Introduction
One of the most dramatic modifications associated with human aging is the progressive decline in skeletal muscle mass and function, known as sarcopenia. Sarcopenia causes physical function impairment, leading to a loss of functional independence and to an increased incidence of adverse health outcomes [1,2]. The prevalence of this condition, which is progressively increasing due to the prolonged life expectancy, has led in the recent decades to increased needs for health care services and resources to support older people [3]. The rate of skeletal muscle loss is estimated at 8% per decade from the 4th until 7th decade, with about 15% lost each decade after 70 years of age and loss of strength is estimated to be even longer [4].
The cause of sarcopenia is widely regarded as multifactorial, with chronic diseases, hormonal and inflammatory changes, mitochondrial dysfunction, physical inactivity, and malnutrition, being the main risk factors [5]. In addition to these causes, loss of muscle has been related to the imbalance between muscle protein synthesis (MPS) and muscle protein breakdown (MPB), associated with alterations in muscle anabolic responses to nutritional stimuli and physical activity, the so-called “anabolic resistance” according to which in the elderly process the “anabolic threshold” required to maximize anabolic pathways is increased [6,7].
Amino acids (AAs) availability is among the most important anabolic signals for MPS [8], and essential amino acids (EAAs), leucine in particular, has been shown to be critical in the regulation of skeletal muscle protein synthesis and degradation in the elderly [9]. There is growing evidence that AA transport into muscle cells may be a rate-limiting step in the process of AA induced stimulation of skeletal muscle protein metabolism, and that AA transporters may have a key role in such process as AA sensors [10]. This is likely linked to the ability of AA transporters to act as both transporter and receptor (transceptor) [11,12] able to transduce a signal reflecting AA availability and leading to activation of the mammalian/mechanistic target of rapamycin complex 1 (mTORC1), the major nutrient-sensitive signalling pathway [13]. Thus, AA transporters may be an important link in the ability for AAs to stimulate MPS. This has been supported by studies in human skeletal muscle showing that the expression of AA transporters is highly dynamic and responsive to different anabolic stimuli [14]. In particular, in 2010, Drummond and colleagues [15] characterized the expression of selected AA transporters in the human skeletal muscle following AA ingestion in a group of healthy young individuals. They found increased mRNA expression of L-type AA transporter SLC7A5 (LAT1), SLC3A2 (CD98), sodium-coupled neutral AA transporter SLC38A2 (SNAT2), and proton-coupled amino acid transporter SLC36A1 (PAT1), which are transporters thought to have key roles in mTORC1 signalling regulation and muscle protein synthesis and muscle growth. These authors argue that, changes in the expression levels of AA transporters, possibly mediated by mTORC1 activity, could serve as an adaptive response for improving AA intracellular delivery and for regulating the rate of muscle protein synthesis in response to anabolic stimuli or during periods of decreased muscle protein synthesis [15]. Consistently with this hypothesis, the same authors showed that an up-regulation of the above transporters occurred after resistance exercise in both young and older adults, likely regulated in an age-dependent manner [16], as well as in older persons in conditions of short-term bed rest [17]. Soon after that, Dickinson and co-workers [14] reported that aging differentially affected the expression of SLC7A5 and SLC38A2 in the skeletal muscle when EAAs were ingested after exercise, suggesting that aging may influence the function of specific AA transporters, and possibly age-related phenotypes, such as sarcopenia [14,18].
Based on these evidences, to shed a light on the impact of the above AA transporters on the age-related loss of muscle strength and physical performance, we screened genetic variants occurring in their genes in a population of elderly subjects (< 90 years), analysing them in combination with parameters of physical status. Moreover, beside these four genes [SLC7A5 (LAT1), SLC3A2 (CD98), SLC38A2 (SNAT2) and SLC36A1 (PAT1)], we also included variants located in SLC7A8 (LAT2) and SLC43A1 (LAT3) as leucine transporters like LAT1, SLC1A5 (ASCT2) as a glutamine transporter like SNAT2, SLC38A3 (SNAT3) and SLC38A7 (SNAT7), preferentially transporting glutamine and arginine, SLC38A9 (SNAT9), as an arginine transporter [19,20].
To investigate whether variants in the selected genes also affect the chance to survive and/or reach very advanced ages, we exploited both the cross-sectional and longitudinal approach by taking advantage of the available dataset, which also included very old subjects (≥90 years), and survival data within 10-year follow-up period from the baseline visit.
Results
Demographic, clinical, and anthropometric characteristics of the analysed sample are presented in Table 1. Among the selected SNPs, 10 did not pass the QC phase. In particular, four SNPs were excluded due to a MAF (Minor Allele Frequency) lower than ten percent per locus (rs17112008, rs7968173, rs1175, rs7735053), while six were excluded because they showed a significant deviation from HWE in control subjects (rs11749532, rs7736177, rs2897968, rs17794251, rs7193392, rs8058969).
