Introduction
Breast cancer (BC) is an enormous public health burden worldwide and ranks as the main cause of cancer deaths in China [1]. With the advances of the comprehensive therapeutic strategies, the 5-year overall survival (OS) rate of BC has been improve dramatically. However, BC kills about 1.2 million people in China each year [2]. Prognostic evaluation is vital for making appropriate therapeutic decisions and follow-up strategies in BC patients. Currently, tumor-node-metastasis (TNM) stage is a key tool for prognostic assessment and a specific treatment choices. However, BC patients at same TNM staging can have very different clinical outcomes. The traditional TNM staging system is mainly on the basis of anatomical information, which is unable to sufficient prediction for prognosis of individual patients because it could not display the biological heterogeneity of BC [3]. Therefore, a predictive tool that can integrate molecular biomarkers into the TNM staging system may improve the accuracy of survival prediction for BC patients.
MicroRNAs (miRNAs) are small, non-coding single-stranded RNAs (18–25 nucleotides) and negatively regulate gene expression by base-pair matching with the 3′UTRs of target mRNAs [4]. Accumulating evidence shows that miRNAs play critical roles in various physiological and pathological processes, including metabolism, carcinogenesis, and proliferation [4, 5]. In addition, previous reports have suggested the important prognostic value of miRNAs signature in a variety of cancers [6–20]. But most of these studies were based on limited number of patients and different miRNAs platforms, lack a normalized standard. Thus, the Cancer Genome Atlas (TCGA) database provides us with a comprehensive catalogue of large-scale miRNAs expression data. Besides, the prognostic value of miRNAs signature to predict 5-year OS of BC patients has not been fully illustrated. With the ability of incorporating diverse independent prognostic variables to provide an individual probability of survival outcome, nomogram is widely applied for cancer prognosis [3, 21].
Therefore, this study aimed to construct a novel miRNA-based model to improve survival prediction and effectively pick out the high-risk patients based on TCGA miRNA sequencing data. Such a practical tool has the potential to guide more effective individualized treatment decisions for BC patients.
Results
Baseline characteristics of patients
A total of 984 BC patients from TCGA database were included. The detailed baseline characteristics of the primary and validation cohort were listed in Table 1. No significant difference of baseline characteristics were displayed between the two independent cohort in Table 1 (all P>0.05). The median age of the 984 BC patients was 58 year (interquartile range [IQR]: 48–67). The 5-year OS rate of the 984 BC patients was 82.6%.
Table 1. Baseline characteristics of study patients
| Variables | Primary cohort No. (%) | Validation cohort No. (%) | P-value |
|---|---|---|---|
| No. of patients | 984 | 492 | |
| Age (years) | 58(48,67) | 58(48,67) | 0.767 |
| T stage | 0.620 | ||
| T1 | 262(26.6) | 132(26.8) | |
| T2 | 570(57.9) | 295(59.9) | |
| T3 | 123(12.5) | 50(10.2) | |
| T4 | 29(3.0) | 15(3.1) | |
| N stage | 0.987 | ||
| N0 | 453(46.0) | 222(45.1) | |
| N1 | 341(34.7) | 169(34.4) | |
| N2 | 107(10.9) | 56(11.4) | |
| N3 | 72(7.3) | 39(7.9) | |
| Unknown | 11(1.1) | 6(1.2) | |
| TNM stage | 0.980 | ||
| I | 170(17.3) | 82(16.7) | |
| II | 569(57.8) | 284(57.7) | |
| III | 226(23.0) | 117(23.8) | |
| IV | 19(1.9) | 9(1.8) | |
| ER status | 0.925 | ||
| Negative | 206(20.9) | 101(20.5) | |
| Positive | 738(75.0) | 369(75.0) | |
| Unknown | 40(4.1) | 22(4.5) | |
| PR status | 0.878 | ||
| Negative | 262(26.6) | 137(27.9) | |
| Positive | 598(60.8) | 295(59.9) | |
| Unknown | 124(12.6) | 60(12.2) | |
| HER2 status | 0.980 | ||
| Negative | 679(69.0) | 337(68.5) | |
| Positive | 152(15.5) | 77(15.6) | |
| Unknown | 153(15.5) | 78(15.9) |
Development of risk score formula and six-miRNA-based prognostic model
To facilitate the utility of the identified prognostic miRNAs in routine clinical practice, the following formula was developed to generate risk score for each patient: Risk score=(0.289×expressionmiR-549a)+ (0.072× expressionmiR-6715a)+(0.026×expressionmiR-4501)+ (0.158× expressionmiR-7974)+(0.068×expressionmiR-4675)-(0.054× expressionmiR-147b). Thus, patients were classified into the low-risk group and the high-risk group via the same median risk score as the cut-off point in the two independent cohorts. The distributions of the miRNA-based risk scores, OS, OS status, and six-miRNA expression profiles of the training cohort and validation cohort are showed in Figure 2. The heat map suggests that the five risky miRNAs (hsa-miR-549a, hsa-miR-6715a, hsa-miR-4501, hsa-miR-7974, hsa-miR-4675) have high expression in the high-risk group, while the one protective miRNA (hsa-miR-147b) exhibits high expression in low-risk group (Figure 2). Besides, compared with the low-risk group, Kaplan–Meier survival analysis shows that the high-risk group has a obvious poorer prognosis (P<0.0001) (Figure 3).
