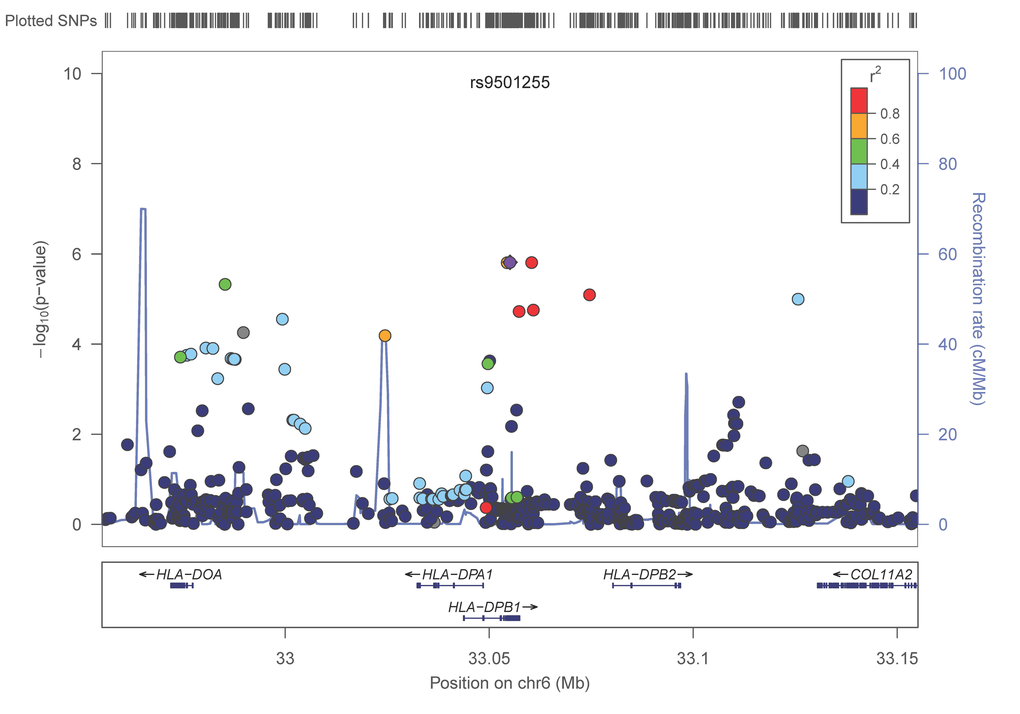
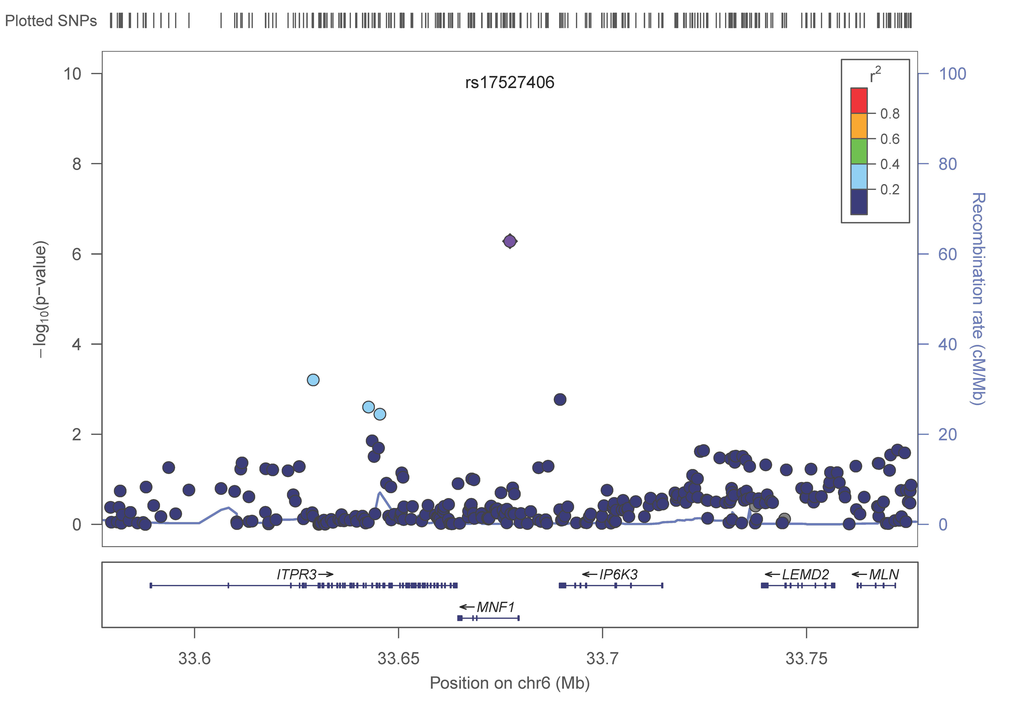
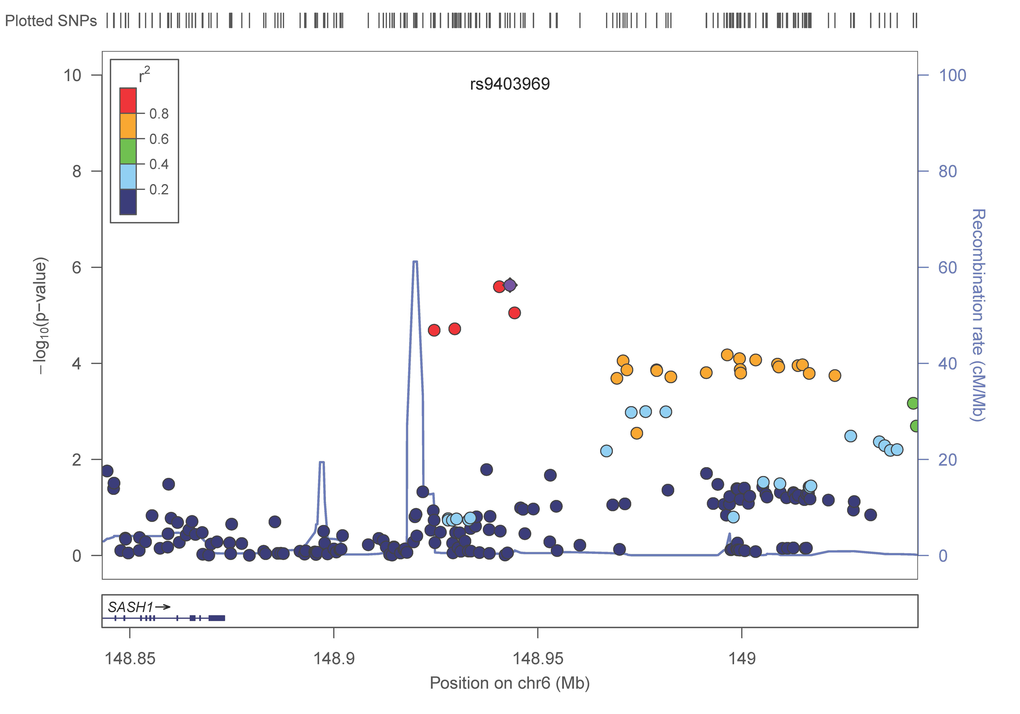
In a meta-analysis of 31,478 subjects from 17 cohorts (Table 1, Supplementary Text) with ~2.5M imputed SNPs (Supplementary Table 1) 536 SNPs (202 were independent (LD, r2 < 0.8) based on the HaploReg tool [31]) with p< 1×10−4 of which 88 (48 were independent signals) had a p-value less than 1×10−5 and one SNP attained a p-value of less than p< 1×10−6 (Table 2, Supplementary Table 2). The Q-Q plot (Supplementary Figure 1) did not provide evidence of inflation of test statistics. The Manhattan plot (fig. 1), highlighted 2 regions on chromosome 6 with high LD and suggestive association with gait speed (Regional plots [32] are displayed in Figure 2). These suggestive regions were further interrogated. Although none of the analyzed SNPs were genome wide significant (p< 5×10−8), one was present in the top ten (POM121L2), and 7 other genes (CEP112, PHACTR1, CNTN5, PTPRT, FHOD3, ADAMTS18, PRIM2) were highlighted based on the presence of SNPs with suggestive significant associations (p<0.0001) as well as low recombination rate and linkage disequilibrium r2 >0.8 which may indicate significant signals in the segment (Figure 2, Supplementary Table 2, Supplementary Figure 2). The 536 suggestive SNPs (p< 1×10−4 in the screening group) were tested for validation in four additional cohorts, GENOA, LLS, MrOSGBG and MrOSMalmo (2588 subjects). Among the top 10 SNPs (six independent) only three exceeded nominal significance which slightly improved the combined meta-analysis significance for HLA-DPB1 SNPs (rs9501255, rs7763822 & rs3749985), however genome-wide levels of significance were not attained (Table 2).