Table 1. Socio-demographic characteristics and functional parameters in the sample stratified for group membership.
| 50-89 (N = 475) | 90-108 (N = 290) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age (year) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mean (SD) | 70.40 ± 8.71 | 96.86 ± 3.86 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Male (%) | 50% | 37.24% | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Height | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mean (SD) | 161.54 ± 9.28 | 151.39 ± 9.52 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Range | 138-190 | 125-175 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BMI (SD) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mean (SD) | 27.24 ± 4.19 | 23.12 ± 4.03 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Range | 17.80-45.35 | 12.98-40.54 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HG strength | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mean (SD) | 22.86 (10.0) | 13.09 (6.22) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Range | 4-55 | 1-36 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ADL [n (%)] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Non disable (=5) | 84.4% | 31.9% | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Disable (<5) | 15.6% | 68.1% | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Walking 4 meters [sec] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mean (SD) | 7.77 (4.35) | 12.3 (7.26) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Range | 2.65-50.00 | 3.65-43.0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviations: BMI: Body mass index; ADL, Activity Daily Living; HG, Hand Grip Strength. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Association with longevity
To investigate whether variants in genes encoding AA transporters also concur to determine the chance to reach very advanced age we applied a cross-sectional study, by including 271 more subjects aged 90 years and older.
Complete results of case-control analysis are reported in Figure S1 (D), while Table 3 shows the statistically significant associations. Among the SNPs showing association with at least one muscle-related phenotype, in accordance with the negative effect of the minor C allele of rs1873793 (SLC38A2) on ADL performance, subjects carrying this allele were also significantly less frequent in the older population compared to the younger one (OR=0.70, CI 0.5-0.95; p=0.035). In addition to this, we found that the minor allele of rs1858828 in SLC38A3 (OR=1.33 (1.06-1.68), pAdd=0.01) was positively correlated with longevity. A lower chance to reach very advanced ages was found for the minor allele of rs12794763 in SLC3A2 (OR=0.42, 0.26-0.68; pD=0.0002), and for the minor alleles of three SNPs in SLC38A9: two of them, rs4865615 and rs7704138, are in LD with each other (r2>0.8) and showed respectively OR=0.54 [(0.33-0.88), pRec=0.011] and OR=0.56 [(0.34-0.92), pRec=0.018], while rs10056358 showed a OR=0.66 [(0.45-0.96), pDom=0.029)].
Table 3. SNPs showing significant associations with longevity under a nominal level (p Model < 0.05).
| Gene | SNP | MAF | OR (CI) | pModel | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SLC3A2 CD98 | rs12794763 | G=0.15 | 0.42 (0.26-0.68) | 0.0002D | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SLC38A2 SNAT2 | rs1873793 | C=0.48 | 0.70 (0.50-0.98) | 0.035D | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SLC38A3 SNAT3 | rs1858828 | T=0.38 | 1.33 (1.06-1.68) | 0.014 A | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SLC38A9 SNAT9 | rs4865615 | C=0.4 | 0.54 (0.33-0.88) | 0.011R | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs7704138 | C=0.39 | 0.56 (0.34-0.92) | 0.018R | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| rs10056358 | A=0.14 | 0.66 (0.45-0.96) | 0.029 D | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviations: MAF, Minor allele frequency; OR, odds ratio; CI, confidence interval. p Model is the p value of the best genetic model, where R is recessive, D is dominant, and A is additive model. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The variable sex was considered a covariate. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Association with survival
Next, using 10-year of follow-up survival data, we investigated the association with survival of the analysed SNPs, estimating their quantitative effects. An increased risk of death was found for subjects carrying the C allele at rs14160 (SLC36A1) compared to those with the TT genotype (adjusted HR = 1.59, 95% CI: 1.025–2.47; p=0.038) (Figure 1a). In SLC38A9, the presence of the A allele for the rs10056358 variation confers a survival advantage before 90 years of age, compared to those with the TT genotype (adjusted HR=0.48, 95% CI 0.27-0.84, p=0.009) (Figure 1b), and a disadvantageous effect later, (adjusted HR = 1.43, 95% CI 1.01-2.03; p= 0.045) (Figure 1c).