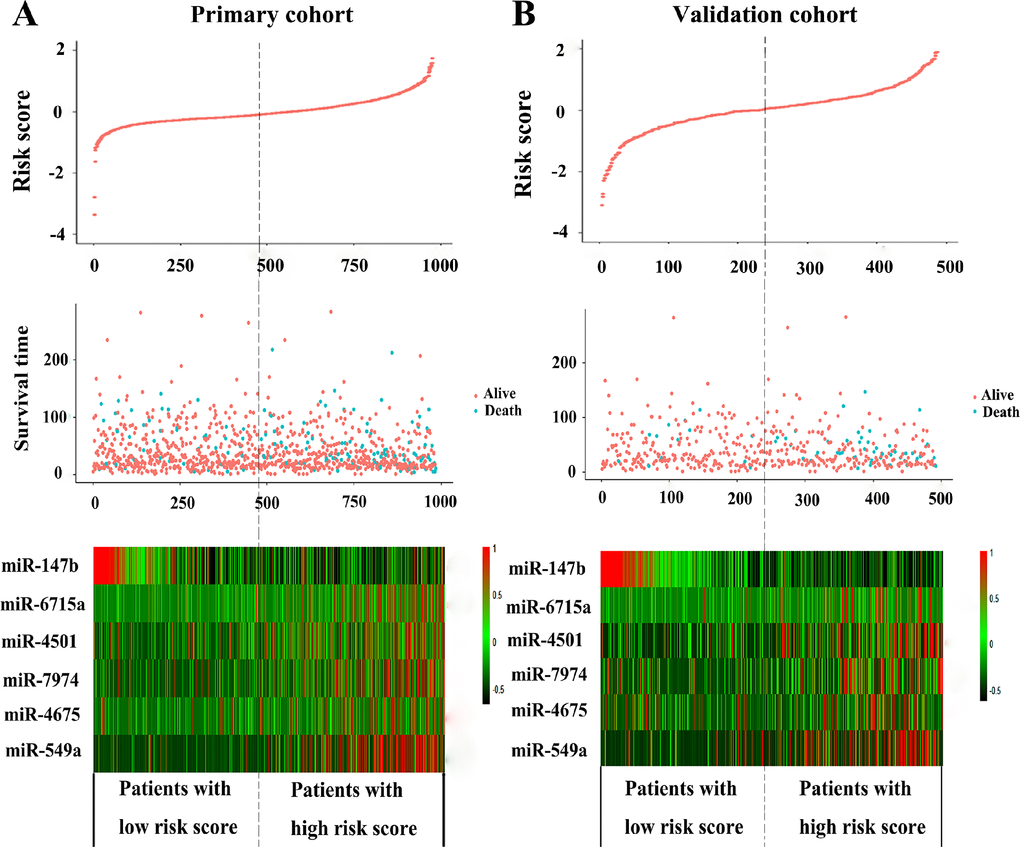
Figure 2. The distribution of risk score, OS, and OS status and the heat map of prognostic six-miRNA signature in the primary cohort (A) and validation cohort (B). The dotted line indicates the cutoff point of the median risk score used to stratify patients into the low-risk group and high-risk group. OS, overall survival.
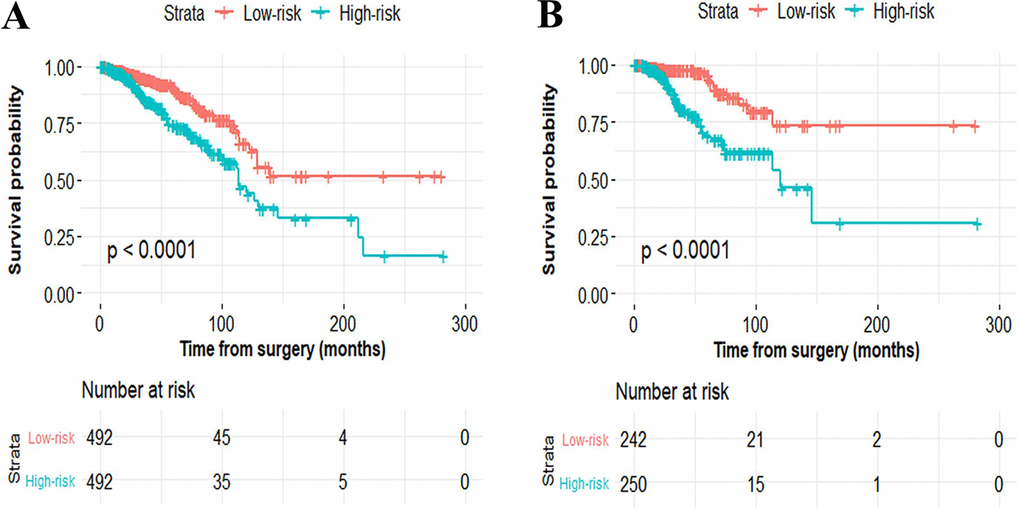
Figure 3. Kaplan–Meier curves of overall survival for breast cancer patients based on the six-miRNA signature in the primary cohort (A) and validation cohort (B) Kaplan–Meier curves of overall survival for breast cancer patients based on the six-miRNA signature in the primary cohort (A) and validation cohort (B).
According to the results of univariate and multivariate CPHR analyses (Table 3), the six-miRNA signature and three clinical risk factors (age, TNM stage and ER status) were identified as independent prognostic variables of OS. T stage and N stage were not entered into multivariate CPHR analysis, because they were associated with TNM stage, known as multicollinearity, could lead to spurious associations and unreliable results [21]. To construct a more sensitive predictive tool in clinical practice, we built a novel six-miRNA-based prognostic model integrating the six-miRNA signature and three clinical risk factors (age, TNM stage and ER status) to predict 5-year OS of BC patients (Figure 4). The six-miRNA-based nomogram revealed the six-miRNA signature and TNM stage as the largest contribution to 5-year OS, followed by the age and ER status. Each variable was acquired a nomogram score on the point scale. After calculating the total nomogram score, we could easily obtain the nomogram-predicted probability of 5-year OS for each patient.
Table 3. Univariate and multivariate Cox proportional hazards regression analyses in the primary cohort
| Variables | Univariate analysis | Multivariate analysis | ||
|---|---|---|---|---|
| Hazard ratios (95%CI) | P-value | Hazard ratios (95%CI) | P-value | |
| Age | 1.029(1.016-1.042) | <0.001 | 1.031(1.017-1.045) | <0.001 |
| T stage | ||||
| T1 | Referent | |||
| T2 | 1.620(1.037-2.530) | 0.034 | ||
| T3 | 1.623(0.911-2.889) | 0.100 | ||
| T4 | 4.402(2.194-8.830) | <0.001 | ||
| N stage | ||||
| N0 | Referent | |||
| N1 | 2.035(1.350-3.069) | 0.001 | ||
| N2 | 2.977(1.745-5.081) | <0.001 | ||
| N3 | 4.246(2.287-7.882) | <0.001 | ||
| Unknown | 7.666(3.001-19.582) | <0.001 | ||
| TNM stage | ||||
| I | Referent | Referent | ||
| II | 2.011 | 0.021 | 2.142(1.179-3.893) | 0.012 |