Table 1. Demography of the screening and validation cohorts
| Cohort | Age, y | %Female | N with gait and GWAS | Gait protocol | |
| Screening | AGES | >65 | 58.9 | 3,166 | 6 meter walk |
| ARIC | >60 | 59.5 | 445 | 7.6 meter walk | |
| BLSA | >60 | 49.5 | 334 | 6 meter walk | |
| CHS | ≥65 | 60.9 | 3,184 | 4.6 meter walk | |
| FHS | >65 | 56.1 | 2,384 | 4 meter walk | |
| HABC | >70 | 47.1 | 1,482 | 6 meter walk | |
| HRS | >65 | 56.4 | 5,073 | 2.5 meter walk | |
| InCHIANTI | > 60 | 55.8 | 898 | 4 meter walk | |
| LBC1921 | 77-80 | 58.4 | 510 | 6 meter walk | |
| LBC1936 | 67-71 | 49.5 | 1,001 | 6 meter walk | |
| MrOS | ≥65 | None | 4,643 | 6 meter walk | |
| ROSMAP | >60 | 69.2 | 1,646 | 2.5 meter walk | |
| RS-I | >55 | 53 | 706 | 6 meter walk | |
| RS-II | >55 | 51.8 | 813 | 6 meter walk | |
| RS-III | >45 | 56.0 | 1,392 | 6 meter walk | |
| SOF | ≥65 | 100 | 3,441 | 6 meter walk | |
| TASCOG | >60 | 42 | 360 | 6 meter walk | |
| Total Screening | 31,478 | ||||
| Validation | GENOA | >60 | 55 | 471 | 7.6 meter walk |
| LLS | >60 | 47.2 | 235 | 4 meter walk | |
| MrOSGBG | >69 | None | 960 | 6 meter walk | |
| MrOSMalmo | >69 | None | 922 | 6 meter walk | |
| Total Validation | 2,588 | ||||
Table 2. Top 10 association meta-analysis results for gait speed
| Screening Set (n=31,478) | Validation Set (n=2,588) | Screening + Validation Set (n=34,066) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SNP | Chr.:Position | E/NE Allele | F E Allele | Closest Gene | Δ(kb)/ gene location | Beta (SE) | P | HetPVal | Beta (SE) | P | Beta (SE) | P | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| rs17527406 | 6:33709545 | C/G | 0.016 | UQCC2 MNF1 | intron | 0.040(0.007) | 5.22E-7 | 0.2669 | 0.014(0.032) | 0.65 | 0.037(0.007) | 6.883e-7 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs9501255* | 6:33087321 | T/C | 0.038 | HLA-DPB1 | 3’ UTR | 0.023(0.005) | 1.53e-6 | 0.5853 | 0.048(0.023) | 0.04 | 0.024(0.005) | 3.326e-7 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs7763822* | 6:33092651 | T/C | 0.038 | HLA-DPB1 | 3 | 0.023(0.005) | 1.54e-6 | 0.5704 | 0.047(0.023) | 0.04 | 0.024(0.005) | 3.440e-7 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs3749985* | 6:33086656 | C/G | 0.038 | HLA-DPB1 | 3’ UTR | 0.023(0.005) | 1.55e-6 | 0.5856 | 0.048(0.023) | 0.04 | 0.024(0.005) | 3.385e-7 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs7746199# | 6:27293545 | C/T | 0.166 | POM121L2 | 15 | 0.011(0.002) | 1.58E-6 | 0.9658 | 0.011(0.008) | 0.19 | 0.011(0.002) | 7.125e-7 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs12155739 | 8:102084750 | C/T | 0.030 | NCALD | intron | -0.041(0.008) | 2.04E-6 | 0.2076 | -0.024(0.032) | 0.45 | -0.039(0.008) | 1.858e-6 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs3800318# | 6:27295862 | A/T | 0.830 | POM121L2 | 13 | -0.011(0.002) | 2.07E-6 | 0.966 | -0.011(0.008) | 0.20 | -0.011(0.002) | 9.686e-7 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs13211166 | 6:27298161 | A/T | 0.190 | POM121L2 | 11 | 0.011(0.002) | 2.12E-6 | 0.9688 | 0.010(0.009) | 0.24 | 0.010(0.002) | 1.136e-6 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs9403969 | 6:148622038 | T/G | 0.737 | SASH1 | 70 | 0.009(0.002) | 2.34e-6 | 0.4004 | 0.005(0.007) | 0.50 | 0.009(0.002) | 2.351e-6 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs16897515# | 6:27310241 | A/C | 0.161 | POM121L2 | missense | 0.011(0.002) | 2.41E-6 | 0.9505 | 0.007(0.009) | 0.42 | 0.010(0.002) | 2.080e-6 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| *First gene segment, #second gene segment. E/NE-Effect-, Non-Effect allele; F E-Frequency of Effect Allele; Δ-distance to proximal gene; HetPVal- Heterogeneity P Value. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
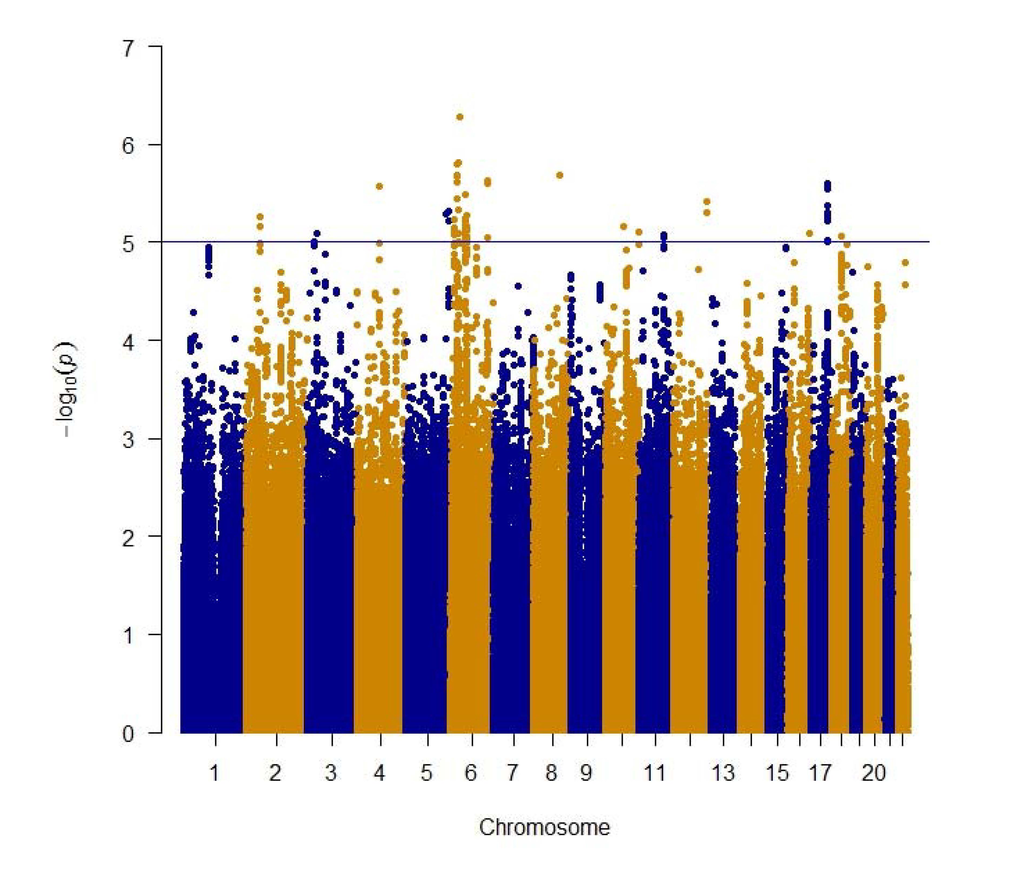
Figure 1. Manhattan plot of meta-analysis of genome wide association studies of gait speed for ~2.5 million genotype and imputed SNPs. The blue line indicates the threshold used to select the 536 suggestive genome wide significant SNPs.
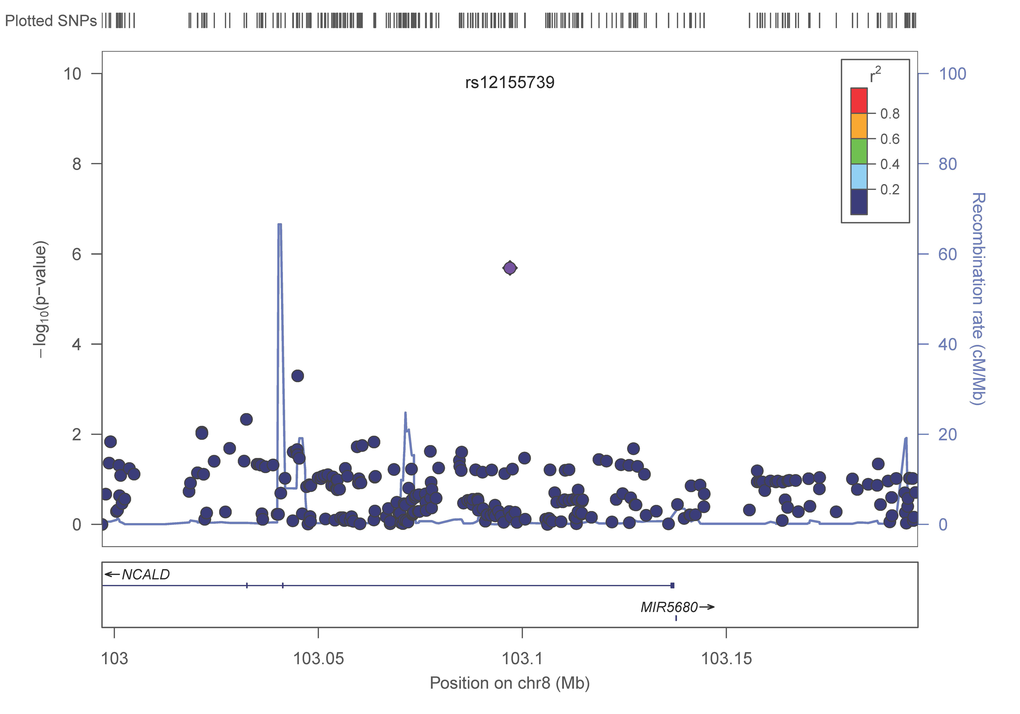
Figure 2E. LocusZoom plots for the suggested top 10 SNPs (5 genes) associated with gait speed of the combined analysis. (A) POM121L2; (B) HLA-DPB1, (C) UQCC2 (MNF1), (D) SASH1, (E) NCALD. In each plot, the −log10 of p values are on the left y-axis; the SNP genomic position (HG19) on the x-axis; the estimated recombination rate from 1000 genomes Nov. 2014 EUR are on the right y-axis and plotted in blue. The most significant SNP is in purple diamond and plotted using the p value attained from the meta-analysis. SNPs are colored to reflect linkage disequilibrium (LD) with the most significant SNP in red (pairwise r2 from 1000 genomes Nov. 2014 EUR). Gene annotations are from the SeattleSeqAnnotation141.
Candidate gene approach
None of the imputed variants previously reported as gait speed candidate genes such as ACE, ACTN3, COMT and APOE reached a nominally significant (p<0.05) threshold (Supplementary Table 3).