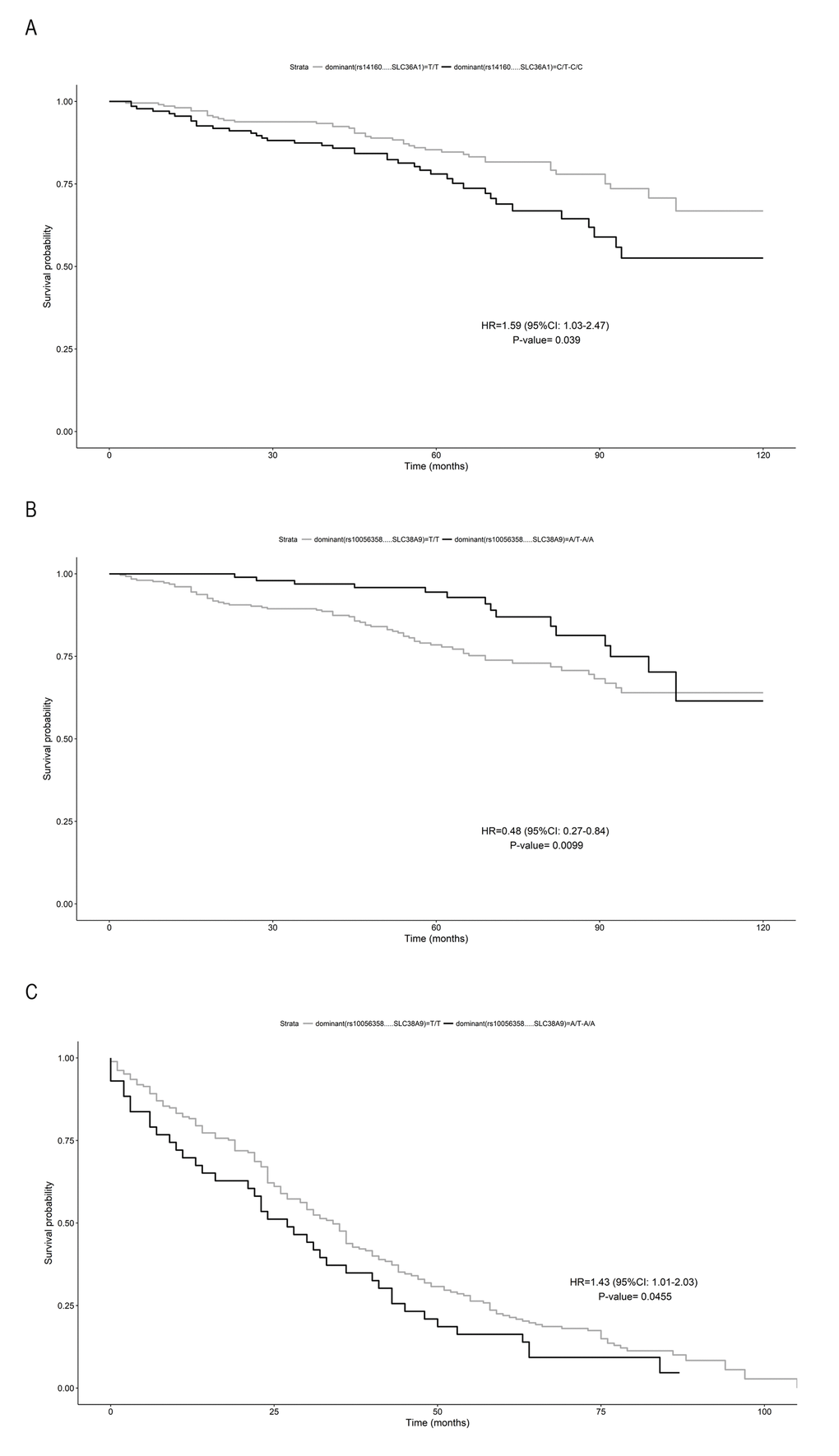
Figure 1. Survival function of carriers of minor allele (black) vs non carriers (grey). (A) rs14160 (SLC36A1) in subjects aged <90 years; (B) rs10056358 (SLC38A9) in subjects aged ≥90 years; (C) rs10056358 (SLC38A9) in subjects aged > 90 years. Time is expressed in months, where 0 is considered the time of recruitment, and each individual is followed up for survival status till death. HR value, confidence interval and p-value from Cox regression analysis are reported inside the figure.
Analysis of all other SNPs showed no statistically significant or suggestive differences in survival between genotypes.
Imputation of functional impact of SNPs
Finally, to explore the functional consequences of the associated polymorphisms we used regulatory information from ENCODE data using HaploReg v4.1 and RegulomeDB databases. Results from the functional annotation analysis are shown in Table S2. By using HaploReg v4.1 database, we found promising functional implications for these SNPs (i.e. promoter and enhancer histone marks, transcription factor binding sites, and eQTL hits), indicating a regulatory potential. The functional implication of some SNPs was further corroborated by RegulomeDB results showing for rs357629 (SLC36A1) a "likely to affect binding and linked to expression of a gene target (score 1f)" and for rs4329925 and rs731710 (SLC7A5), and for rs1873793 (SLC38A2) a "likely to affect binding (score 2b)". LD patterns showed that all SNPs, except two (rs12794763, SLC3A2 and rs999165, SLC7A8), were in LD with several variants that can collectively capture the casual variant.
Discussion
The goal of the current study was to investigate the impact of SNPs in selected AA transporters genes on physical performance and survival at old age.