| III | 3.269 | <0.001 | 3.850(2.042-7.256) | <0.001 |
| IV | 12.784 | <0.001 | 15.909(7.474-33.865) | <0.001 |
| ER status | ||||
| Negative | Referent | Referent | ||
| Positive | 0.652(0.444-0.958) | 0.029 | 0.508(0.342-0.754) | 0.001 |
| Unknown | 1.918(0.921-3.997) | 0.082 | 1.755(0.837-3.678) | 0.137 |
| PR status | ||||
| Negative | Referent | |||
| Positive | 0.737(0.498-1.090) | 0.127 | ||
| Unknown | 1.260(0.757-2.098) | 0.374 | ||
| HER2 status | ||||
| Negative | Referent | |||
| Positive | 1.288(0.778- 2.134) | 0.325 | ||
| Unknown | 1.459(0.969-2.196) | 0.071 | ||
| Six-miRNA signature | 1.194(1.140-1.252) | <0.001 | 1.193(1.138-1.251) | <0.001 |
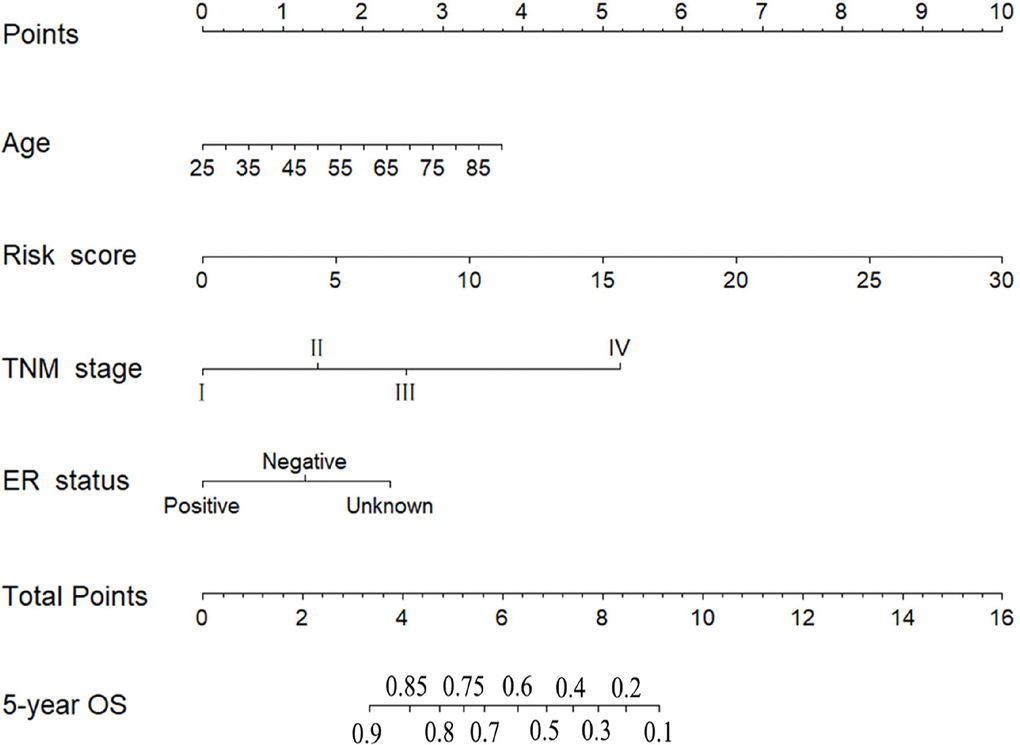
Figure 4. Six-miRNA-based prognostic model to predict 5-year overall survival in breast cancer patients Six-miRNA-based prognostic model to predict 5-year overall survival in breast cancer patients.
Assessment of the six-miRNA-based signature and prognostic model
To test whether the six-miRNA signature could predict OS regardless of stages, we performed risk stratification in patients with TNM stage, T stage, and N stage. The patients with low-risk scores had significantly better OS than patients with high-risk scores in TNM stage II (P=0.00063), TNM stage III (P=0.001), T2 (P=0.00015), T3 (P=0.0076), N1 (P=0.021), N2 (P=0.021) and N3 (P=0.018) (Figure 5). To assess the predictive performance of the six-miRNA-based signature and prognostic nomogram, we conducted a time-dependent ROC curve analysis by comparing the respective AUC value. Then, the AUC values of the six-miRNA signature at 5 years were 0.701 (95%CI: 0.633–0.768) and 0.789 (95%CI: 0.715–0.880) in the primary cohort and validation cohort, respectively (Figure 6A–6B). And the AUC values of the six-miRNA-based prognostic model at 5 years were 0.758 (95%CI: 0.686–0.830) and 0.777 (95%CI: 0.687–0.867) in the primary cohort and validation cohort, respectively (Figure 6C–6D). Importantly, these AUC values revealed that six-miRNA-based signature and prognostic nomogram had favorable discrimination performance for BC patients. In addition, calibration plots of the six-miRNA-based prognostic model fitted well in the training cohort and validation cohort, which indicated good calibration ability (Figure 7).
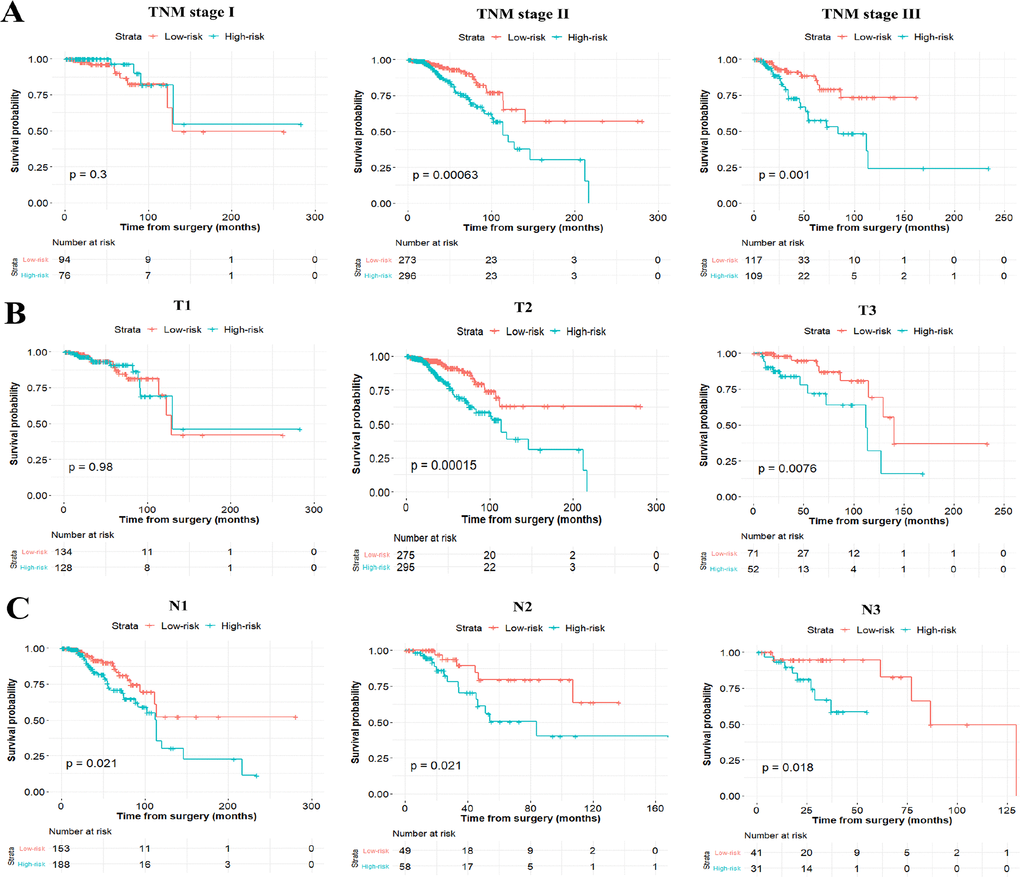
Figure 5. Stratified analysis of the six-miRNA signature for breast cancer patients in TNM stage (A), T stage (B), and N stage (C) Stratified analysis of the six-miRNA signature for breast cancer patients in TNM stage (A), T stage (B), and N stage (C).