Pathway analysis
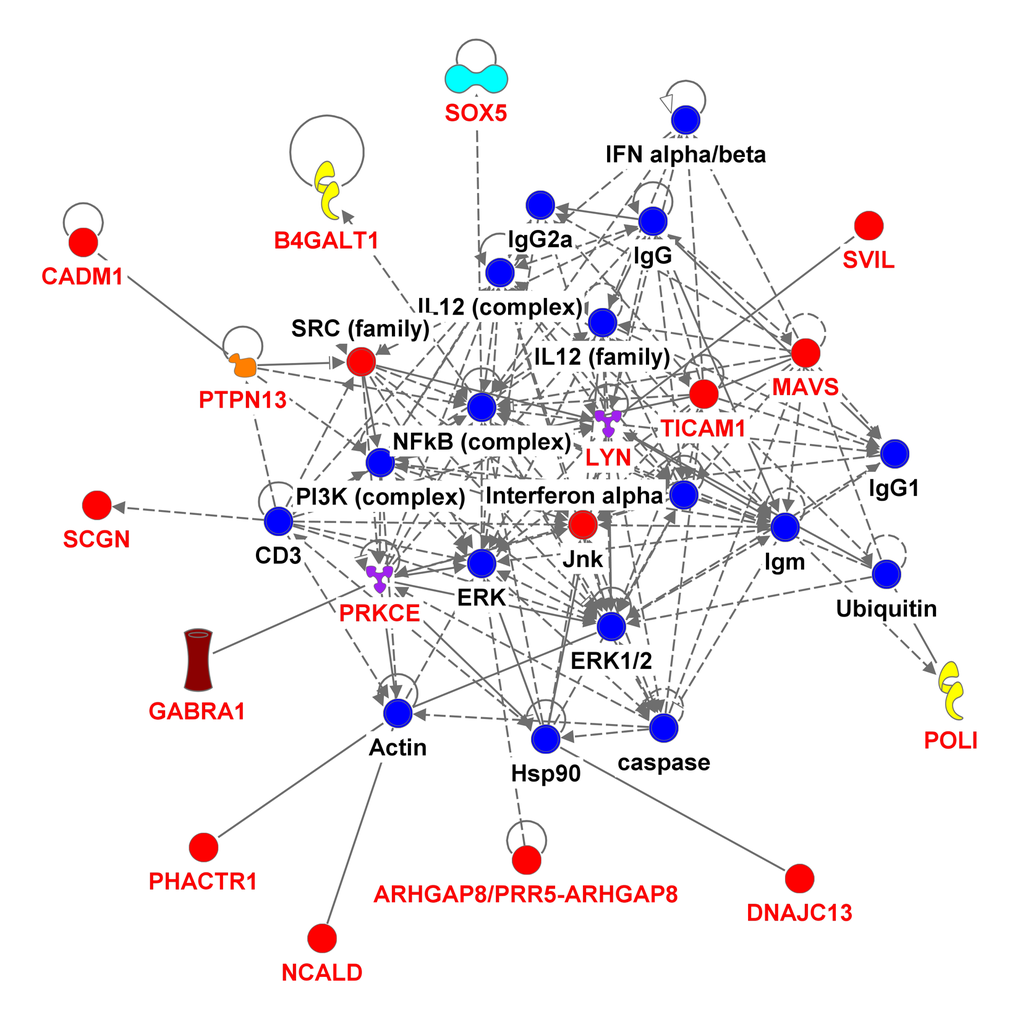
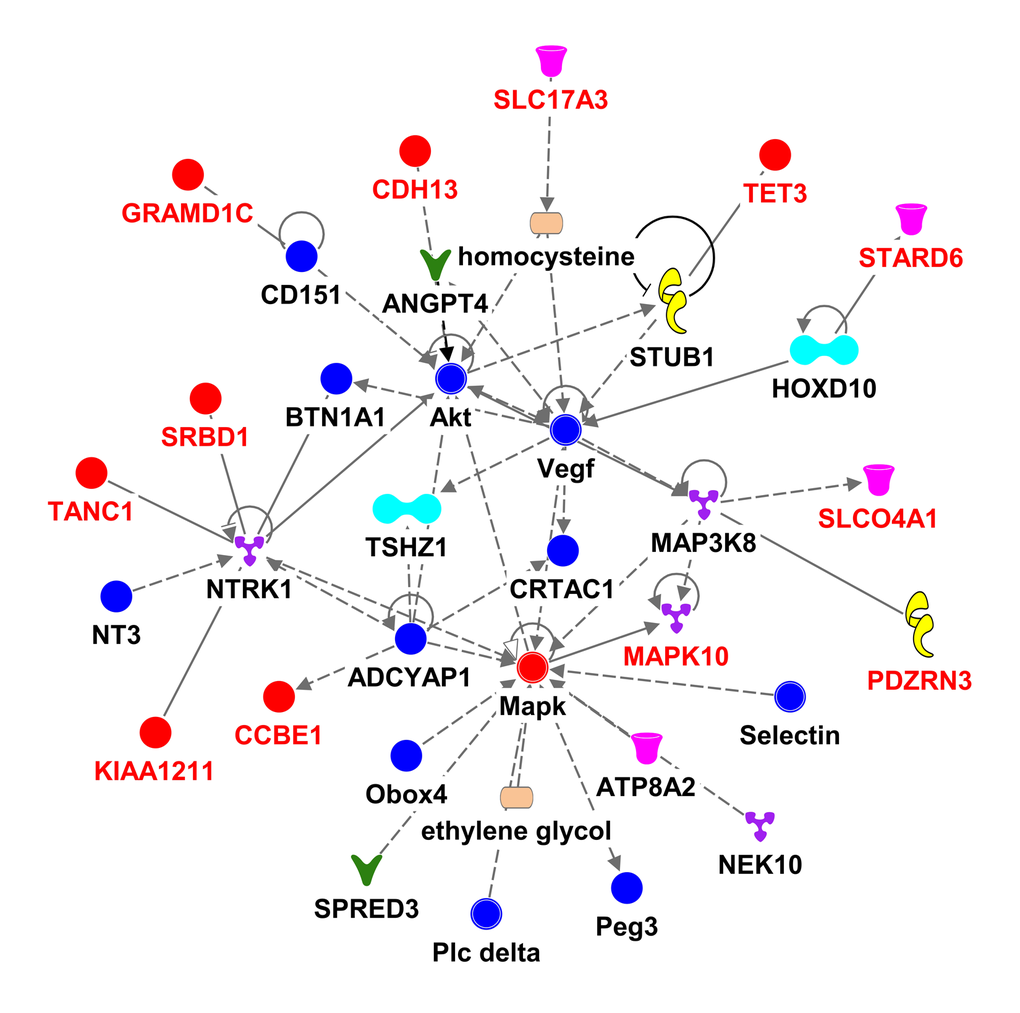
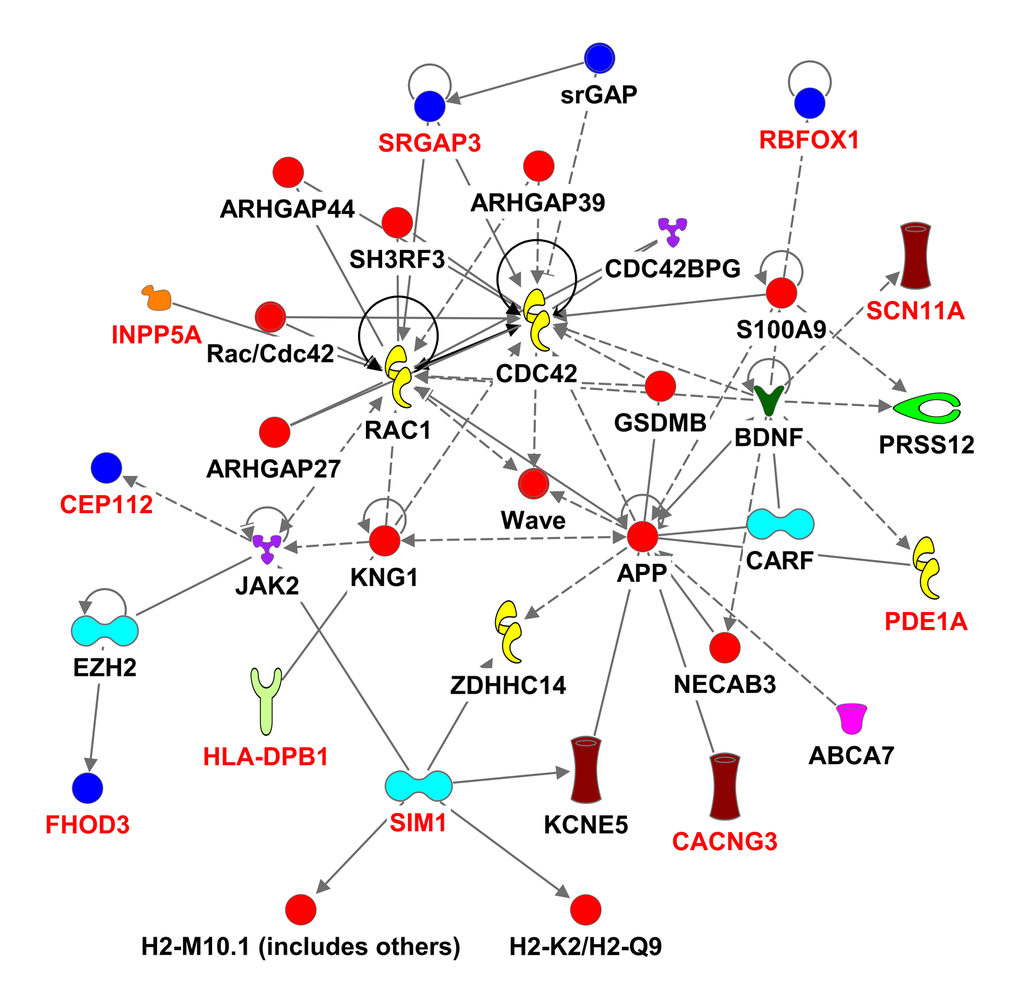
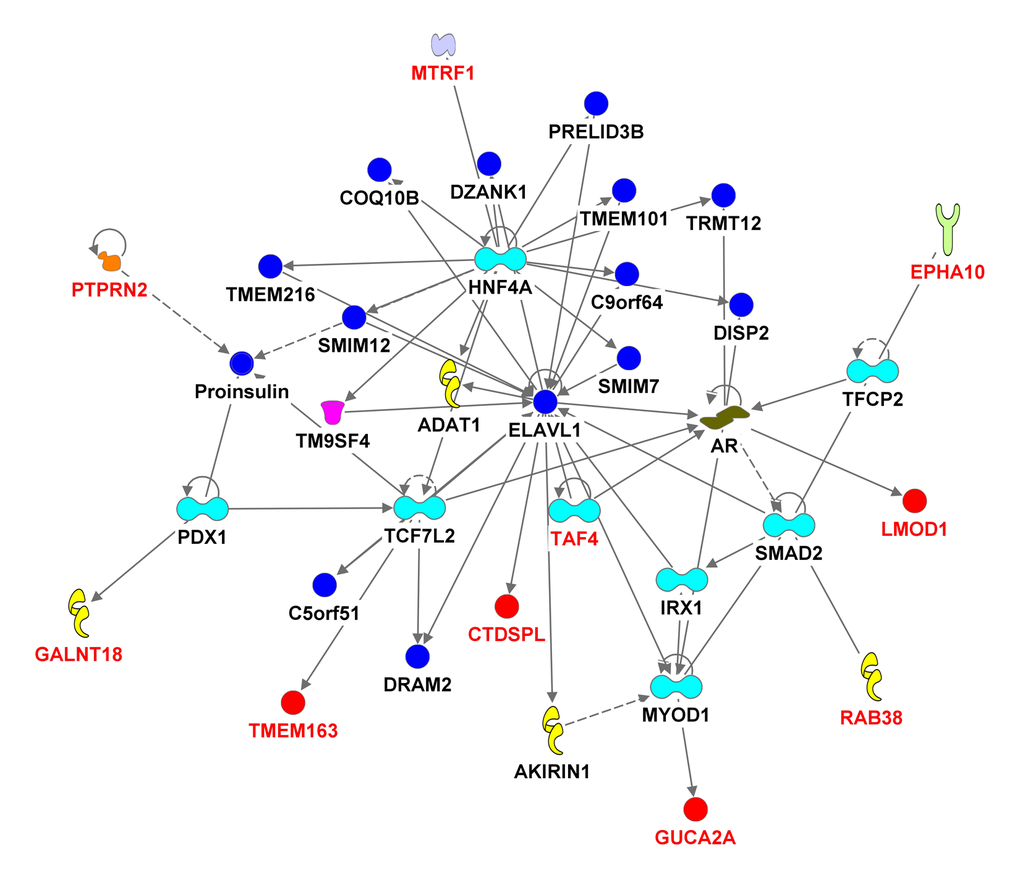
We used the 536 suggestive SNPs to generate the network analysis, in which 283 SNPs representing 68 genes (Supplementary Table 4) were located in both the IPA dataset and the SeattleSeqAnnotation141 for SNP annotation (the remaining 253SNPs did not map to a gene). Among the genes having the highest number of defining SNPs, were CEP112 (38 SNPs), PHACTR1 (23 SNPs), CNTN5 (19 SNPs), PTPRT (18 SNPs), FHOD3 (17 SNPs), ADAMTS18 (12 SNPs) and PRIM2 (11 SNPs). The vast majority of these genes’ products are located in the cytoplasm and plasma membrane while the rest are in the nucleus, extracellular space and other cellular spaces. Ten types of protein actions (enzyme, transporter, phosphatase, transcription regulator, kinase, ion channel, transmembrane receptor, translation regulator, ligand-dependent nuclear receptor and peptidase) are enumerated in Supplementary Table 5. Five of them serve as a biomarker for diagnosis, disease progression, prognosis, and unspecified application and five of them were targets for drug development including PRIM2, GABRA1, LYN, PRKCE and SCN11A. Five major putative disease and function networks were established using the candidate genes (based on the IPA software analysis significance classification) and were classified accordingly to cancer, gastrointestinal disease, organismal injury and abnormalities, neurological disease, cell and tissue morphology, cellular function, development and maintenance, amino acid metabolism, small molecule biochemistry, gene expression, cell-to-cell signaling and interaction, nervous system development and function, cellular assembly and organization. Seven genes were not mapped to any network (see Figure 3, Supplementary Table 6).
