Introduction
In studies of aging, changes in the length of life are usually analyzed by comparing average (mean) or median longevity. Frequently, some estimate of maximal longevity is also considered. While values of the standard deviations or standard errors of the mean are routinely reported, the distribution of individual age at death is rarely analyzed or discussed. This contrasts with the recent interest in analyzing the distribution of biomarkers of aging using the statistical distance measure [1] to estimate the level of physiological dysregulation and to relate it to resilience and robustness during aging [2].
In a recent publication based on analysis of demographic data, Van Raalte and her colleagues reported that socio-economic status influences not only the mean longevity but also the variability of human life-span [3]. Using an example of Finnish women, these investigators showed that reduced mean longevity of less educated and less affluent people is associated with greater variability of life-span. The inverse relationship of life expectancy and life-span variation was seen also in other human populations [3]. The practical implication of these findings is that for both the individuals and the health care systems it is more difficult to predict the age at death of the less privileged people than of the more privileged strata or for the entire population.
Because of the potential significance of this relationship for the analysis of mortality data and physiological biomarkers in studies of aging as well as for various public health considerations, we thought that it would be of interest to determine whether interventions known to extend the average (or the average and the maximal) longevity of experimental animals have any effect on the variability of life-span. We hypothesized that extension of longevity by genetic, dietary or pharmacological means leads to reduction of life-span variability. However, inspection of data from the National Institute of Aging Interventions Testing Program (ITP) [4] and from our studies of the interactions of murine longevity genes with calorie restriction (CR) [5–7] indicated that reciprocal changes of longevity and its variability