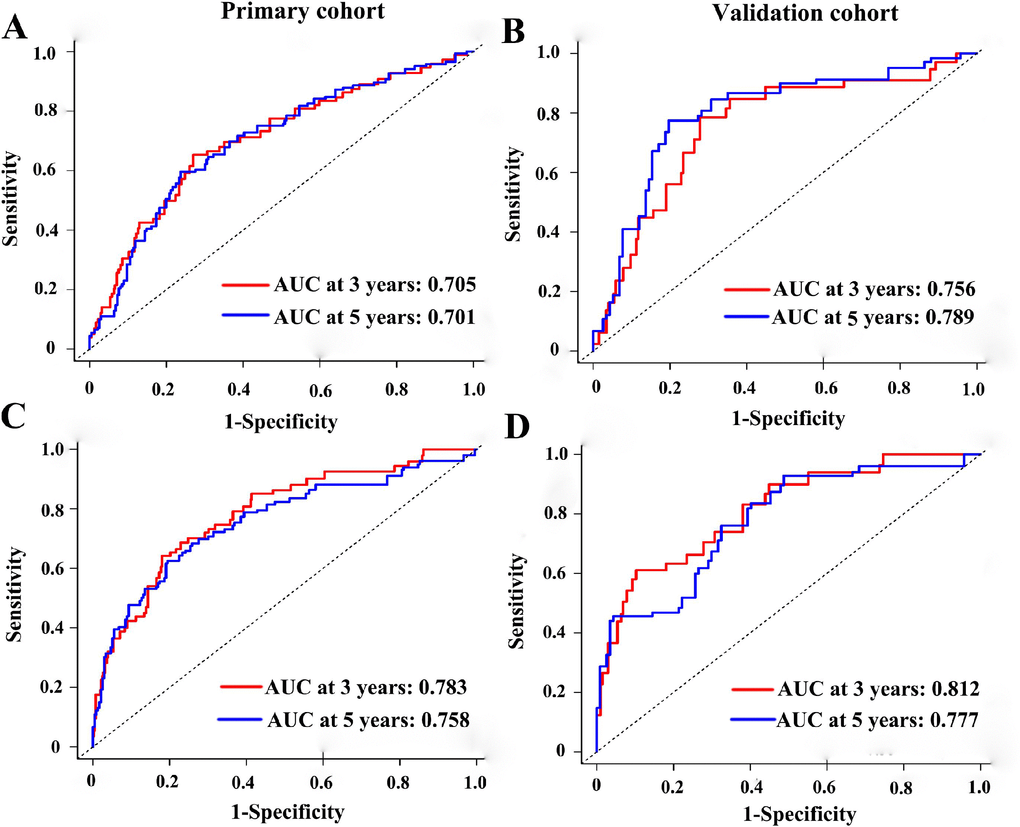
Figure 6. Time-dependent receiver operating characteristic curves at 3-, 5-years based on the six-miRNA signature in the primary cohort (A) and validation cohort (B). Time-dependent receiver operating characteristic curves at 3-, 5-years based on the six-miRNA-based prognostic model in the primary cohort (C) and validation cohort (D).
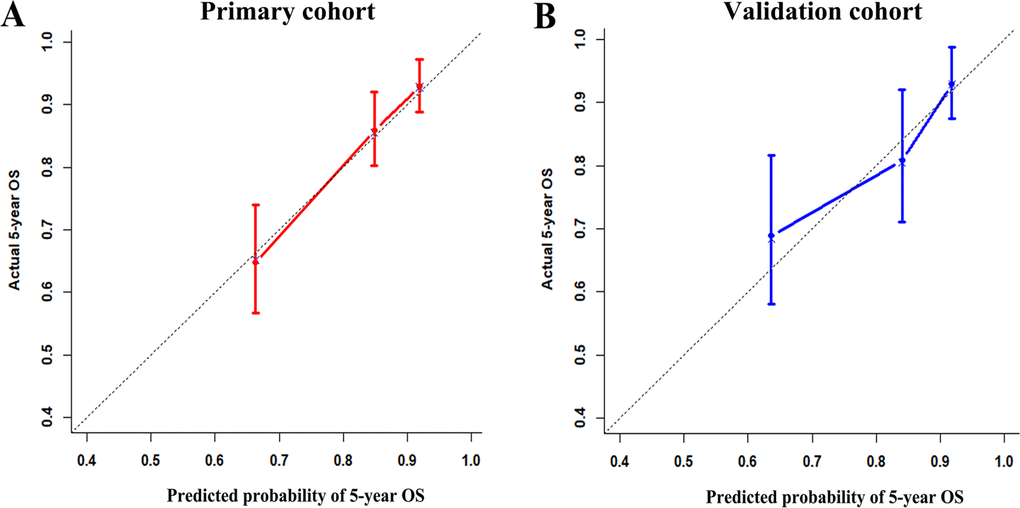
Figure 7. Calibration plots of the six-miRNA-based prognostic model in the primary cohort (A) and validation cohort (B) Calibration plots of the six-miRNA-based prognostic model in the primary cohort (A) and validation cohort (B).
Comparison with other prognostic factors
In ROC analysis to compare predictive accuracy of different prognostic factors, the six-miRNA signature suggested higher prognostic accuracy than clinical risk factors, or single miRNA alone (Figure 8A–8B). Thus, the six-miRNA signature can outperform the clinical prognostic features. More importantly, the six-miRNA-based prognostic nomogram had significantly better predictive performance than TNM stage (0.758 vs 0.650, P<0.001) (Figure 8C).
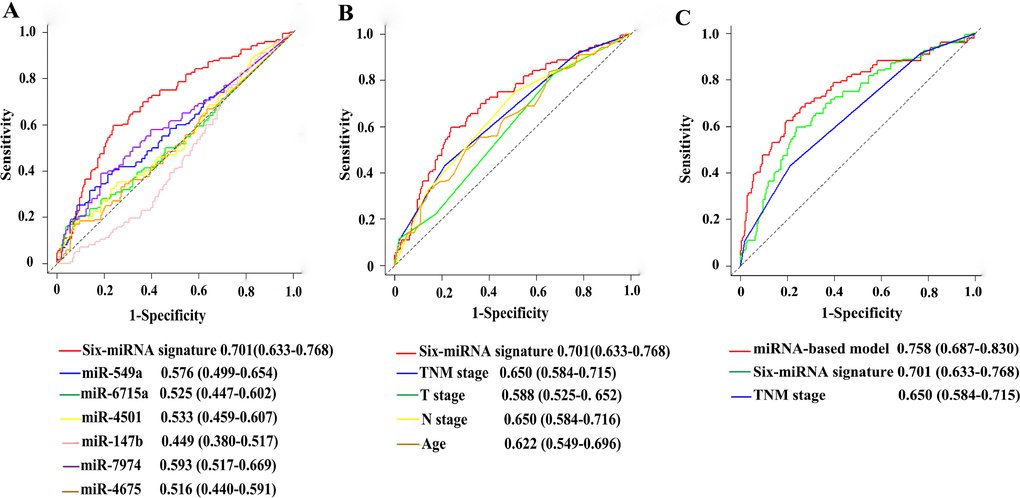
Figure 8. Comparisons of the prognostic accuracy at 5-years using time-dependent receiver operating characteristic curves in the six-miRNA signature with single miRNA (A), the six-miRNA signature with clinical risk factors (B), and the six-miRNA-based prognostic model with six-miRNA signature, TNM stage (C) Comparisons of the prognostic accuracy at 5-years using time-dependent receiver operating characteristic curves in the six-miRNA signature with single miRNA (A), the six-miRNA signature with clinical risk factors (B), and the six-miRNA-based prognostic model with six-miRNA signature, TNM stage (C).