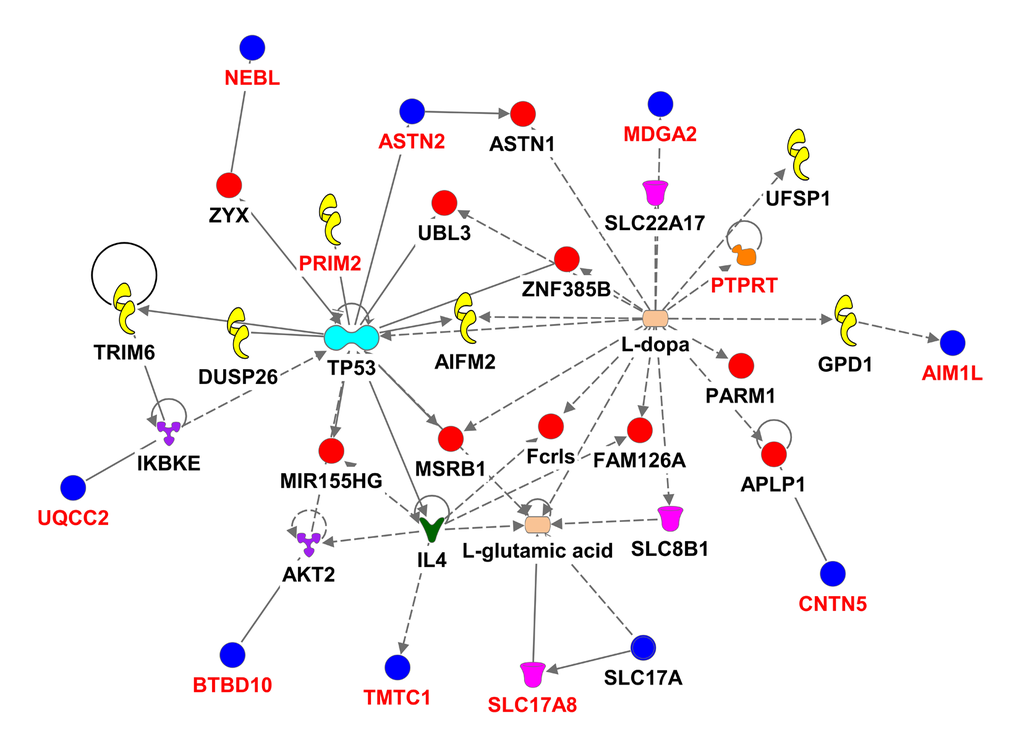
Figure 3E. Ingenuity pathway analysis of genes associated with gait speed. Genes are represented as nodes; solid lines indicate direct- and hatched lines indirect- interaction. Gene functions are color-coded as follows: Red= other, Navy Blue =Group/Complex, Yellow= Enzyme, Turquoise= transcription regulator, Brown= Ion Channel, Orange= Phosphatase, Purple = Kinase, Magenta= Transporter, Beige=chemical-endogenous mammalian, Hunter Green (Dark Green) = Growth factor, light Green= Transmembrane Receptor, Light Purple= Translation Regulator, Olive Green=Ligand-dependent nuclear receptor, Bright green= Peptidase.
eQTL analysis
By querying a large collection of eQTL results (listed in Supplementary Text), we obtained a long list of possible SNP relationships with gene expression (Supplementary Table 7). We also identified the strongest eQTL SNP for each particular transcript in each study. Those SNPs with low p-values (for association with gene expression, p<10-8) and high LD (D'>0.9) with the functional variant, were picked as candidates of signal concordance between the eQTL signals and gait speed signal. Following this analysis, several transcripts including PRSS16 and WDSUB1 were highlighted (Supplementary Table 7). We also observed a relationship between a SNP and PTPRT expression (in liver tissue), which in addition to the meta-analysis and pathway analysis emphasized its potential functional link through its synaptic function and neuronal development, both of which may contribute to [33] gait speed. By emphasizing a strong relationship of the best eQTL with our queried SNPs, we likely underreport SNP-expression relationships due to missing LD information and the inability to project LD relationships for trans-eQTLs in the region.
Applying HaploReg v4.1 analysis to the 536 variants resulted in 9 categories (Supplementary Table 8): miscRNA (1 variant); snoRNA (2 variants); microRNA (4 variants); snRNA (9 variants); pseudogenes (14 variants); sequencing in progress (43 variants); LINC RNA (86 variants); and 372 variants within protein coding genes. In addition, some variants annotate to the same gene resulting in a total of 139 genes (protein-coding or non-coding). Of those genes, 6 are exceptionally long, containing over a million base-pairs, the longest of which is PTPRT coded by 1117219bp. The shortest genes are the ones coding for micro (MIR3143) or small nuclear (U7) RNAs at 63bp each. There is only partial information regarding the chromatin state of each variant. However, from the information gathered in the analysis we observed 14 transcription start sites and 245 enhancers (Supplementary Table 8).
Discussion
In this genome-wide association study of gait speed in 31,478 adults ages 60 and older from 17 different cohorts in the USA, Europe and Australia and 2,588 individuals in four validation cohorts, we did not discover any genome-wide significant association with gait speed nor did we confirm gait speed associations with previously reported candidate genes (i.e. ACE, ACTN3, COMT and APOE) (Supplementary Table 3). However, our analyses revealed some potentially relevant SNPs that could be targeted for further analyses regarding their associations with gait speed.
Our results shed light on several candidate genetic polymorphisms that did not achieve genome wide significance but which had multiple signals on the gene segment, an observation that supported the association with the trait of interest. In addition, these SNPs map to genes that were either linked to physiologic functions expected to influence gait speed (such as neuromuscular function, cardiac function and muscle health or brain function) ADAMTS18, a gene associated with bone mineral density, could be associated with gait speed if individuals with variants in this gene had suffered from fracture leading to slowing of gait [34]. In functional studies ADAMTS18 levels were significantly lower in subjects with non-healing skeletal fractures compared to normal subjects [35]. POM121L2 - an ion transport gene [36] - was listed in the top ten meta-analysis genes with four variants, making it a potential candidate for our study. This gene has been linked to schizophrenia [37], suggesting a potential brain-related association with gait speed. One of the top candidates in our analysis was UQCC2 (also known as M19 or MNF1), a mitochondrial membrane protein that regulates skeletal muscle differentiation and insulin secretion [38]. Although UQCC2 function has a clear link to gait speed, the fact that in this study only one SNP found within UQCC2 demonstrated suggestive significance, which provides less confidence of a true association. NCALD, a calcium-binding protein, has been associated with diabetic nephropathy [39]. The region that was highlighted next to SASH1, a tumor suppressor gene, has multiple signals associated with gait speed. However, there is a high recombination rate between this region and the candidate gene (figure 2), suggesting a higher dissociation between the gene and the signaled region. The last candidate from the top 10 SNP association list is HLA-DPB1, an immune response gene that has been linked to rheumatoid and inflammatory myopathies [40,41]. Interestingly, one of its variants (rs7763822) was indicated in systemic sclerosis susceptibility in Korean subjects [42] suggesting a pleiotropic effect.
CEP112 involved in proper cell cycle progression [43] was not listed among the top 10 SNPs (Table 2) however its clear dominancy (38 SNPs) among the 536 suggestive SNPs make it an attractive candidate for further functional association studies with gait speed. Similar to CEP112 variants, PHACTR1 regulates cardiac α-actin isoform ratio [44] and actomyosin assembly [45]; CNTN5 is associated with neuron function [46]; PTPRT regulates synaptic function and neuronal development [33] and serves as a genuine susceptibility locus for rheumatoid arthritis [33]; FHOD3, is a key regulator in the cardiac muscle [47] and sarcomere organization in striated muscle cells [48]; and PRIM2 is involved in DNA replication and transcription and is crucial for normal growth and development [49]. This list of genes repeatedly implicates associated signals that are important for neuromuscular function, cardiac function and muscle health, which could reasonably contribute to the complex trait of gait speed.