are not consistently observed. This suggested that our hypothesis would most likely need to be rejected and brought up a new question, namely, what factors influence variability of the lifespan. Here we report results of a study aimed at analyzing the effects of sex, strain, life-extending interventions and their interactions on life-span variation.
Results
When the longevity of the mice in the ITP dataset was correlated to the variance in longevity, a strong decrease in variance with increasing longevity was observed. Skewness also decreased, while kurtosis, i.e., non-flatness, increased (Table 1). However, when gender was included in these relationships, we found that it strongly affected the results (Fig. 1; Table 2).
Table 1. Estimates of the regression coefficients of the fitted line of longevity against variance, skewness and kurtosis.
| Estimate | SE | t-value | df | 𝛘2 (LRT) | p-value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Variance (MAD) | -0.425 | 0.047 | -9.125 | 1 | 71.57 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Skewness | -0.442 | 0.047 | -9.472 | 1 | 79.98 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kurtosis | 0.325 | 0.049 | 6.681 | 1 | 42.19 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| In each test group, longevity is measured as the median age at death, variance as the Median Absolute Deviation (MAD) of age at death, skewness as the skewness of age at death, and kurtosis as the kurtosis of age at death. Estimate, standard error (SE) and t-value are from linear mixed models with research site as random effect variable. 𝛘2 and p-value are from a Likelihood Ratio Test (LRT) where the model with the independent variable is compared to the model without that variable. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
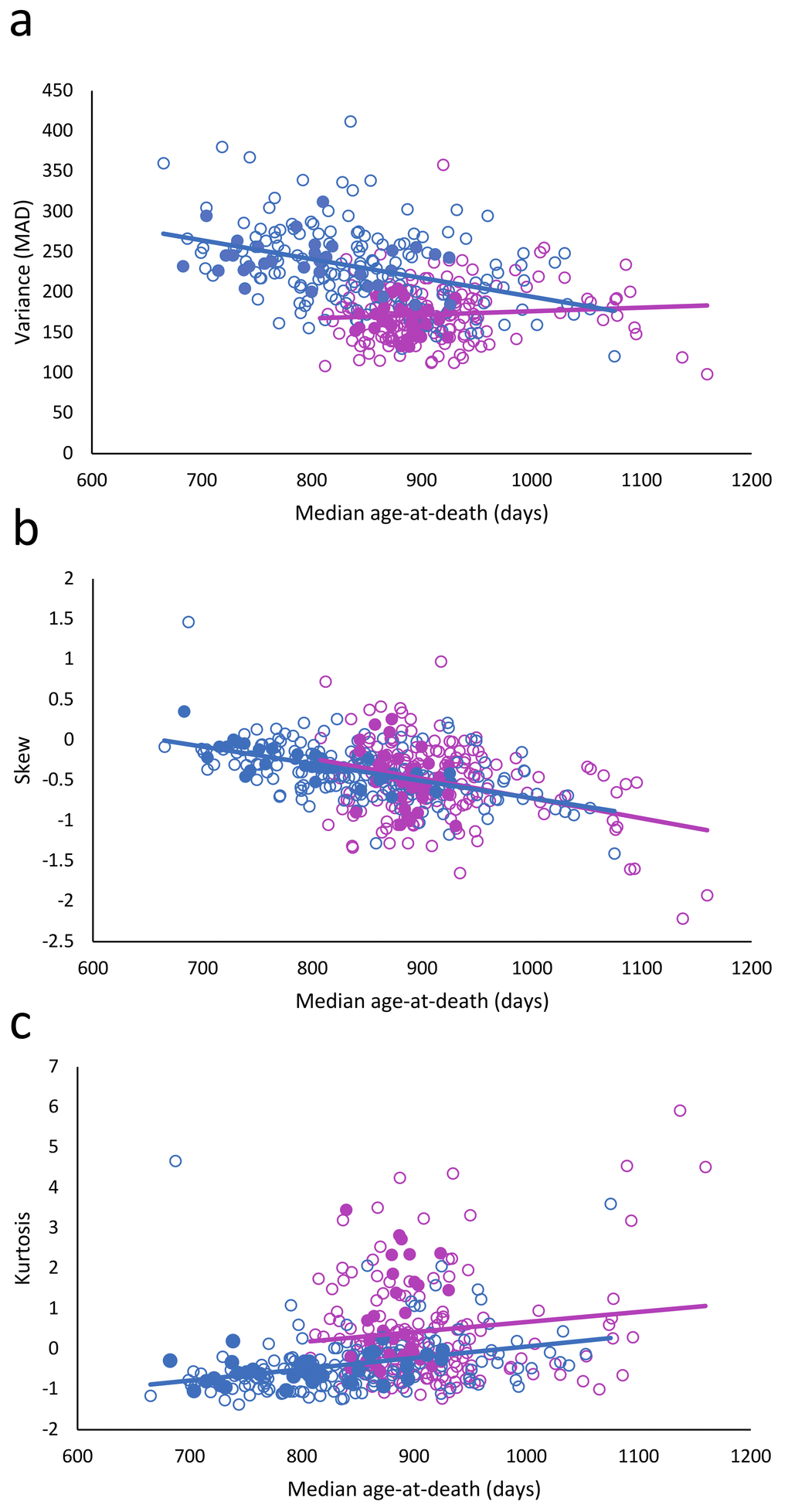
Figure 1. Relationship between longevity and variance (a), skewness (b), and kurtosis (c). Females: red circles; males: blue circles; filled circles: treated; and unfilled circles; untreated.
Table 2. Estimates of contrasts of models of longevity against variance, skewness and kurtosis, including gender and the interaction between gender and independent variables.