GO and KEGG pathway analyses of predicted target genes
To evaluate the potential function of the six-miRNAs, a total of 39 target genes of the six-miRNAs were predicted using TargetScan, miRTarBase and miRDB database, respectively. GO analysis included molecular function (MF), biological process (BP), and cellular component (CC). The 39 target genes were mainly related with protein binding (MF), transcription and DNA-templated (BP), cytoplasm and nucleus (CC) (Figure 9A). And KEGG pathway analysis revealed that the 39 genes mainly enriched in MAPK signaling pathway, transcriptional misregulation in cancer and cAMP signaling pathway (Figure 9B).
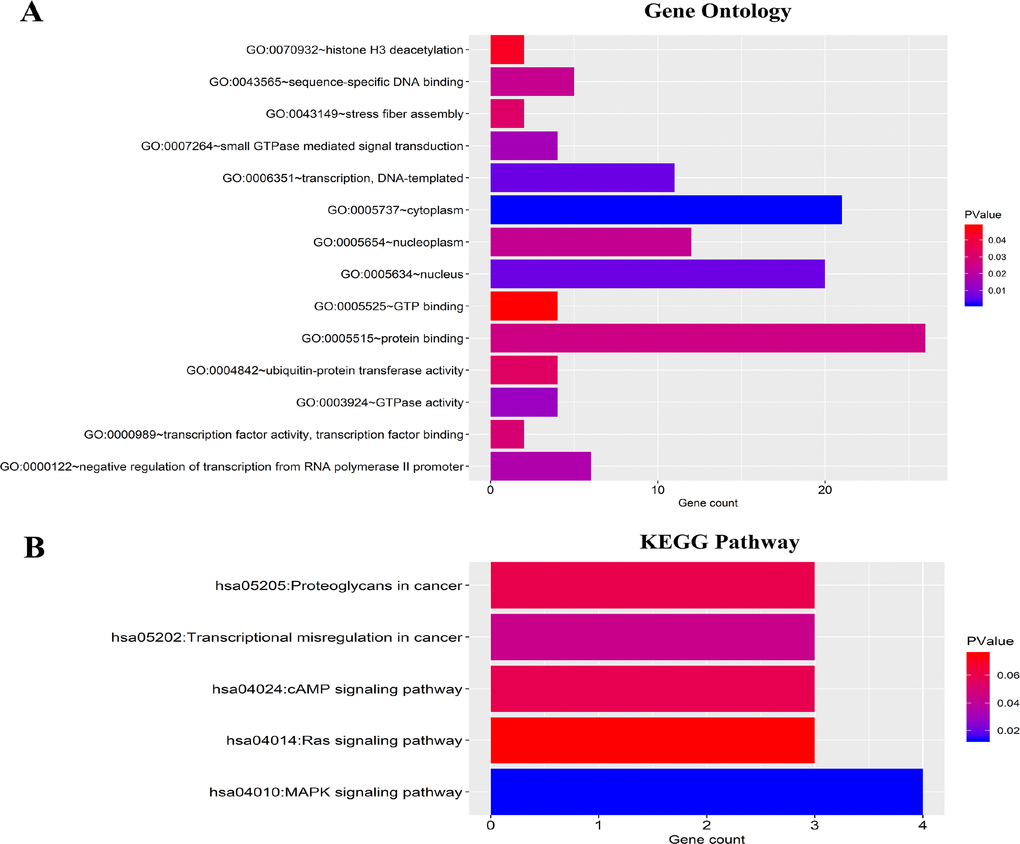
Figure 9. Functional enrichment analysis for predicted target genes of the six miRNAs. (A) Gene ontology (GO) enrichment analysis. (B) Kyoto Encyclopedia of Genes and Genomes analyses (KEGG) enrichment analysis. The x-axis indicates the number of genes, and the y-axis represents the GO terms and KEGG pathway names. The color represents the P-value.
Discussion
A molecular marker-based approach to accurately predict survival in BC patients is urgently needed in the era of precision medicine. Accumulating evidence indicates that miRNAs play a vital role in BC prognosis[10, 22–24]. In the present study, we confirmed six-miRNA signature that was significantly associated with OS in BC patients based on the TCGA database. Furthermore, this six-miRNA signature enabled to stratify patients into the low-risk and high-risk groups with distinct differences in 5-year OS. Moreover, a novel six-miRNA-based prognostic model combining six-miRNA signature and clinical risk factors was established and validated to improve survival prediction for BC patients. The six-miRNA-based nomogram consisted of four independent prognostic variables, including age, TNM stage, ER status, and six-miRNA signature. The proposed tool was significantly superior to the traditional TNM stage in predicting 5-year OS for BC patients. The AUC value of the six-miRNA-based prognostic model was 0.758, which indicating favorable discrimination performance. Therefore, our six-miRNA-based nomogram might be a vital tool for survival prediction in BC patients, aiding in personalized therapeutic treatment strategies and postoperative counseling. Further bioinformatics analysis helps us understand the biological function of the six OS-related miRNAs. On the basis of the GO and KEGG pathway analyses, the six-miRNAs may play crucial roles in protein binding, transcription and DNA-templated, cytoplasm, nucleus, MAPK signaling pathway, transcriptional misregulation in cancer and cAMP signaling pathway.
Previous reports about DEMs have indicated that the miRNA-based signature is a important marker for survival or relapse in a variety of cancers [8–10, 12, 14, 23–27]. Recently, Gong et al built a miRNA-based classifier to predict relapse in Hormone Receptor-Positive HER2-Negative BC patients [24]. However, this study has been limited by small sample size and small number of miRNAs screened to mine miRNA expression profiling. In addition, many researches were inconsistent in these sets of prognostic miRNAs because of the heterogeneous of BC and variations in the methods for miRNAs selection. TCGA database provides a robust platform to systematically analyze the large-scale miRNA sequencing data. Consequently, compared with above previous study, a total of 1601 miRNAs were initially selected in our study, which could provide a more comprehensive analysis. Besides, the miRNA signature from the TCGA database regarding to the 5-year OS of BC patients has not been reported.