A second tier of locus with repetitive signals established among the 536 suggestive SNPs included PDZN3, which is implicated in muscle function and regeneration [50–52], CACNG3, a voltage-dependent calcium channel subunit [53] that was previously linked to ataxic phenotype in mice [54], ASTN2 that functions in neuronal migration [55] and that was associated with hip osteoarthritis susceptibility [56], SIM1 involved in coordinating muscle activity and generating rhythmic activity [57] and also associated with obesity [58], and MDGA2, which is required for proper development of cranial motoneuron subtypes [59].
The eQTL analysis (various tissues and cell types, listed in Supplementary Text) of the 536 suggestive SNPs reported a couple of candidate genes such as PRSS16, a gene encoding serine protease expressed exclusively in the thymus. PRSS16 was associated with exercise [60] and was linked to COMT (a candidate gene for gait speed (20)). Both are regulated by ZNF804a [61]. This link between the two genes (PRSS16 and COMT) may support our gait speed association results. Another candidate gene from our eQTL analysis was WDSUB1 a U-box ubiquitin ligases encoded protein which was associated with sudden cardiac death [62]. A link with cardiovascular diseases may indicate a potential cardiovascular effect on gait speed. The last candidate in this analysis is PTPRT, a gene that regulates synaptic function and neuronal development. It is possible that its link to gait speed (operates through its role in diabetes [63]). The fact that it was present in all three sets of analysis results may suggest a stronger candidate for further analysis.
The lead motif of the network analysis in all 5 disease networks was “cellular function”, however, the candidate SNPs from the multiple analysis strategies strongly suggested links to bone, skeleton, muscle and brain, incorporating development, structure and function. While our SNP associations did not achieve genome wide significance, we believe that we demonstrated a potential link to gait speed. To exclude false positive signals, these associations should be pursued further in controlled experiments as well as animal models, which will increase our understanding of the biology of gait speed deterioration with aging. Such efforts would provide us with an increased knowledge of the biological systems which impact on gait speed; this may contribute to improved treatment strategies and drug development to promote aging with grace.
This study did not provide conclusive evidence for the genetics contributing to gait speed. While the large sample is a strength (and we have the power to detect smaller effects), the observed associations suggest that an even larger sample is required to establish genetic contributions to the gait speed phenotype. The individual effects of common SNPs for complex traits such as gait speed are expected to be very small. From studies of other polygenic complex traits, it has been observed that the number of discovered variants is strongly correlated with experimental sample size [64]. Another potential explanation why we did not observe genome wide significant associations is that there are many potential pathways that contribute to gait speed, including nervous system function (neuromuscular, central nervous system), musculoskeletal conditions such as sarcopenia and osteoarthritis, cardiovascular disease, visual function, psychological factors and other contributors. This complexity of phenotype may make it difficult to discover associations. Phenotype refinement may be a future approach to explore.
In summary, the lack of genome-wide significant signals from this moderately large sample of older adults suggests that larger samples (or study to sub-classify the gait speed phenotype) will be needed to identify SNP-based associations. Also, it may suggest that downstream mechanisms are more likely to make more important contributions to gait speed. Gait speed is a complex phenotype with many potential contributors; it is not unsurprising that it should be governed by multiple genes. However, we were able to use network analyses to define some potential networks of genes that might be of relevance for this phenotype. Future studies may be best positioned to focus on one network in more detail and to examine gene-environment or gene-behavior- environment interactions.
Methods
Subjects
The Aging and Longevity Working Group of the CHARGE Consortium [65,66], was formed to facilitate genome-wide association study meta-analyses of age associated diseases and phenotypes among multiple large and well-phenotyped cohorts of older individuals who underwent genotyping.
Screening cohorts
A combined cohort of 31,478 subjects age 60 years and older with timed walks constituted our discovery sample (Table 1). Timed walk at usual pace was converted to gait speed (m/s) to harmonize the phenotype across cohorts. Participants of the following 17 European descendent cohorts were included (Supplementary Text):
The Age, Gene/Environment Susceptibility-Reykjavik (AGES), The Atherosclerosis Risk in Communities (ARIC), Baltimore Longitudinal study on Aging (BLSA), Cardiovascular Health Study (CHS), Framingham Heart Study (FHS), Health, Aging, and Body Composition Study (HABC), Health and Retirement Study (HRS), Invecchiare in Chianti (InCHIANTI), Lothian Birth Cohorts 1921 (LBC1921) and 1936 (LBC1936), Osteoporotic Fractures in Men Study (MrOS), The Religious Orders Study and Rush Memory and Aging Project (ROSMAP), Rotterdam Study (RS-I, -II, -III), Study of Osteoporotic Fractures (SOF), Tasmanian Study of Cognition and Gait (TASCOG) (Table 1, Supplementary Text). All participants with gait speed assessments including participants who were able to walk with assistance of a cane were included in this analysis. Exclusion criteria included missing gait assessments and inability to walk (Supplement text).
Validation cohorts
The validation cohort consisted of 2,588 subjects (>60 years) from the Genetic Epidemiology Network of Arteriopathy (GENOA), Leiden Longevity Study (LLS), Osteoporotic Fractures in Men Study (MrOS) Sweden, Malmö[MrOSMalmo] and Gothenburg [MrOSGBG] studies (Table 1, Supplementary Text). Together these cohorts reach the minimum number of subjects required for sufficient statistical power (Our power calculation shows that given a fixed sample size (n=2500) our analysis will have >80% power to detect MAF=0.01, alpha<0.0001) to validate significant signal(s) from the screening cohort using the same harmonized gait speed phenotype. Results from the screening and validation cohorts were meta-analyzed.
Phenotype definition
The different methods of assessing gait speed in individual cohorts are described in Table 1 and Supplementary Text. Variability in the methods of assessing gait speed in the participating cohorts included differences in distance walked (8 to 25 feet) and measurement techniques (instrumented walkway versus stopwatch). Previous reports including 4 cohorts from this report (CHS, HABC, InCHIANTI and SOF) have suggested that there is a high correlation (r2>0.9) between the different methods of measuring gait speed [9,24,67]. The mean overall gait speed was 1.13± 0.25 m/sec, and varied from 0.66 ± 0.16 m/sec to 1.66 ± 0.41 m/sec in the individual cohorts (Supplementary Table 9, Supplementary Figure 3).
Genotyping
A structured, pre-specified analytical plan was applied to each of the 17 cohorts included in the screening sample. Genome-wide analysis of imputed genotypes, summarized in Supplementary Text, were conducted in each cohort. Imputation (using either BimBam or MACH) resulted in approximately 2.5 million HapMap SNPs being available for analysis. Imputation details, QC and SNP count per cohort can be found in Supplementary Text and Supplementary Table 1. Exclusion criteria for SNP in each of the 21 cohorts (screening and validation) included: 1) minor allele frequency (MAF) < 0.005); 2) imputation quality (R2 or oevar_imp < 0.3); and for the meta-analysis, SNPs with average MAF ≤ 0.01 and total N < 15,000.
Cohort-specific analyses
Multiple linear regression of imputed SNP dosages on gait speed was performed using an additive model, i.e. as a count of the number of variant alleles present (1 degree of freedom). Sex-combined analysis was performed. Adjustment for age (at time of exam), sex, study site (for cohorts with multiple sites), principal components to control for population stratification, height, and presence of osteoarthritis (yes/no) if available were included. For cohorts with osteoarthritis data available, the analysis was done excluding participants with osteoarthritis (Supplementary Text and Supplementary Table 10).
Meta-analysis
Inverse variance weighted meta-analysis was performed on summary statistics of the cohort-level association analyses. Meta-analysis of gait speed (Screening and validation cohorts were analyzed separately as well as together (joint meta-analysis)) was performed using METAL [68] with a fixed effects model of beta estimates and standard errors from each cohort. In addition, we applied heterogeneity test between studies (on both screening and validation cohorts) using METAL. A p-value threshold (Bonferroni-adjusted) of p<5x10-8 was used to indicate genome-wide statistical significance.
Pathway analysis
We assembled a list of 536 meta-analyzed SNPs (representing 69 genes) that were highly suggestively associated (p < 1 × 10−4) with gait speed. This list resulted in 67 candidate genes (Annotated by Ingenuity Pathway Analysis (IPA) and SeattleSeqAnnotation) being identified which were used in the IPA analysis (www.ingenuity.com). The resulting classification of networks, pathways, biological processes and molecular functions are represented in tables and graphic format (Figure 3, Supplementary Table 4, Supplementary Table 5 and Supplementary Table 6).
Expression quantitative trait loci (eQTL) analysis
We examined existing eQTL resources for the candidate suggestive list of 536 SNPs (p<10-4) to further explore their biological and functional relevance to gait speed (Supplementary Text). We queried these SNPs against an eQTL database (listed in Supplementary Text) containing eQTL results from over 100 studies across a wide range of tissues. A general overview of a subset of >50 eQTL studies has been published [69], with specific citations for the included datasets included in the Supplementary Material.
Further we applied the HaploReg v4.1 annotation tool for TF analysis of 536 SNPs suggestively associated with gait speed.
AGES
This study has been funded by NIH contracts N01-AG-1-2100 and 271201200022C, the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association), and the Althingi (the Icelandic Parliament). The study is approved by the Icelandic National Bioethics Committee, VSN: 00-063. The researchers are indebted to the participants for their willingness to participate in the study.
ARIC
The Atherosclerosis Risk in Communities Study is carried out as a collaborative study supported by National Heart, Lung, and Blood Institute contracts (HHSN268201100005C, HHSN268201100006C, HHSN268201100007C, HHSN268201100008C, HHSN268201100009C, HHSN268201100010C, HHSN268201100011C, and HHSN268201100012C), R01HL087641, R01HL59367 and R01HL086694; National Human Genome Research Institute contract U01HG004402; and National Institutes of Health contract HHSN268200625226C. The authors thank the staff and participants of the ARIC study for their important contributions. Infrastructure was partly supported by Grant Number UL1RR025005, a component of the National Institutes of Health and NIH Roadmap for Medical Research. This project was also partially supported from NIH grant R01-HL093029.
BLSA
The BLSA was supported by the Intramural Research Program of the NIH, National Institute on Aging.