Our analysis of a group of elderly individuals in the age ranges of 50-89 years, showed that genetic variants in SLC7A5 (rs4329925) and SLC7A8 (rs999165 and rs12588118), negatively affected HG strength, a reliable marker of the muscle performance and an indicator of quality of aging [21]. Additionally, different SNPs in the same genes (rs731710 in SLC7A5 and rs3783436 in SLC7A8) were significantly associated with ADL levels. SLC7A5 and SLC7A8 code, respectively, for the plasma membrane proteins LAT1 and LAT2. Each of them forms with CD98 (SLC3A2) a heterodimeric bidirectional antiporter that regulates the simultaneous transport of leucine into cells and efflux of glutamine out of cells. Intriguingly, we found that two variants in SLC3A2 (rs12804553 and rs4726) were associated, with opposite effects, with walking ability. These findings point to a role of the LAT1(LAT2)/CD98 complex in affecting muscle strength and physical performance in the elderly, likely through regulation of the anabolic signal mediated by the intracellular leucine/glutamine levels, and downstream activation of mTORC1. On the other hand, it is well known that leucine provides direct anabolic stimuli to skeletal muscle, so that the post-exercise leucine assumption is considered a potential treatment in numerous muscle wasting conditions [18]. Similarly, glutamine status is considered a hallmark of catabolic states and muscle loss [22].
Further supportive evidence to the above hypothesis is provided by the association with genes that mediate the transport of glutamine, such as SLC38A2 (SNAT2) and SLC38A7 (SNAT7). We found, indeed, significantly worse ADL scores associated to rs1873793 (SLC38A2), a finding supported by a disadvantage in attaining longevity for subjects carrying the same allelic variant, and a better walking performance associated to rs9806843 (SLC38A7). For the above transporters an important role in mTORC1 activation was documented. For instance, a reduction of SNAT2 activity in muscle cells is associated with reduced levels of glutamine and leucine and impaired protein synthesis through mTORC1 [23]; furthermore, it was found that the expression of system SNAT2, LAT1 and CD98 was upregulated after leucine availability in L6 myotubes [24]. The importance of these AA transporters is further bolstered by the fact that the heterodimer LAT1-CD98 also regulates the leucine flux into the lysosome [25], the major cellular compartment for mTORC1 activation [26], and that SNAT2 may act as a transceptor involved in both amino acid transport and signal transduction [27].
Although the mechanisms by which aging alters leucine-induced protein synthesis are currently under investigation, our data are in accordance with the hypothesis that a concerted activation of AA transporters may occur differently in young and older adults, finally influencing the anabolic response of skeletal muscle to AA availability. This hypothesis is supported by Drummond and Dickinson’s work showing that aging differentially affects the expression of SLC7A5 and SLC38A2 in the skeletal muscle, in response to resistance exercise and essential AA supplementation [14,17]. These evidences are also in keeping with our findings regarding SLC36A1 (PAT1), which has been reported to function as transceptor on the lysosomal membrane [28]. For this gene, we found the minor alleles at rs357618 and rs357629 (in LD with each other) associated with a better ADL performance. A different variant of SLC36A1 (minor allele of rs14160), was found, instead, to increase the risk of death along a 10 year follow up of survival in our younger sample. Similarly, different SLC3A2 variants showed cross-phenotype associations on different age-related phenotypes [29]. As to SNAT9 (SLC38A9), another lysosomal transceptor [30–32], we found a complex and dynamic association with survival chance. In fact, while a survival advantage was found for carriers of the rs10056358 A allele in the sample group aged 50-89 years, 90-plus subjects carrying this allele showed a lower probability to survive. Accordingly, the same minor allele A negatively influenced the probability to achieve longevity. Thus, this SNP manifests an antagonistic pleiotropic effect on survival at old ages, with the rs10056358-A allele conferring a positive effect on survival at ages before 90 and negative effect afterwards. The absence of correlation between SLC38A9 gene and the parameters of physical performance suggests that it functions as a pleiotropic gene with age-dependent effects, affecting survival and longevity but not quality of aging [33,34]. On the contrary, conditional effects of these genes on quality of aging but not on longevity are shown for the most part of the genes studied in this work.
Overall, our data provide evidence that the genetic variability of these genes may impact on muscle performance and/or physical decline in adulthood, as well as on the probability to survive at old age. Thus, taking into account that AA transporters may act as sensors of AA availability and that this availability is closely related to skeletal muscle metabolism, AA transporter genetic variability may act at forefront of individual susceptibility to anabolic resistance experienced with age, acting as risk factor for the onset of muscular decline in the elderly population.
Hence, understanding the functional implications of the associated SNPs may represent an important indication for elucidating the possible molecular mechanism underlying the associations found. However, the majority of the associated SNPs are non-coding variants, except two SNPs located in the coding regions. Our bioinformatic analyses identified, for some of them, a number of features (histone modifications, DNase I hypersensitivity clusters and transcription factor binding sites) consistent with a possible regulatory function, although we cannot exclude the possibility that the associated variants represent proxies for unknown causal variants as a result of LD, apart from those not showing any LD in their genomic region (rs12794763 and rs999165).
We are aware that our study has some weaknesses that should be addressed. A first limitation of the study is the lack of proper correction for multiple testing. Since this study was exploratory, a Bonferroni correction would have eliminated potentially important findings if applied. Another possible limitation could be the time of the survival follow up, not sufficient to draw long-term conclusions on the effect of genetic variants with minor effect on survival. Furthermore, the sample size could be increased and further explorations in additional study populations are needed before conclusions can be drawn.