Although the miRNA-based model performs well in BC survival prediction, there are several shortcomings should be acknowledged in the present study. First, experimental studies should be conducted to deeply explore the molecular mechanisms of these miRNAs in the future. Second, the TCGA database lacks some important postoperative variables (chemotherapy, radiotherapy, hormone therapy), thus we could not carry out a comprehensive analysis and identify the low-risk patients to tailor adjuvant chemotherapy. Third, multicenter, large-scale, prospective studies should be performed to validate this predictive tool before application in routine clinical practice. Fourth, the risk score could accurately discriminate patients with N1, N2 and N3 status. However, the risk score did not accurately discriminate patients with N0 status. Indeed, 21-gene expression Oncotype DX was considered as an accurate molecular tool to discriminate high-risk patients with N0 status and put the indication of adjuvant chemotherapy [28]. Thus, the risk score and 21-gene expression Oncotype DX could be used to identify the low-risk patients whether benefit from adjuvant chemotherapy, regardless of N status.
In conclusion, the current study showed a novel, robust six-miRNA-based prognostic model incorporating six-miRNA signature and clinical risk factors to predict 5-year OS in BC patients. The six-miRNA-based nomogram had higher prognostic value than the conventional TNM stage in BC patients. Furthermore, the six-miRNA signature can effectively identify the low risk patients from the high risk group in BC patients. Therefore, this practical tool has the potential to facilitate individualized treatment decisions for BC patients.
Material and methods
Patients and study design
In this study, the raw counts of BC dataset (Level 3 miRNA expression profiles), including 1044 BC samples and 102 adjacent normal breast tissues were acquired from TCGA data portal in September 2, 2018. The inclusion criteria were included: (1) histologically confirmed invasive BC; (2) both miRNA expression profile and complete survival information available; (3) OS time was more than 1 month. Finally, a total of 984 BC patients with the corresponding clinical features including, age, T stage, N stage, TNM stage, estrogen receptor (ER) status, progestrone receptor (PR) status, human epithelial growth factor receptor 2 (HER2) status were enrolled as primary cohort in this study. And we acquired data from 984 patients randomly assigned 492 patients as the validation cohort based on a computer-generated allocation sequence. Because the application of data abided by the TCGA publication guidelines, the approval of institutional ethics committees was not required.
Construction of risk score formula and miRNA-based prognostic model
Prognostic miRNAs which were distinctly associated with OS in the multivariate CPHR analysis (P<0.05) were pointed out to develop the risk score formula. The formula was carried to compute the prognostic risk score for each patient. Using the coefficients obtained from the multivariable CPHR analysis, a risk score formula was built as following: Risk score (miRNA-based classifier) = sum of coefficients × expression level of miRNAs. Moreover, the BC patients were stratified into the high-risk group and the low-risk group via the median risk score as the cutoff value. To provide the oncologists and patients with a quantitative method to achieve individualized survival prediction, we constructed a prognostic nomogram that incorporated both the miRNA-based signature and clinical risk factors using Cox regression model.
Evaluation of risk score formula and miRNA-based prognostic model
To further assess the predictive performance of the miRNA-based classifier and prognostic model, we measured the area under the curve (AUC) based on time-dependent receiver operating characteristic (ROC) analysis [30]. Furthermore, stratified analysis was conducted to test whether the miRNA-based classifier was associated with OS independent of stages. In addition, calibration curve was used to evaluate the agreement between model predicted outcome and actual outcome. The predictive accuracy of miRNA-based classifier and prognostic model were compared with other risk factors using ROC analysis.
Target gene prediction and functional enrichment analysis
Potential target genes of prognostic miRNAs were predicted via three online databases, including TargetScan, miRTarBase and miRDB [31–33]. Thus, we confirmed the overlapping miRNA target genes from the three online databases to perform enrichment analysis. The Database for Annotation, Visualization, and Integrated Discovery 6.8 Bioinformatics Tool (DAVID 6.8) was carried out to Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis.
Statistical analysis
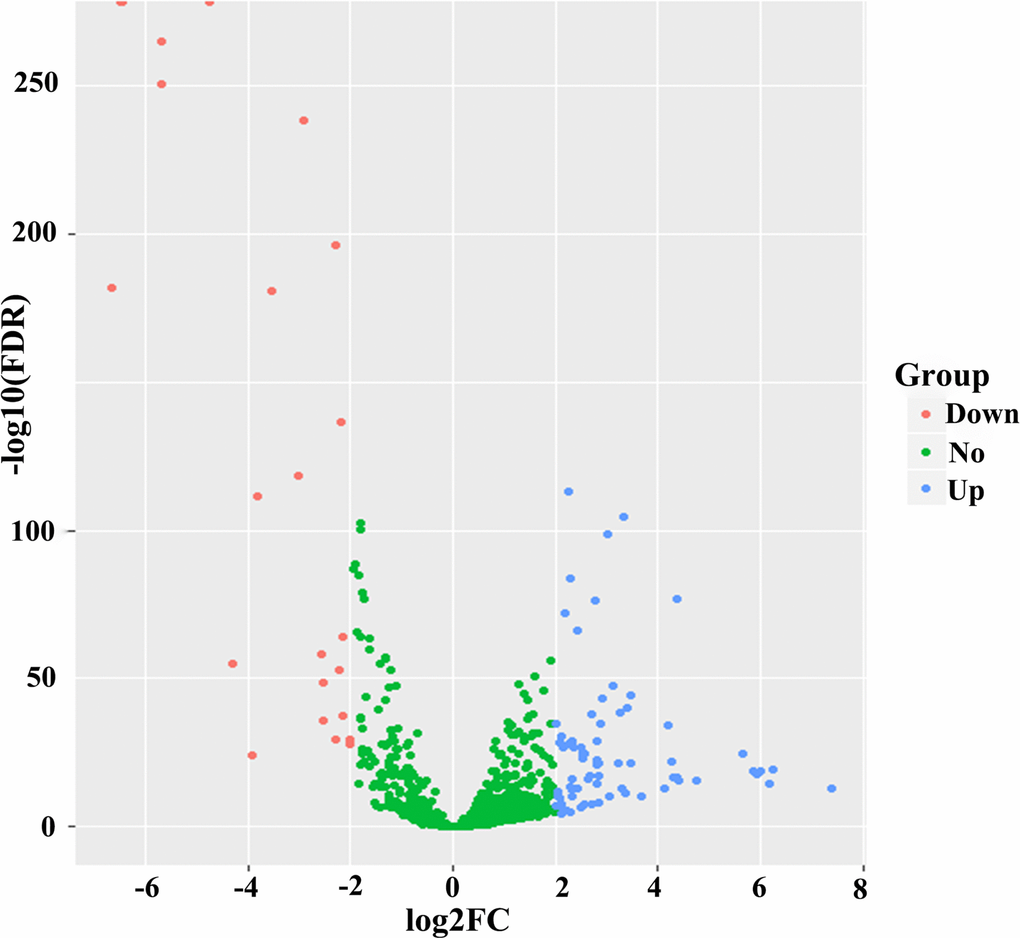
The Mann-Whitney U test and the χ2 test were implemented to compare the associations of continuous and categorical variables between the primary cohort and validation cohort, respectively. Univariate and multivariate CPHR analyses were executed to screen the independent prognostic variables of OS (P <0.05). Then, we used the Cox regression coefficients to establish a risk score formula and miRNA-based nomogram. For survival analyses, Kaplan-Meier method was carried out to plot survival curves, which were compared using log-rank tests. The predictive accuracy of each variable was tested via time-dependent ROC analysis. Time-dependent ROC curve analysis is extensively applied in biomedical reports for assessing the predictive accuracy of the six-miRNA signature. It is a graphical display which plots sensitivity estimates (probability of a true positive) against one minus specificity (probability of a false positive) of the six-miRNA signature for all possible threshold values. In a time-dependent ROC analysis, the sensitivity and specificity are determined at each time point to guide important medical decisions [34]. A volcano plot and heat map were drawn using the “ggplot2” package of R software. The primary end point was OS, which was computed the interval from surgery to the date of death from any cause. A value of P<0.05 was determined statistically significant. All statistical analyses were conducted with Stata/MP, version 14.0 (StataCorp LP, College Station, TX) and R version 3.4.4 were applied to the statistical analyses.