CHS
The CHS research was supported by NHLBI contracts HHSN268201200036C, HHSN268200800007C, HHSN268200960009C, N01HC55222, N01HC85079, N01HC85080, N01HC85081, N01HC85082, N01HC85083, N01HC85086; and NHLBI grants HL080295, HL087652, HL105756 with additional contribution from the National Institute of Neurological Disorders and Stroke (NINDS). Additional support was provided through AG023629 from the National Institute on Aging (NIA). A full list of CHS investigators and institutions can be found at http://chs-nhlbi.org/. The provision of genotyping data was supported in part by the National Center for Advancing Translational Sciences, CTSI grant UL1TR000124, and the National Institute of Diabetes and Digestive and Kidney Disease Diabetes Research Center (DRC) grant DK063491 to the Southern California Diabetes Endocrinology Research Center.
FHS
The Framingham Heart Study phenotype-genotype analyses were supported by NIA R01AG29451 (JMM, KLL). The Framingham Heart Study of the National Heart Lung and Blood Institute of the National Institutes of Health and Boston University School of Medicine was supported by the National Heart, Lung and Blood Institute's Framingham Heart Study Contract No. N01-HC-25195 and its contract with Affymetrix, Inc for genotyping services (Contract No. N02-HL-6-4278). Analyses reflect intellectual input and resource development from the Framingham Heart Study investigators participating in the SNP Health Association Resource (SHARe) project. A portion of this research was conducted using the Linux Cluster for Genetic Analysis (LinGA-II) funded by the Robert Dawson Evans Endowment of the Department of Medicine at Boston University School of Medicine and Boston Medical Center. The investigators thank the participants and the staff for their support of the Framingham Heart Study. Dr. Kiel's effort was supported by NIAMS grant RO1 AR41398.
HABC
This research was supported by NIA contracts N01AG62101, N01AG62103, and N01AG62106. The genome-wide association study was funded by NIA grant 1R01AG032098-01A1 to Wake Forest University Health Sciences and genotyping services were provided by the Center for Inherited Disease Research (CIDR). CIDR is fully funded through a federal contract from the National Institutes of Health to The Johns Hopkins University, contract number HHSN268200782096C. This research was supported in part by the Intramural Research Program of the NIH, National Institute on Aging.
HRS
HRS is supported by the National Institute on Aging (NIA U01AG009740). Genotyping was funded separately by NIA (RC2 AG036495, RC4 AG039029). Our genotyping was conducted by the NIH Center for Inherited Disease Research (CIDR) at Johns Hopkins University. Genotyping quality control and final preparation of the data were performed by the Genetics Coordinating Center at the University of Washington.
InCHIANTI
The InCHIANTI study baseline (1998-2000) was supported as a "targeted project" (ICS110.1/RF97.71) by the Italian Ministry of Health and in part by the U.S. National Institute on Aging (Contracts: 263 MD 9164 and 263 MD 821336).
LBC1921 and LBC1936
We thank the cohort participants and team members who contributed to these studies. Phenotype collection in the Lothian Birth Cohort 1921 was supported by the UK Biotechnology and Biological Sciences Research Council (BBSRC), The Royal Society and The Chief Scientist Office of the Scottish Government. Phenotype collection in the Lothian Birth Cohort 1936 was supported by Research Into Ageing (continues as part of Age UK The Disconnected Mind project). Genotyping of the cohorts was funded by the BBSRC. The work was undertaken by The University of Edinburgh Centre for Cognitive Ageing and Cognitive Epidemiology, part of the cross council Lifelong Health and Wellbeing Initiative (MR/K026992/1). Funding from the BBSRC and Medical Research Council (MRC) is gratefully acknowledged.
MrOS
The Osteoporotic Fractures in Men (MrOS) Study is supported by National Institutes of Health funding. The following institutes provide support: the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS), the National Institute on Aging (NIA), the National Center for Research Resources (NCRR), and NIH Roadmap for Medical Research under the following grant numbers: U01 AR45580, U01 AR45614, U01 AR45632, U01 AR45647, U01 AR45654, U01 AR45583, U01 AG18197, U01-AG027810, and UL1 RR024140. The National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) provides funding for the MrOS ancillary study ‘Replication of candidate gene associations and bone strength phenotype in MrOS’ under the grant number R01-AR051124. The National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) provides funding for the MrOS ancillary study ‘GWAS in MrOS and SOF’ under the grant number RC2ARO58973.
ROSMAP
The data from the Rush Alzheimer’s Disease Center used in these analyses was supported by National Institute on Aging grants P30AG10161, R01AG17917, R01AG15819, R01AG30146, the Illinois Department of Public Health, R01NS078009 and the Translational Genomics Research Institute.
RS-I, -II, -III
The generation and management of GWAS genotype data for the Rotterdam Study are supported by the Netherlands Organisation of Scientific Research NWO Investments (nr. 175.010.2005.011, 911-03-012). This study is funded by the Research Institute for Diseases in the Elderly (014-93-015; RIDE2), the Netherlands Genomics Initiative (NGI)/Netherlands Organisation for Scientific Research (NWO) project nr. 050-060-810. The Rotterdam Study is funded by Erasmus Medical Center and Erasmus University, Rotterdam, Netherlands Organization for the Health Research and Development (ZonMw), the Research Institute for Diseases in the Elderly (RIDE), the Ministry of Education, Culture and Science, the Ministry for Health, Welfare and Sports, the European Commission (DG XII), and the Municipality of Rotterdam. This research is supported by the Dutch Technology Foundation STW, which is part of the NWO, and which is partly funded by the Ministry of Economic Affairs. MAI is supported by ZonMW grant number 916.13.054.
SOF
The Study of Osteoporotic Fractures (SOF) is supported by National Institutes of Health funding. The National Institute on Aging (NIA) provides support under the following grant numbers: R01 AG005407, R01 AR35582, R01 AR35583, R01 AR35584, R01 AG005394, R01 AG027574, and R01 AG027576.
TASCOG
All participants and research staff who took part in TASCOG are duly acknowledged, in addition to funding bodies including the National Health and Medical Research Council (NHMRC) of Australia, and the National Institutes of Health (NIH) USA.
GENOA
Support for the Genetic Epidemiology Network of Arteriopathy (GENOA) was provided by the National Heart, Lung and Blood Institute (HL054457, HL087660, HL119443) and the National Institute of Neurological Disorders and Stroke (NS041558) of the National Institutes of Health. Genotyping was performed at the Mayo Clinic (S.T.T, Mariza de Andrade, Julie Cunningham) and was made possible by the University of Texas Health Sciences Center (Eric Boerwinkle, Megan Grove-Gaona). We would also like to thank the families that participated in the GENOA study.
LLS
The research leading to these results has received funding from the European Union’s Seventh Framework Programme (FP7/2007-2011) under grant agreement number 259679. This study was financially supported by the Innovation-Oriented Research Program on Genomics (SenterNovem IGE05007), the Centre for Medical Systems Biology and the Netherlands Consortium for Healthy Ageing (grant 050-060-810), all in the framework of the Netherlands Genomics Initiative, Netherlands Organization for Scientific Research (NWO), by Unilever Colworth and by BBMRI-NL, a Research Infrastructure financed by the Dutch government (NWO 184.021.007).
MrOS Sweden
This study was supported by the Swedish Research Council and by grants from the Swedish government (under the Avtal om Läkarutbildning och Medicinsk Forskning [Agreement for Medical Education and Research]), the Lundberg Foundation, the Torsten Söderberg Foundation, the Novo Nordisk Foundation and the Knut and Alice Wallenberg Foundation.
Supplementary Materials
Conflicts of Interest
The authors have no conflict of interests to declare.
Funding
Acknowledgments and funding for the participating studies are listed below.