| Estimate | SE | t-value | df | 𝛘2 (LRT) | p-value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Variance (MAD) | Intercept (F) | -0.623 | 0.067 | -9.288 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender (M) | 1.035 | 0.089 | 11.659 | 1 | 115.83 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age | 0.071 | 0.072 | 0.978 | 1 | 0.95 | 0.329 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Interaction Gender x Median | -0.437 | 0.091 | -4.782 | 1 | 20.47 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Skewness | Intercept (F) | 0.068 | 0.083 | 0.822 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender (M) | -0.106 | 0.104 | -1.024 | 1 | 1.03 | 0.310 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age | -0.505 | 0.084 | -5.986 | 1 | 33.74 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Interaction Gender x Median | 0.064 | 0.107 | 0.591 | 1 | 0.35 | 0.556 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kurtosis | Intercept (F) | 0.299 | 0.077 | 3.887 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender (M) | -0.579 | 0.105 | -5.505 | 1 | 29.16 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age | 0.182 | 0.085 | 2.129 | 1 | 4.51 | 0.034 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Interaction Gender x Median | 0.023 | 0.108 | 0.216 | 1 | 0.05 | 0.829 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Estimate, standard error and t-value are from linear mixed models with research site as random effect variable. 𝛘2 and p-value are from a Likelihood Ratio Test (LRT) where the model with the independent variable is compared to the model without that variable. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The most striking finding was that the variance of longevity is strongly sexually dimorphic (Fig. 1) (Table 2: 𝛘2 = 20.47; df = 1; p < 0.001). In females the variance did not decrease with the increased longevity (Table 2: 𝛘2 = 0.95; df = 1; p =0.329), while that in males did (𝛘2 = 20.47; df = 1; p < 0.001). The skewness decreased (𝛘2 = 33.74; df = 1; P < 0.001), equally in both female and males (𝛘2 = 0.35; df = 1; p = 0.556). Kurtosis increased slightly (𝛘2 = 4.51; df = 1; p = 0.034), but was significantly lower in males than females (𝛘2 = 29.16; df = 1; p < 0.001).
A graphic representation of the variance in longevity of control mice vs mice treated with effective anti-aging compounds showed that, apart from the effect of the compounds on median longevity, they mainly seemed to decrease early death in males (Fig. 2).
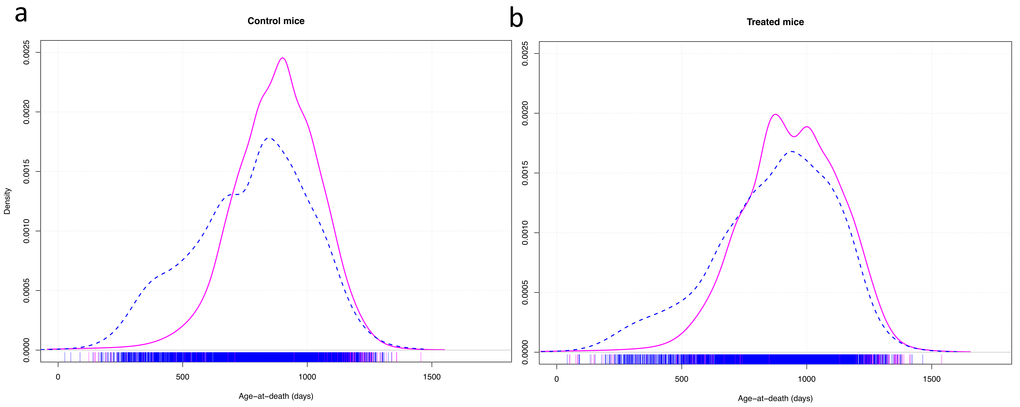
Figure 2. Density plots of the age at death of control mice (a) and mice treated with effective anti-aging pharmaceuticals (b). Cox-test for difference between females (solid red line) and males (dashed blue line) in survival with research site as random effect variable: control mice: z = 11.02, p < 0.001; treated mice: z = 6.95, p < 0.001.
The dataset of the longevity genes/CR studies showed no overall correlation between the median longevity per test group and the variance in longevity, measured as MAD per test group (𝛘2 = 0.25; df = 1; p = 0.617). When gender was taken into consideration and the studies were split according to effect of diet, genotype, or diet plus genotype, it was found that when mice had longevity extending genotypes and were treated with CR, variance of females increased near statistical significance (𝛘2 = 3.25; df = 1; p = 0.071), while in males this seemed not the case (Figure 3; Table 3).