Author Contributions
FXS, JGL, and ZHP made substantial contribution to conception and design, analysis and interpretation of data, wrote and revised the manuscript. HLW collected and analyzed the data, and also took part in the drafting of the manuscript. All authors contributed toward data analysis, drafting and revising the paper and agree to be accountable for all aspects of the work. All authors have read and approved the final version of the manuscript.
Acknowledgements
This work benefited from the Cancer Genome Atlas (TCGA) database. We were grateful to the access to the resources and the efforts of the staff to expand and improve the databases.
Conflicts of Interest
The authors declare that there is no conflict of interest to disclose.
Funding
This study was supported by the National Natural Science Foundation of China (grants 81402201 and 81372817), National Natural Science Foundation of Guangdong Province (grant 2014A030310070), and grant 163 (2013) from the Key Laboratory of Malignant Tumor Molecular Mechanism and Translational Medicine of the Guangzhou Bureau of Science and Information Technology.
References
- 1. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015.CA: a cancer journal for clinicians.2016;66:115-132. https://doi.org/10.3322/caac.21338 [PubMed]
- 2. Fan L, Strasser-Weippl K, Li J-J, St Louis J, Finkelstein DM, Yu K-D, Chen W-Q, Shao Z-M, Goss PE. Breast cancer in China.The Lancet Oncology.2014;15:e279-e289. https://doi.org/10.1016/S1470-2045(13)70567-9 [PubMed]
- 3. Balachandran VP, Gonen M, Smith JJ, DeMatteo RP. Nomograms in oncology: more than meets the eye.The Lancet Oncology.2015;16:e173-e180. https://doi.org/10.1016/S1470-2045(14)71116-7 [PubMed]
- 4. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function.Cell.2004;116:281-297. https://doi.org/10.1016/S0092-8674(04)00045-5 [PubMed]
- 5. Harfe BD. MicroRNAs in vertebrate development.Current opinion in genetics & development.2005;15:410-415. https://doi.org/10.1016/j.gde.2005.06.012 [PubMed]
- 6. Jones KB, Salah Z, Del Mare S, Galasso M, Gaudio E, Nuovo GJ, Lovat F, LeBlanc K, Palatini J, Randall RL, Volinia S, Stein GS, Croce CM, et al. miRNA signatures associate with pathogenesis and progression of osteosarcoma.Cancer Res.2012;72:1865-1877. https://doi.org/10.1158/0008-5472.CAN-11-2663 [PubMed]
- 7. Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE, Iorio MV, Visone R, Sever NI, Fabbri M, Iuliano R, Palumbo T, Pichiorri F, et al. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia.N Engl J Med.2005;353:1793-1801. https://doi.org/10.1056/NEJMoa050995 [PubMed]
- 8. Zhang J-X, Song W, Chen Z-H, Wei J-H, Liao Y-J, Lei J, Hu M, Chen G-Z, Liao B, Lu J, Zhao H-W, Chen W, He Y-L, et al. Prognostic and predictive value of a microRNA signature in stage II colon cancer: a microRNA expression analysis.The Lancet Oncology.2013;14:1295-1306. https://doi.org/10.1016/S1470-2045(13)70491-1 [PubMed]
- 9. Tang X-R, Li Y-Q, Liang S-B, Jiang W, Liu F, Ge W-X, Tang L-L, Mao Y-P, He Q-M, Yang X-J, Zhang Y, Wen X, Zhang J, et al. Development and validation of a gene expression-based signature to predict distant metastasis in locoregionally advanced nasopharyngeal carcinoma: a retrospective, multicentre, cohort study.The Lancet Oncology.2018; https://doi.org/10.1016/S1470-2045(18)30080-9 [PubMed]
- 10. Li H, Liu J, Chen J, Wang H, Yang L, Chen F, Fan S, Wang J, Shao B, Yin D, Zeng M, Li M, Li J, et al. A serum microRNA signature predicts trastuzumab benefit in HER2-positive metastatic breast cancer patients.Nature communications.2018;9:1614 https://doi.org/10.1038/s41467-018-03537-w [PubMed]
- 11. Ji D, Qiao M, Yao Y, Li M, Chen H, Dong Q, Jia J, Cui X, Li Z, Xia J, Gu J. Serum-based microRNA signature predicts relapse and therapeutic outcome of adjuvant chemotherapy in colorectal cancer patients.EBioMedicine.2018; https://doi.org/10.1016/j.ebiom.2018.08.042 [PubMed]
- 12. Bagnoli M, Canevari S, Califano D, Losito S, Maio MD, Raspagliesi F, Carcangiu ML, Toffoli G, Cecchin E, Sorio R, Canzonieri V, Russo D, Scognamiglio G, et al. Development and validation of a microRNA-based signature (MiROvaR) to predict early relapse or progression of epithelial ovarian cancer: a cohort study.The Lancet Oncology.2016;17:1137-1146. https://doi.org/10.1016/S1470-2045(16)30108-5 [PubMed]
- 13. Zhang MY, Li SH, Huang GL, Lin GH, Shuang ZY, Lao XM, Xu L, Lin XJ, Wang HY, Li SP. Identification of a novel microRNA signature associated with intrahepatic cholangiocarcinoma (ICC) patient prognosis.BMC Cancer.2015;15:64 https://doi.org/10.1186/s12885-015-1067-6 [PubMed]
- 14. Shu X, Hildebrandt MA, Gu J, Tannir NM, Matin SF, Karam JA, Wood CG, Wu X. MicroRNA profiling in clear cell renal cell carcinoma tissues potentially links tumorigenesis and recurrence with obesity.British journal of cancer.2017;116:77-84. https://doi.org/10.1038/bjc.2016.392 [PubMed]