References
- 1. Fritz S, Lusardi M. White paper: “walking speed: the sixth vital sign”. J Geriatr Phys Ther. 2009; 32:46–49. https://doi.org/10.1519/00139143-200932020-00002 [PubMed]
- 2. Bohannon RW, Williams Andrews A. Normal walking speed: a descriptive meta-analysis. Physiotherapy. 2011; 97:182–89. https://doi.org/10.1016/j.physio.2010.12.004 [PubMed]
- 3. Bollard E, Fleming H. A study to investigate the walking speed of elderly adults with relation to pedestrian crossings. Physiother Theory Pract. 2013; 29:142–49. https://doi.org/10.3109/09593985.2012.703760 [PubMed]
- 4. Dommes A, Cavallo V, Dubuisson JB, Tournier I, Vienne F. Crossing a two-way street: comparison of young and old pedestrians. J Safety Res. 2014; 50:27–34. https://doi.org/10.1016/j.jsr.2014.03.008 [PubMed]
- 5. Bijleveld-Uitman M, van de Port I, Kwakkel G. Is gait speed or walking distance a better predictor for community walking after stroke? J Rehabil Med. 2013; 45:535–40. https://doi.org/10.2340/16501977-1147 [PubMed]
- 6. Elbers RG, van Wegen EE, Verhoef J, Kwakkel G. Is gait speed a valid measure to predict community ambulation in patients with Parkinson’s disease? J Rehabil Med. 2013; 45:370–75. https://doi.org/10.2340/16501977-1123 [PubMed]
- 7. Perry SB, Woollard J, Little S, Shroyer K. Relationships among measures of balance, gait, and community integration in people with brain injury. J Head Trauma Rehabil. 2014; 29:117–24. https://doi.org/10.1097/HTR.0b013e3182864f2f [PubMed]
- 8. Kuh D, Karunananthan S, Bergman H, Cooper R. A life-course approach to healthy ageing: maintaining physical capability. Proc Nutr Soc. 2014; 73:237–48. https://doi.org/10.1017/S0029665113003923 [PubMed]
- 9. Studenski S, Perera S, Patel K, Rosano C, Faulkner K, Inzitari M, Brach J, Chandler J, Cawthon P, Connor EB, Nevitt M, Visser M, Kritchevsky S, et al. Gait speed and survival in older adults. JAMA. 2011; 305:50–58. https://doi.org/10.1001/jama.2010.1923 [PubMed]
- 10. Abellan van Kan G, Rolland Y, Andrieu S, Bauer J, Beauchet O, Bonnefoy M, Cesari M, Donini LM, Gillette Guyonnet S, Inzitari M, Nourhashemi F, Onder G, Ritz P, et al. Gait speed at usual pace as a predictor of adverse outcomes in community-dwelling older people an International Academy on Nutrition and Aging (IANA) Task Force. J Nutr Health Aging. 2009; 13:881–89. https://doi.org/10.1007/s12603-009-0246-z [PubMed]
- 11. Ortega-Alonso A, Pedersen NL, Kujala UM, Sipilä S, Törmäkangas T, Kaprio J, Koskenvuo M, Rantanen T. A twin study on the heritability of walking ability among older women. J Gerontol A Biol Sci Med Sci. 2006; 61:1082–85. https://doi.org/10.1093/gerona/61.10.1082 [PubMed]
- 12. Pajala S, Era P, Koskenvuo M, Kaprio J, Alén M, Tolvanen A, Tiainen K, Rantanen T. Contribution of genetic and environmental factors to individual differences in maximal walking speed with and without second task in older women. J Gerontol A Biol Sci Med Sci. 2005; 60:1299–303. https://doi.org/10.1093/gerona/60.10.1299 [PubMed]
- 13. Christensen K, McGue M, Yashin A, Iachine I, Holm NV, Vaupel JW. Genetic and environmental influences on functional abilities in Danish twins aged 75 years and older. J Gerontol A Biol Sci Med Sci. 2000; 55:M446–52. https://doi.org/10.1093/gerona/55.8.M446 [PubMed]
- 14. Carmelli D, Kelly-Hayes M, Wolf PA, Swan GE, Jack LM, Reed T, Guralnik JM. The contribution of genetic influences to measures of lower-extremity function in older male twins. J Gerontol A Biol Sci Med Sci. 2000; 55:B49–53. https://doi.org/10.1093/gerona/55.1.B49 [PubMed]
- 15. Tiainen KM, Perola M, Kovanen VM, Sipilä S, Tuononen KA, Rikalainen K, Kauppinen MA, Widen EI, Kaprio J, Rantanen T, Kujala UM. Genetics of maximal walking speed and skeletal muscle characteristics in older women. Twin Res Hum Genet. 2008; 11:321–34. https://doi.org/10.1375/twin.11.3.321 [PubMed]
- 16. Ayers E, Barzilai N, Crandall JP, Milman S, Verghese J. Association of exceptional parental longevity and physical function in aging. Age (Dordr). 2014; 36:9677. https://doi.org/10.1007/s11357-014-9677-5 [PubMed]
- 17. Newman AB, Glynn NW, Taylor CA, Sebastiani P, Perls TT, Mayeux R, Christensen K, Zmuda JM, Barral S, Lee JH, Simonsick EM, Walston JD, Yashin AI, Hadley E. Health and function of participants in the Long Life Family Study: A comparison with other cohorts. Aging (Albany NY). 2011; 3:63–76. https://doi.org/10.18632/aging.100242 [PubMed]
- 18. Eynon N, Hanson ED, Lucia A, Houweling PJ, Garton F, North KN, Bishop DJ. Genes for elite power and sprint performance: ACTN3 leads the way. Sports Med. 2013; 43:803–17. https://doi.org/10.1007/s40279-013-0059-4 [PubMed]
- 19. Lima RM, Leite TK, Pereira RW, Rabelo HT, Roth SM, Oliveira RJ. ACE and ACTN3 genotypes in older women: muscular phenotypes. Int J Sports Med. 2011; 32:66–72. https://doi.org/10.1055/s-0030-1267229 [PubMed]
- 20. Pereira A, Costa AM, Leitão JC, Monteiro AM, Izquierdo M, Silva AJ, Bastos E, Marques MC. The influence of ACE ID and ACTN3 R577X polymorphisms on lower-extremity function in older women in response to high-speed power training. BMC Geriatr. 2013; 13:131. https://doi.org/10.1186/1471-2318-13-131 [PubMed]
- 21. Holtzer R, Ozelius L, Xue X, Wang T, Lipton RB, Verghese J. Differential effects of COMT on gait and executive control in aging. Neurobiol Aging. 2010; 31:523–31. https://doi.org/10.1016/j.neurobiolaging.2008.05.011 [PubMed]
- 22. Kulminski A, Ukraintseva SV, Arbeev KG, Manton KG, Oshima J, Martin GM, Yashin AI. Association between APOE epsilon 2/epsilon 3/epsilon 4 polymorphism and disability severity in a national long-term care survey sample. Age Ageing. 2008; 37:288–93. https://doi.org/10.1093/ageing/afn003 [PubMed]
- 23. Soerensen M, Dato S, Tan Q, Thinggaard M, Kleindorp R, Beekman M, Suchiman HE, Jacobsen R, McGue M, Stevnsner T, Bohr VA, de Craen AJ, Westendorp RG, et al. Evidence from case-control and longitudinal studies supports associations of genetic variation in APOE, CETP, and IL6 with human longevity. Age (Dordr). 2013; 35:487–500. https://doi.org/10.1007/s11357-011-9373-7 [PubMed]
- 24. Verghese J, Holtzer R, Wang C, Katz MJ, Barzilai N, Lipton RB. Role of APOE genotype in gait decline and disability in aging. J Gerontol A Biol Sci Med Sci. 2013; 68:1395–401. https://doi.org/10.1093/gerona/glt115 [PubMed]
- 25. Witte JS. Genome-wide association studies and beyond. Annu Rev Public Health. 2010; 31:9–20, 4, 20. https://doi.org/10.1146/annurev.publhealth.012809.103723 [PubMed]
- 26. Adams HH, Verlinden VJ, Callisaya ML, van Duijn CM, Hofman A, Thomson R, Uitterlinden AG, Vernooij MW, van der Geest JN, Srikanth V, Ikram MA. Heritability and Genome-Wide Association Analyses of Human Gait Suggest Contribution of Common Variants. J Gerontol A Biol Sci Med Sci. 2015. [PubMed]
- 27. Verghese J, Holtzer R, Lipton RB, Wang C. Quantitative gait markers and incident fall risk in older adults. J Gerontol A Biol Sci Med Sci. 2009; 64:896–901. https://doi.org/10.1093/gerona/glp033 [PubMed]
- 28. Cummings SR, Studenski S, Ferrucci L. A diagnosis of dismobility--giving mobility clinical visibility: a Mobility Working Group recommendation. JAMA. 2014; 311:2061–62. https://doi.org/10.1001/jama.2014.3033 [PubMed]
- 29. Verghese J, Wang C, Lipton RB, Holtzer R, Xue X. Quantitative gait dysfunction and risk of cognitive decline and dementia. J Neurol Neurosurg Psychiatry. 2007; 78:929–35. https://doi.org/10.1136/jnnp.2006.106914 [PubMed]
- 30. Verghese J, Wang C, Allali G, Holtzer R, Ayers E. Modifiable Risk Factors for New-Onset Slow Gait in Older Adults. J Am Med Dir Assoc. 2016; 17:421–25. https://doi.org/10.1016/j.jamda.2016.01.017 [PubMed]
- 31. Ward LD, Kellis M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 2012; 40:D930–34. https://doi.org/10.1093/nar/gkr917 [PubMed]
- 32. Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP, Boehnke M, Abecasis GR, Willer CJ. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics. 2010; 26:2336–37. https://doi.org/10.1093/bioinformatics/btq419 [PubMed]
- 33. Julià A, Ballina J, Cañete JD, Balsa A, Tornero-Molina J, Naranjo A, Alperi-López M, Erra A, Pascual-Salcedo D, Barceló P, Camps J, Marsal S. Genome-wide association study of rheumatoid arthritis in the Spanish population: KLF12 as a risk locus for rheumatoid arthritis susceptibility. Arthritis Rheum. 2008; 58:2275–86. https://doi.org/10.1002/art.23623 [PubMed]
- 34. Xiong DH, Liu XG, Guo YF, Tan LJ, Wang L, Sha BY, Tang ZH, Pan F, Yang TL, Chen XD, Lei SF, Yerges LM, Zhu XZ, et al. Genome-wide association and follow-up replication studies identified ADAMTS18 and TGFBR3 as bone mass candidate genes in different ethnic groups. Am J Hum Genet. 2009; 84:388–98. https://doi.org/10.1016/j.ajhg.2009.01.025 [PubMed]
- 35. Wei J, Liu CJ, Li Z. ADAMTS-18: a metalloproteinase with multiple functions. Front Biosci (Landmark Ed). 2014; 19:1456–67. https://doi.org/10.2741/4296 [PubMed]
- 36. Shang Y, Das S, Rabold R, Sham JS, Mitzner W, Tang WY. Epigenetic alterations by DNA methylation in house dust mite-induced airway hyperresponsiveness. Am J Respir Cell Mol Biol. 2013; 49:279–87. https://doi.org/10.1165/rcmb.2012-0403OC [PubMed]
- 37. Aberg KA, Liu Y, Bukszár J, McClay JL, Khachane AN, Andreassen OA, Blackwood D, Corvin A, Djurovic S, Gurling H, Ophoff R, Pato CN, Pato MT, et al. A comprehensive family-based replication study of schizophrenia genes. JAMA Psychiatry. 2013; 70:573–81. https://doi.org/10.1001/jamapsychiatry.2013.288 [PubMed]