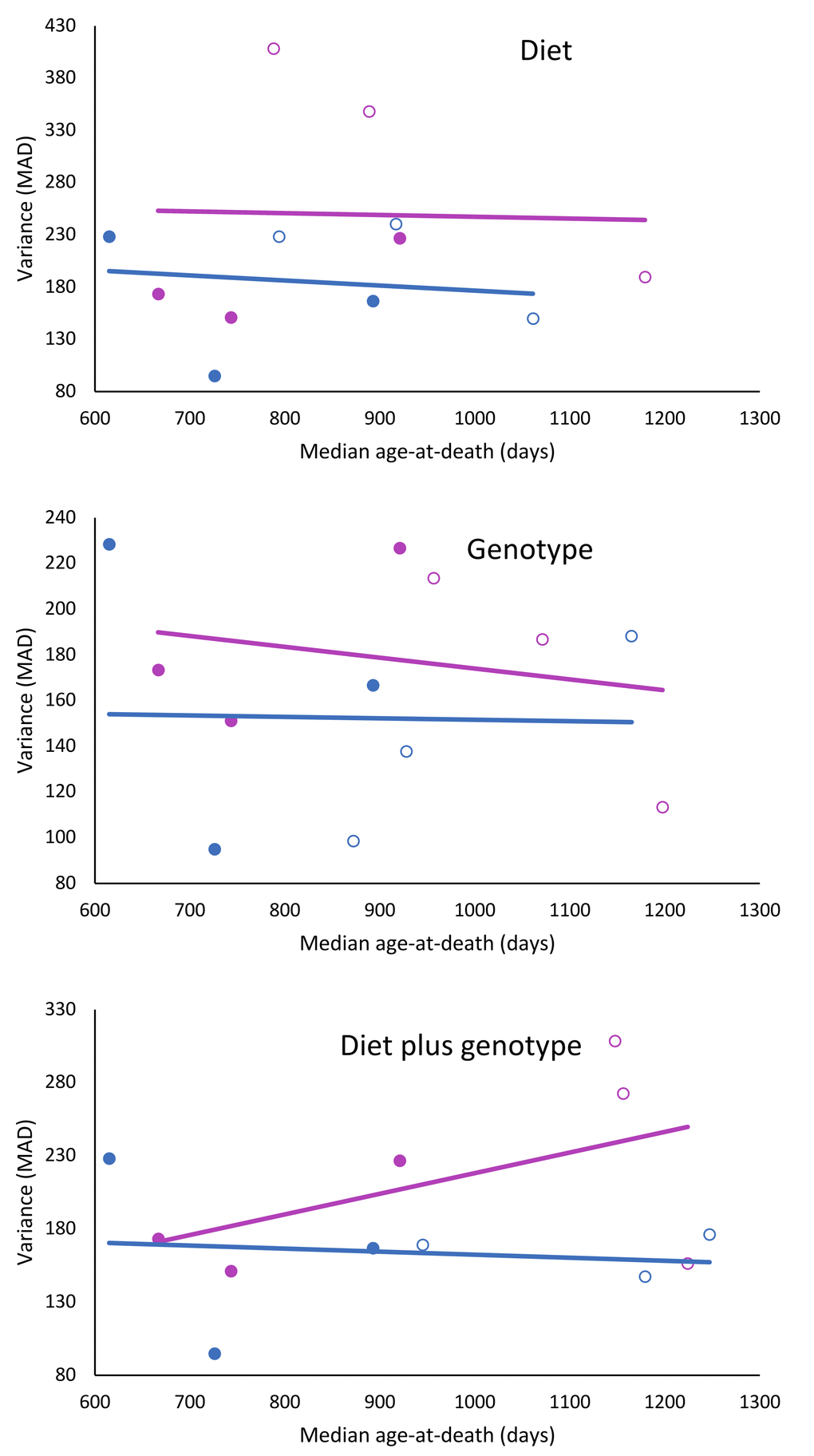
Figure 3. Effects of CR, life extending mutations or both on the relationship to longevity and variance in female and male mice. Females: red circles; males: blue circles; filled circles: treated; and unfilled circles; untreated.