- 15. Sana J, Radova L, Lakomy R, Kren L, Fadrus P, Smrcka M, Besse A, Nekvindova J, Hermanova M, Jancalek R, Svoboda M, Hajduch M, Slampa P, et al. Risk Score based on microRNA expression signature is independent prognostic classifier of glioblastoma patients.Carcinogenesis.2014;35:2756-2762. https://doi.org/10.1093/carcin/bgu212 [PubMed]
- 16. Li X, Shi Y, Yin Z, Xue X, Zhou B. An eight-miRNA signature as a potential biomarker for predicting survival in lung adenocarcinoma.Journal of translational medicine.2014;12:159 https://doi.org/10.1186/1479-5876-12-159 [PubMed]
- 17. Han ZB, Zhong L, Teng MJ, Fan JW, Tang HM, Wu JY, Chen HY, Wang ZW, Qiu GQ, Peng ZH. Identification of recurrence-related microRNAs in hepatocellular carcinoma following liver transplantation.Molecular oncology.2012;6:445-457. https://doi.org/10.1016/j.molonc.2012.04.001 [PubMed]
- 18. Becerra C, Kopetz S, Wang X, Goel A, Chen L. Prediction of radiotherapy response with a 5-microRNA signature-based nomogram in head and neck squamous cell carcinoma.Cancer medicine.2018;7:726-735. https://doi.org/10.1002/cam4.1369 [PubMed]
- 19. Li X, Zhang Y, Zhang Y, Ding J, Wu K, Fan D. Survival prediction of gastric cancer by a seven-microRNA signature.Gut.2010;59:579-585. https://doi.org/10.1136/gut.2008.175497 [PubMed]
- 20. Gattolliat CH, Le Teuff G, Combaret V, Mussard E, Valteau-Couanet D, Busson P, Benard J, Douc-Rasy S. Expression of two parental imprinted miRNAs improves the risk stratification of neuroblastoma patients.Cancer medicine.2014;3:998-1009. https://doi.org/10.1002/cam4.264 [PubMed]
- 21. Iasonos A, Schrag D, Raj GV, Panageas KS. How to build and interpret a nomogram for cancer prognosis.Journal of clinical oncology: official journal of the American Society of Clinical Oncology.2008;26:1364-1370. https://doi.org/10.1200/JCO.2007.12.9791 [PubMed]
- 22. Krishnan P, Ghosh S, Wang B, Li D, Narasimhan A, Berendt R, Graham K, Mackey JR, Kovalchuk O, Damaraju S. Next generation sequencing profiling identifies miR-574-3p and miR-660-5p as potential novel prognostic markers for breast cancer.BMC genomics.2015;16:735 https://doi.org/10.1186/s12864-015-1899-0 [PubMed]
- 23. Kawaguchi T, Yan L, Qi Q, Peng X, Edge SB, Young J, Yao S, Liu S, Otsuji E, Takabe K. Novel MicroRNA-Based Risk Score Identified by Integrated Analyses to Predict Metastasis and Poor Prognosis in Breast Cancer.Ann Surg Oncol.2018; https://doi.org/10.1245/s10434-018-6859-x [PubMed]
- 24. Gong C, Tan W, Chen K, You N, Zhu S, Liang G, Xie X, Li Q, Zeng Y, Ouyang N, Li Z, Zeng M, Zhuang S, et al. Prognostic Value of a BCSC-associated MicroRNA Signature in Hormone Receptor-Positive HER2-Negative Breast Cancer.EBioMedicine.2016;11:199-209. https://doi.org/10.1016/j.ebiom.2016.08.016 [PubMed]
- 25. Zhu R, Lin W, Zhao W, Fan F, Tang L, Hu Y. A 4-microRNA signature for survival prognosis in pediatric and adolescent acute myeloid leukemia.Journal of cellular biochemistry.2018; https://doi.org/10.1016/S1470-2045(13)70491-1 [PubMed]
- 26. Zhao Y, Schetter AJ, Yang GB, Nguyen G, Mathe EA, Li P, Cai H, Yu L, Liu F, Hang D, Yang H, Wang XW, Ke Y, et al. microRNA and inflammatory gene expression as prognostic marker for overall survival in esophageal squamous cell carcinoma.International journal of cancer.2013;132:2901-2909. https://doi.org/10.1002/ijc.27954 [PubMed]
- 27. Yang Y, Qu A, Zhao R, Hua M, Zhang X, Dong Z, Zheng G, Pan H, Wang H, Yang X, Zhang Y. Genome-wide identification of a novel miRNA-based signature to predict recurrence in patients with gastric cancer.Molecular oncology.2018; https://doi.org/10.1002/1878-0261.12385 [PubMed]
- 28. Sparano JA, Gray RJ, Makower DF, Pritchard KI, Albain KS, Hayes DF, Geyer CE Jr, Dees EC, Perez EA, Olson JA Jr, Zujewski J, Lively T, Badve SS, et al. Prospective Validation of a 21-Gene Expression Assay in Breast Cancer.N Engl J Med.2015;373:2005-2014. https://doi.org/10.1056/NEJMoa1510764 [PubMed]
- 29. Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data.Bioinformatics (Oxford, England).2010;26:139-140. https://doi.org/10.1093/bioinformatics/btp616 [PubMed]
- 30. Heagerty PJ, Lumley T, Pepe MS. Time-dependent ROC curves for censored survival data and a diagnostic marker.Biometrics.2000;56:337-344. https://doi.org/10.1111/j.0006-341X.2000.00337.x [PubMed]
- 31. Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs.eLife.2015;4 https://doi.org/10.7554/eLife.05005 [PubMed]
- 32. Wong N and Wang X. miRDB: an online resource for microRNA target prediction and functional annotations.Nucleic acids research.2015;43:D146-152. https://doi.org/10.1093/nar/gku1104 [PubMed]
- 33. Chou CH, Shrestha S, Yang CD, Chang NW, Lin YL, Liao KW, Huang WC, Sun TH, Tu SJ, Lee WH, Chiew MY, Tai CS, Wei TY, et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions.Nucleic acids research.2018;46:D296-d302. https://doi.org/10.1093/nar/gkx1067 [PubMed]
- 34. Kamarudin AN, Cox T, Kolamunnage-Dona R. Time-dependent ROC curve analysis in medical research: current methods and applications.BMC medical research methodology.2017;17:53 https://doi.org/10.1186/s12874-017-0332-6 [PubMed]