Notwithstanding, and considering that this is the first study reporting genetic variants in AA transporter genes associated to the age-associated decline of muscle performance, we believe that our findings can open future investigations on the role of AA transporters in the quality of aging and longevity. This could provide valuable insights into potential targets for risk stratification in the population, and for therapeutic interventions aimed at increasing muscle mass and strength at old age.
Methods
Study population
The study was conducted on a sample of 765 subjects in the age range 50–107 years. The younger sample (age-range 50-89 years; mean age 70.40 ± 8.71) included 475 subjects (238 males and 237 females), the oldest old one (age range 90-108 years; mean age 96.86 ± 3.86) included 290 subjects (108 males and 182 females). See Table 1 for a complete description of the sample studied. All the subjects were born in Calabria (southern Italy) and their ancestry in the region had been ascertained up to the grandparents’ generation. Samples were collected within the framework of several recruitment campaigns carried out for monitoring the quality of aging in the whole Calabria region from 2002 onwards. Subjects older than 90 years were identified through the population registers and then contacted by specialised personnel and invited to join the study. Younger subjects were contacted through general physicians. Finally, each subject was recruited after a complete multidimensional geriatric assessment with detailed clinical history, including anthropometric measures and a set of the most common tests to assess cognitive functioning, functional activity, physical performance and depression. In addition, common clinical haematological tests were performed. White blood cells (WBC) from blood buffy coats were used as source of DNA.
Ethic statement
Recruitment campaigns and subsequent analyses received the approval of the relevant ethical committee. All the subjects provided written informed consent for the permission to collect blood samples and usage of register-based information for research purposes.
Physical performance
Hand Grip (HG) strength was measured by a handheld dynamometer (SMEDLEY's dynamometer TTM) while the subject was sitting with the arm close to his/her body. The test was repeated three times with the stronger hand and the maximum of these values was considered in the analyses. When a test was not carried out, it was specified if it was due to physical disabilities or because the subject refused to participate. Since HG strength is affected by age, sex, and height, the scores were corrected for these factors.
Walking time was measured as the best performance (shortest time in seconds) of two walks along a 4-meter distance.
Functional activity
The management of Activities of Daily Living (bathing, dressing, toileting, transfer from bed to chair, and feeding) was assessed using a modification of the Katz Index of ADL [35]. The assessment was based on what the subject was able to do at the time of the visit. The score is given counting the number of activities in which the participant is dependent or independent at the time of the visit. For the analyses, ADL scores were dichotomized as one if the subject was not independent in all five items and zero otherwise.
SNP selection and genotyping
A total of 58 SNPs mapping within and nearby genes encoding 10AA transporters were prioritized by a tagging approach, attempting to choose those most likely to be of functional relevance (nonsynonymous SNPs, SNPs located in the 5′ and 3′ UTR regions). Supplementary Table S1 reports the complete list of selected SNPs, their position (relative to the chromosome and to the gene), and putative functional annotation.
Multiplex SNP genotyping was performed using PCR followed by primer extension and MALDI-TOF mass spectrometry using iPLEX Gold technology from Sequenom (Sequenom Inc, San Diego, USA). Sequenom MassARRAY Assay Designer software (version 3) was used to design primers for PCR and single base extension. Standard procedures were used to amplify PCR products, and unincorporated nucleotides were deactivated with the shrimp alkaline phosphatase (SAP). A primer extension reaction was implemented by mass extension primer and terminator. The primer extension products were then desalted on resin, and spotted onto the 384-element SpectroCHIP (Sequenom) for MALDI-TOF analysis using Spec-troACQUIRE v3.3.1.3 (Sequenom). Spectra were analyzed using MassARRAY Typer v3.4 Software (Sequenom). Approximately 10% of the samples were analysed in duplicate, and the concordance rate of the genotypes was higher than 99%.
Quality-control
After genotype calling, the dataset went through a battery of quality-control (QC) tests. At sample level, subjects with a proportion of missing genotypes higher than 10% were excluded from the study. At SNP level, SNPs were excluded if they had a significant deviation from Hardy-Weinberg equilibrium (HWE, p<0.05), a Missing Frequency (MiF) higher than 10% and a Minor Allele Frequency (MAF) lower than 5%.
Statistical analyses
Continuous and categorical variables were compared by using the independent samples t-test and the chi-square test as appropriate. For each SNP, allele and genotype frequencies were estimated by gene counting from the observed genotypes. Hardy Weinberg Equilibrium (HWE) was tested by Fisher’s exact test. Pairwise measures of linkage disequilibrium (LD) between the analysed loci was estimated by Haploview (https://www.broadinstitute.org/haploview/haploview). Linear and logistic regression models were applied to estimate the impact of genetic variability on parameters of muscle strength (HG) and physical performance (WS and ADL), including as covariates age, gender, and height in the formulated regression models. A logistic regression model was also used to evaluate the effect of genetic variability on the chance to reach very advanced age. In these models, genetic data were coded with respect to a dominant, recessive, and additive fashion. Then, for each SNP the most likely genetic model was estimated on the basis of minimum level of statistical significance (Wald’ test p-value).