- 38. Cambier L, Rassam P, Chabi B, Mezghenna K, Gross R, Eveno E, Auffray C, Wrutniak-Cabello C, Lajoix AD, Pomiès P. M19 modulates skeletal muscle differentiation and insulin secretion in pancreatic β-cells through modulation of respiratory chain activity. PLoS One. 2012; 7:e31815. https://doi.org/10.1371/journal.pone.0031815 [PubMed]
- 39. Kamiyama M, Kobayashi M, Araki S, Iida A, Tsunoda T, Kawai K, Imanishi M, Nomura M, Babazono T, Iwamoto Y, Kashiwagi A, Kaku K, Kawamori R, et al. Polymorphisms in the 3′ UTR in the neurocalcin delta gene affect mRNA stability, and confer susceptibility to diabetic nephropathy. Hum Genet. 2007; 122:397–407. https://doi.org/10.1007/s00439-007-0414-3 [PubMed]
- 40. van Steenbergen HW, Raychaudhuri S, Rodríguez-Rodríguez L, Rantapää-Dahlqvist S, Berglin E, Toes RE, Huizinga TW, Fernández-Gutiérrez B, Gregersen PK, van der Helm-van Mil AH. Association of valine and leucine at HLA-DRB1 position 11 with radiographic progression in rheumatoid arthritis, independent of the shared epitope alleles but not independent of anti-citrullinated protein antibodies. Arthritis Rheumatol. 2015; 67:877–86. https://doi.org/10.1002/art.39018 [PubMed]
- 41. Chinoy H, Lamb JA, Ollier WE, Cooper RG. An update on the immunogenetics of idiopathic inflammatory myopathies: major histocompatibility complex and beyond. Curr Opin Rheumatol. 2009; 21:588–93. https://doi.org/10.1097/BOR.0b013e3283315a22 [PubMed]
- 42. Zhou X, Lee JE, Arnett FC, Xiong M, Park MY, Yoo YK, Shin ES, Reveille JD, Mayes MD, Kim JH, Song R, Choi JY, Park JA, et al. HLA-DPB1 and DPB2 are genetic loci for systemic sclerosis: a genome-wide association study in Koreans with replication in North Americans. Arthritis Rheum. 2009; 60:3807–14. https://doi.org/10.1002/art.24982 [PubMed]
- 43. Kumar A, Rajendran V, Sethumadhavan R, Purohit R. CEP proteins: the knights of centrosome dynasty. Protoplasma. 2013; 250:965–83. https://doi.org/10.1007/s00709-013-0488-9 [PubMed]
- 44. Kelloniemi A, Szabo Z, Serpi R, Näpänkangas J, Ohukainen P, Tenhunen O, Kaikkonen L, Koivisto E, Bagyura Z, Kerkelä R, Leosdottir M, Hedner T, Melander O, et al. The Early-Onset Myocardial Infarction Associated PHACTR1 Gene Regulates Skeletal and Cardiac Alpha-Actin Gene Expression. PLoS One. 2015; 10:e0130502. https://doi.org/10.1371/journal.pone.0130502 [PubMed]
- 45. Wiezlak M, Diring J, Abella J, Mouilleron S, Way M, McDonald NQ, Treisman R. G-actin regulates the shuttling and PP1 binding of the RPEL protein Phactr1 to control actomyosin assembly. J Cell Sci. 2012; 125:5860–72. https://doi.org/10.1242/jcs.112078 [PubMed]
- 46. Ashrafi S, Betley JN, Comer JD, Brenner-Morton S, Bar V, Shimoda Y, Watanabe K, Peles E, Jessell TM, Kaltschmidt JA. Neuronal Ig/Caspr recognition promotes the formation of axoaxonic synapses in mouse spinal cord. Neuron. 2014; 81:120–29. https://doi.org/10.1016/j.neuron.2013.10.060 [PubMed]
- 47. Kan-O M, Takeya R, Abe T, Kitajima N, Nishida M, Tominaga R, Kurose H, Sumimoto H. Mammalian formin Fhod3 plays an essential role in cardiogenesis by organizing myofibrillogenesis. Biol Open. 2012; 1:889–96. https://doi.org/10.1242/bio.20121370 [PubMed]
- 48. Taniguchi K, Takeya R, Suetsugu S, Kan-O M, Narusawa M, Shiose A, Tominaga R, Sumimoto H. Mammalian formin fhod3 regulates actin assembly and sarcomere organization in striated muscles. J Biol Chem. 2009; 284:29873–81. https://doi.org/10.1074/jbc.M109.059303 [PubMed]
- 49. Chung J, Tsai S, James AH, Thames BH, Shytle S, Piedrahita JA. Lack of genomic imprinting of DNA primase, polypeptide 2 (PRIM2) in human term placenta and white blood cells. Epigenetics. 2012; 7:429–31. https://doi.org/10.4161/epi.19777 [PubMed]
- 50. Ko JA, Kimura Y, Matsuura K, Yamamoto H, Gondo T, Inui M. PDZRN3 (LNX3, SEMCAP3) is required for the differentiation of C2C12 myoblasts into myotubes. J Cell Sci. 2006; 119:5106–13. https://doi.org/10.1242/jcs.03290 [PubMed]
- 51. Lu Z, Je HS, Young P, Gross J, Lu B, Feng G. Regulation of synaptic growth and maturation by a synapse-associated E3 ubiquitin ligase at the neuromuscular junction. J Cell Biol. 2007; 177:1077–89. https://doi.org/10.1083/jcb.200610060 [PubMed]
- 52. Sewduth RN, Jaspard-Vinassa B, Peghaire C, Guillabert A, Franzl N, Larrieu-Lahargue F, Moreau C, Fruttiger M, Dufourcq P, Couffinhal T, Duplàa C. The ubiquitin ligase PDZRN3 is required for vascular morphogenesis through Wnt/planar cell polarity signalling. Nat Commun. 2014; 5:4832. https://doi.org/10.1038/ncomms5832 [PubMed]
- 53. Burgess DL, Gefrides LA, Foreman PJ, Noebels JL. A cluster of three novel Ca2+ channel gamma subunit genes on chromosome 19q13.4: evolution and expression profile of the gamma subunit gene family. Genomics. 2001; 71:339–50. https://doi.org/10.1006/geno.2000.6440 [PubMed]
- 54. Sharp AH, Black JL
3rd , Dubel SJ, Sundarraj S, Shen JP, Yunker AM, Copeland TD, McEnery MW. Biochemical and anatomical evidence for specialized voltage-dependent calcium channel gamma isoform expression in the epileptic and ataxic mouse, stargazer. Neuroscience. 2001; 105:599–617. https://doi.org/10.1016/S0306-4522(01)00220-2 [PubMed] - 55. Lionel AC, Tammimies K, Vaags AK, Rosenfeld JA, Ahn JW, Merico D, Noor A, Runke CK, Pillalamarri VK, Carter MT, Gazzellone MJ, Thiruvahindrapuram B, Fagerberg C, et al. Disruption of the ASTN2/TRIM32 locus at 9q33.1 is a risk factor in males for autism spectrum disorders, ADHD and other neurodevelopmental phenotypes. Hum Mol Genet. 2014; 23:2752–68. https://doi.org/10.1093/hmg/ddt669 [PubMed]
- 56. Lindner C, Thiagarajah S, Wilkinson JM, Panoutsopoulou K, Day-Williams AG, Cootes TF, Wallis GA, and arcOGEN Consortium. Investigation of association between hip osteoarthritis susceptibility loci and radiographic proximal femur shape. Arthritis Rheumatol. 2015; 67:2076–84. https://doi.org/10.1002/art.39186 [PubMed]
- 57. Kiehn O, Dougherty KJ, Hägglund M, Borgius L, Talpalar A, Restrepo CE. Probing spinal circuits controlling walking in mammals. Biochem Biophys Res Commun. 2010; 396:11–18. https://doi.org/10.1016/j.bbrc.2010.02.107 [PubMed]
- 58. Swarbrick MM, Evans DS, Valle MI, Favre H, Wu SH, Njajou OT, Li R, Zmuda JM, Miljkovic I, Harris TB, Kwok PY, Vaisse C, Hsueh WC. Replication and extension of association between common genetic variants in SIM1 and human adiposity. Obesity (Silver Spring). 2011; 19:2394–403. https://doi.org/10.1038/oby.2011.79 [PubMed]
- 59. Ingold E, Vom Berg-Maurer CM, Burckhardt CJ, Lehnherr A, Rieder P, Keller PJ, Stelzer EH, Greber UF, Neuhauss SC, Gesemann M. Proper migration and axon outgrowth of zebrafish cranial motoneuron subpopulations require the cell adhesion molecule MDGA2A. Biol Open. 2015; 4:146–54. https://doi.org/10.1242/bio.20148482 [PubMed]
- 60. De Moor MH, Liu YJ, Boomsma DI, Li J, Hamilton JJ, Hottenga JJ, Levy S, Liu XG, Pei YF, Posthuma D, Recker RR, Sullivan PF, Wang L, et al. Genome-wide association study of exercise behavior in Dutch and American adults. Med Sci Sports Exerc. 2009; 41:1887–95. https://doi.org/10.1249/MSS.0b013e3181a2f646 [PubMed]
- 61. Girgenti MJ, LoTurco JJ, Maher BJ. ZNF804a regulates expression of the schizophrenia-associated genes PRSS16, COMT, PDE4B, and DRD2. PLoS One. 2012; 7:e32404. https://doi.org/10.1371/journal.pone.0032404 [PubMed]
- 62. Arking DE, Junttila MJ, Goyette P, Huertas-Vazquez A, Eijgelsheim M, Blom MT, Newton-Cheh C, Reinier K, Teodorescu C, Uy-Evanado A, Carter-Monroe N, Kaikkonen KS, Kortelainen ML, et al. Identification of a sudden cardiac death susceptibility locus at 2q24.2 through genome-wide association in European ancestry individuals. PLoS Genet. 2011; 7:e1002158. https://doi.org/10.1371/journal.pgen.1002158 [PubMed]
- 63. Thomas J, Garg ML, Smith DW. Altered expression of histone and synaptic plasticity associated genes in the hippocampus of streptozotocin-induced diabetic mice. Metab Brain Dis. 2013; 28:613–18. https://doi.org/10.1007/s11011-013-9418-y [PubMed]
- 64. Lango Allen H, Estrada K, Lettre G, Berndt SI, Weedon MN, Rivadeneira F, Willer CJ, Jackson AU, Vedantam S, Raychaudhuri S, Ferreira T, Wood AR, Weyant RJ, et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature. 2010; 467:832–38. https://doi.org/10.1038/nature09410 [PubMed]
- 65. Newman AB, Walter S, Lunetta KL, Garcia M, Karasik D, Rivadeneira F, Tiemeier H, Uitterlinden AG, Walston J, Westendorp R, Harris TB, Lumley T, van Duijn CM, et al. A Meta-analysis of Four Genome-wide Association Studies of Survival to Age 90 Years or Older: The Cohorts for Heart and Aging Research in Genome Epidemiology (CHARGE) Consortium. J Gerontol A Biol Sci Med Sci. Epub ahead of print.
- 66. Walter S, Atzmon G, Demerath E, Garcia ME, Lunetta KL, Christensen K, Arnold AM, Aspelund T, Aulchenko YS, Benjamin EJ, Christiansen L, D’Agostino RB
Sr , Fitzpatrick AL, et al. Neurobiol Aging. 2011. - 67. Guralnik JM, Ferrucci L, Pieper CF, Leveille SG, Markides KS, Ostir GV, Studenski S, Berkman LF, Wallace RB. Lower extremity function and subsequent disability: consistency across studies, predictive models, and value of gait speed alone compared with the short physical performance battery. J Gerontol A Biol Sci Med Sci. 2000; 55:M221–31. https://doi.org/10.1093/gerona/55.4.M221 [PubMed]
- 68. Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010; 26:2190–91. https://doi.org/10.1093/bioinformatics/btq340 [PubMed]
- 69. Zhang X, Gierman HJ, Levy D, Plump A, Dobrin R, Goring HH, Curran JE, Johnson MP, Blangero J, Kim SK, O’Donnell CJ, Emilsson V, Johnson AD. Synthesis of 53 tissue and cell line expression QTL datasets reveals master eQTLs. BMC Genomics. 2014; 15:532. https://doi.org/10.1186/1471-2164-15-532 [PubMed]