The hypothesis tested in this study is based on prior evidences for a role of AA transporters in the mTORC1-mediated induction of muscle protein synthesis. Furthermore, the involvement of genetic variability of AA transporters in relation to age-related muscle loss are lacking: in this sense, this study was exploratory. Thus, the p-values of single SNP analysis are reported without Bonferroni post hoc correction for multiple comparisons, to be not conservative and eliminate potentially important findings if applied.
In order to evaluate if the detected effects of the analysed polymorphisms on both muscle strength and physical performance might finally result in differential patterns of survival of the different relevant genotypes, we evaluated survival after 10 years from the baseline visit. Kaplan-Meier survival curves were estimated for each SNP affecting the analysed muscle-related phenotypes. In order to evaluate their predictive value with respect to mortality risk, the obtained survival curves were then compared by log-rank test. Subjects alive after the follow-up time were considered as censored, and this time was used as the censoring date in the survival analyses. In addition, Hazard ratios (HR) and 95% Confidence Intervals (95% CI) were estimated by using Cox proportional hazard models taking also into account possible confounder variables (age and gender).
Statistical analyses have been performed using SNPassoc and surv packages of R (http://www.R-project.org/).
Bio-informatic analyses
The functionality of theassociated SNPs was explored by retrieving regulatory information from the ENCODE (https://genome.ucsc.edu/ENCODE/) [36] and the Roadmap Epigenome Mapping projects (http://www.ppmroadmap.com/) [37] as implemented in HaploReg (v4.1, www.broadinstitute.org/mammals/haploreg/) [38], and RegulomeDB (www.regulomedb.org/) [39].
Supplementary Materials
Author Contributions
EH and PC carried out the genetic analyses and participated to the interpretation of data. FDR carried out the statistical analyses. AM carried out the statistical analyses and participated to the interpretation of data. SD and GR drafted the manuscript, generated the conception of the study and participated to the interpretation of data. GP participated in the interpretation of data and revision of the manuscript.
Acknowledgements
The work has been made possible by the collaboration with the nursing homes of SADEL S.p.A (San Teodoro, San Raffaele, Villa del Rosario, A.G.I. srl, SAVELLI HOSPITAL, Casa di Cura Madonna dello Scoglio), in the frame of the agreement “SOLUZIONI INNOVATIVE PER L’INNALZAMENTO DELLA SALUTE E DELLA SICUREZZA DELLA POPOLAZIONE” with the University of Calabria.
Conflicts of Interest
The authors have no conflicts of interest to declare.
Funding
This study was supported by grants from the Italian Ministry of University and Research (PRIN: Progetti di Ricerca di rilevante Interesse Nazionale – 2015, Prot. 20157ATSLF) to GR.
References
- 1. Janssen I. The epidemiology of sarcopenia. Clin Geriatr Med. 2011; 27:355–63. https://doi.org/10.1016/j.cger.2011.03.004 [PubMed]
- 2. Cesari M, Landi F, Vellas B, Bernabei R, Marzetti E. Sarcopenia and physical frailty: two sides of the same coin. Front Aging Neurosci. 2014; 6:192. https://doi.org/10.3389/fnagi.2014.00192 [PubMed]
- 3. Janssen I, Baumgartner RN, Ross R, Rosenberg IH, Roubenoff R. Skeletal muscle cutpoints associated with elevated physical disability risk in older men and women. Am J Epidemiol. 2004; 159:413–21. https://doi.org/10.1093/aje/kwh058 [PubMed]
- 4. Malafarina V, Uriz-Otano F, Iniesta R, Gil-Guerrero L. Sarcopenia in the elderly: diagnosis, physiopathology and treatment. Maturitas. 2012; 71:109–14. https://doi.org/10.1016/j.maturitas.2011.11.012 [PubMed]
- 5. Walston JD. Sarcopenia in older adults. Curr Opin Rheumatol. 2012; 24:623–27. https://doi.org/10.1097/BOR.0b013e328358d59b [PubMed]
- 6. Haran PH, Rivas DA, Fielding RA. Role and potential mechanisms of anabolic resistance in sarcopenia. J Cachexia Sarcopenia Muscle. 2012; 3:157–62. https://doi.org/10.1007/s13539-012-0068-4 [PubMed]
- 7. Dardevet D, Rémond D, Peyron MA, Papet I, Savary-Auzeloux I, Mosoni L. Muscle wasting and resistance of muscle anabolism: the “anabolic threshold concept” for adapted nutritional strategies during sarcopenia. Sci World J. 2012; 2012:269531. https://doi.org/10.1100/2012/269531 [PubMed]
- 8. Mitchell WK, Wilkinson DJ, Phillips BE, Lund JN, Smith K, Atherton PJ. Human Skeletal Muscle Protein Metabolism Responses to Amino Acid Nutrition. Adv Nutr. 2016; 7:828S–38S. https://doi.org/10.3945/an.115.011650 [PubMed]
- 9. Volpi E, Kobayashi H, Sheffield-Moore M, Mittendorfer B, Wolfe RR. Essential amino acids are primarily responsible for the amino acid stimulation of muscle protein anabolism in healthy elderly adults. Am J Clin Nutr. 2003; 78:250–58. https://doi.org/10.1093/ajcn/78.2.250 [PubMed]
- 10. Dickinson JM, Rasmussen BB. Amino acid transporters in the regulation of human skeletal muscle protein metabolism. Curr Opin Clin Nutr Metab Care. 2013; 16:638–44. https://doi.org/10.1097/MCO.0b013e3283653ec5 [PubMed]
- 11. Taylor PM. Role of amino acid transporters in amino acid sensing. Am J Clin Nutr. 2014; 99:223S–30S. https://doi.org/10.3945/ajcn.113.070086 [PubMed]
- 12. Goberdhan DC, Wilson C, Harris AL. Amino Acid Sensing by mTORC1: Intracellular Transporters Mark the Spot. Cell Metab. 2016; 23:580–89. https://doi.org/10.1016/j.cmet.2016.03.013 [PubMed]
- 13. Shimobayashi M, Hall MN. Multiple amino acid sensing inputs to mTORC1. Cell Res. 2016; 26:7–20. https://doi.org/10.1038/cr.2015.146 [PubMed]
- 14. Dickinson JM, Drummond MJ, Coben JR, Volpi E, Rasmussen BB. Aging differentially affects human skeletal muscle amino acid transporter expression when essential amino acids are ingested after exercise. Clin Nutr. 2013; 32:273–80. https://doi.org/10.1016/j.clnu.2012.07.009 [PubMed]
- 15. Drummond MJ, Glynn EL, Fry CS, Timmerman KL, Volpi E, Rasmussen BB. An increase in essential amino acid availability upregulates amino acid transporter expression in human skeletal muscle. Am J Physiol Endocrinol Metab. 2010; 298:E1011–18. https://doi.org/10.1152/ajpendo.00690.2009 [PubMed]
- 16. Drummond MJ, Fry CS, Glynn EL, Timmerman KL, Dickinson JM, Walker DK, Gundermann DM, Volpi E, Rasmussen BB. Skeletal muscle amino acid transporter expression is increased in young and older adults following resistance exercise. J Appl Physiol (1985). 2011; 111:135–42. https://doi.org/10.1152/japplphysiol.01408.2010 [PubMed]
- 17. Drummond MJ, Dickinson JM, Fry CS, Walker DK, Gundermann DM, Reidy PT, Timmerman KL, Markofski MM, Paddon-Jones D, Rasmussen BB, Volpi E. Bed rest impairs skeletal muscle amino acid transporter expression, mTORC1 signaling, and protein synthesis in response to essential amino acids in older adults. Am J Physiol Endocrinol Metab. 2012; 302:E1113–22. https://doi.org/10.1152/ajpendo.00603.2011 [PubMed]
- 18. Dickinson JM, Gundermann DM, Walker DK, Reidy PT, Borack MS, Drummond MJ, Arora M, Volpi E, Rasmussen BB. Leucine-enriched amino acid ingestion after resistance exercise prolongs myofibrillar protein synthesis and amino acid transporter expression in older men. J Nutr. 2014; 144:1694–702. https://doi.org/10.3945/jn.114.198671 [PubMed]
- 19. Chapel A, Kieffer-Jaquinod S, Sagné C, Verdon Q, Ivaldi C, Mellal M, Thirion J, Jadot M, Bruley C, Garin J, Gasnier B, Journet A. An extended proteome map of the lysosomal membrane reveals novel potential transporters. Mol Cell Proteomics. 2013; 12:1572–88. https://doi.org/10.1074/mcp.M112.021980 [PubMed]
- 20. Bhutia YD, Ganapathy V. Glutamine transporters in mammalian cells and their functions in physiology and cancer. Biochim Biophys Acta. 2016; 1863:2531–39. https://doi.org/10.1016/j.bbamcr.2015.12.017 [PubMed]
- 21. McLean RR, Shardell MD, Alley DE, Cawthon PM, Fragala MS, Harris TB, Kenny AM, Peters KW, Ferrucci L, Guralnik JM, Kritchevsky SB, Kiel DP, Vassileva MT, et al. Criteria for clinically relevant weakness and low lean mass and their longitudinal association with incident mobility impairment and mortality: the foundation for the National Institutes of Health (FNIH) sarcopenia project. J Gerontol A Biol Sci Med Sci. 2014; 69:576–83. https://doi.org/10.1093/gerona/glu012 [PubMed]
- 22. Meynial-Denis D. Glutamine metabolism in advanced age. Nutr Rev. 2016; 74:225–36. https://doi.org/10.1093/nutrit/nuv052 [PubMed]
- 23. Evans K, Nasim Z, Brown J, Butler H, Kauser S, Varoqui H, Erickson JD, Herbert TP, Bevington A. Acidosis-sensing glutamine pump SNAT2 determines amino acid levels and mammalian target of rapamycin signalling to protein synthesis in L6 muscle cells. J Am Soc Nephrol. 2007; 18:1426–36. https://doi.org/10.1681/ASN.2006091014 [PubMed]
- 24. Luo JQ, Chen DW, Yu B. Upregulation of amino acid transporter expression induced by L-leucine availability in L6 myotubes is associated with ATF4 signaling through mTORC1-dependent mechanism. Nutrition. 2013; 29:284–90. https://doi.org/10.1016/j.nut.2012.05.008 [PubMed]
- 25. Milkereit R, Persaud A, Vanoaica L, Guetg A, Verrey F, Rotin D. LAPTM4b recruits the LAT1-4F2hc Leu transporter to lysosomes and promotes mTORC1 activation. Nat Commun. 2015; 6:7250. https://doi.org/10.1038/ncomms8250 [PubMed]
- 26. Yao Y, Jones E, Inoki K. Lysosomal Regulation of mTORC1 by Amino Acids in Mammalian Cells. Biomolecules. 2017; 7:E51. https://doi.org/10.3390/biom7030051 [PubMed]
- 27. Pinilla J, Aledo JC, Cwiklinski E, Hyde R, Taylor PM, Hundal HS. SNAT2 transceptor signalling via mTOR: a role in cell growth and proliferation? Front Biosci (Elite Ed). 2011; 3:1289–99. [PubMed]
- 28. Zoncu R, Bar-Peled L, Efeyan A, Wang S, Sancak Y, Sabatini DM. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science. 2011; 334:678–83. https://doi.org/10.1126/science.1207056 [PubMed]
- 29. Solovieff N, Cotsapas C, Lee PH, Purcell SM, Smoller JW. Pleiotropy in complex traits: challenges and strategies. Nat Rev Genet. 2013; 14:483–95. https://doi.org/10.1038/nrg3461 [PubMed]
- 30. Jung J, Genau HM, Behrends C. AA dependent mTORC1 regulation by the lysosomal membrane protein SLC38A9. Mol Cell Biol. 2015; 35:2479–94. https://doi.org/10.1128/MCB.00125-15 [PubMed]
- 31. Wang S, Tsun ZY, Wolfson RL, Shen K, Wyant GA, Plovanich ME, Yuan ED, Jones TD, Chantranupong L, Comb W, Wang T, Bar-Peled L, Zoncu R, et al. Metabolism. Lysosomal amino acid transporter SLC38A9 signals arginine sufficiency to mTORC1. Science. 2015; 347:188–94. https://doi.org/10.1126/science.1257132 [PubMed]
- 32. Rebsamen M, Pochini L, Stasyk T, de Araújo ME, Galluccio M, Kandasamy RK, Snijder B, Fauster A, Rudashevskaya EL, Bruckner M, Scorzoni S, Filipek PA, Huber KV, et al. SLC38A9 is a component of the lysosomal amino acid sensing machinery that controls mTORC1. Nature. 2015; 519:477–81. https://doi.org/10.1038/nature14107 [PubMed]
- 33. Yashin AI, Wu D, Arbeeva LS, Arbeev KG, Kulminski AM, Akushevich I, Kovtun M, Culminskaya I, Stallard E, Li M, Ukraintseva SV. Genetics of aging, health, and survival: dynamic regulation of human longevity related traits. Front Genet. 2015; 6:122. https://doi.org/10.3389/fgene.2015.00122 [PubMed]
- 34. Ukraintseva S, Yashin A, Arbeev K, Kulminski A, Akushevich I, Wu D, Joshi G, Land KC, Stallard E. Puzzling role of genetic risk factors in human longevity: “risk alleles” as pro-longevity variants. Biogerontology. 2016; 17:109–27. https://doi.org/10.1007/s10522-015-9600-1 [PubMed]
- 35. Katz S, Downs TD, Cash HR, Grotz RC. Progress in development of the index of ADL. Gerontologist. 1970; 10:20–30. https://doi.org/10.1093/geront/10.1_Part_1.20 [PubMed]
- 36. Consortium EP, and ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012; 489:57–74. https://doi.org/10.1038/nature11247 [PubMed]
- 37. Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, Kheradpour P, Zhang Z, Wang J, Ziller MJ, Amin V, Whitaker JW, Schultz MD, et al, and Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature. 2015; 518:317–30. https://doi.org/10.1038/nature14248 [PubMed]
- 38. Ward LD, Kellis M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016; 44:D877–81. https://doi.org/10.1093/nar/gkv1340 [PubMed]
- 39. Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, Karczewski KJ, Park J, Hitz BC, Weng S, Cherry JM, Snyder M. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012; 22:1790–97. https://doi.org/10.1101/gr.137323.112 [PubMed]