Table 3. Estimates of contrasts of models of longevity with variance including gender and the interaction between gender and independent variables; CR, life extending mutations, or both.
| Estimate | SE | t-value | df | 𝛘2 (LRT) | p-value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diet | Intercept (F) | 0.375 | 0.442 | 0.850 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender (M) | -0.756 | 0.625 | -1.210 | 1 | 2.02 | 0.156 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age | -0.032 | 0.433 | -0.074 | 1 | 0.01 | 0.928 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Interaction Gender x Median | -0.058 | 0.659 | -0.088 | 0.01 | 0.914 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Genotype | Intercept (F) | 0.298 | 0.494 | 0.603 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender (M) | -0.576 | 0.616 | -0.936 | 1 | 1.16 | 0.282 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age | -0.192 | 0.457 | -0.420 | 1 | 0.25 | 0.616 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Interaction Gender x Median | 0.124 | 0.644 | 0.192 | 1 | 0.07 | 0.798 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diet+Gen. | Intercept (F) | 0.379 | 0.450 | 0.843 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gender (M) | -0.810 | 0.454 | -1.787 | 1 | 3.83 | 0.050 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age | 0.567 | 0.347 | 1.636 | 1 | 3.25 | 0.071 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Interaction Gender x Median | -0.573 | 0.477 | -1.202 | 1 | 1.97 | 0.161 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Estimate, standard error and t-value are from linear mixed models with strain as random effect variable. 𝛘2 and p-value are from a Likelihood Ratio Test (LRT) where the model with the independent variable is compared to the model without that variable. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The graphic representation of the variance in longevity of control mice vs long-lived mutant mice treated with CR showed that, apart from the life extending effect of the treatment on median longevity, the gender difference in longevity was due predominantly to some males dying relatively early (Fig. 4).
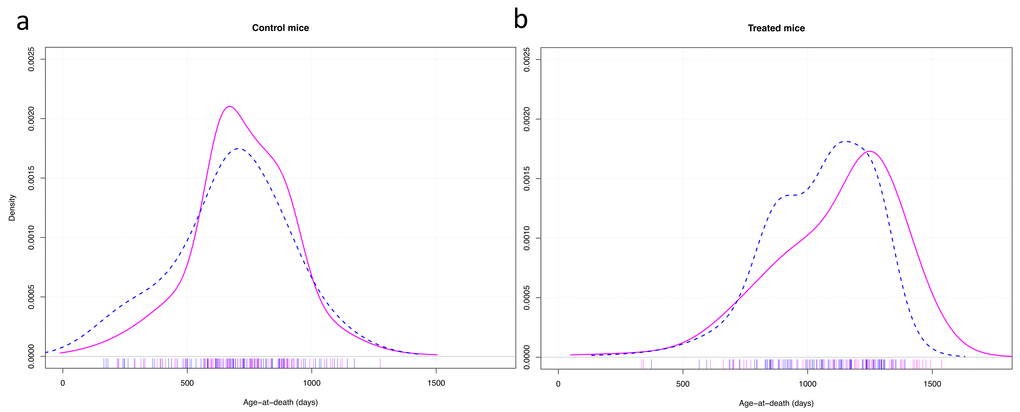
Figure 4. Density plots of the age of death of control mice (a) and mice treated with anti-aging diet as well as having anti-aging genotypes (b). Cox-test for difference between females (solid red) and males (dashed blue) in survival with strain as random effect variable: control mice: z = 0.57, p = 0.57; treated mice: z = 3.97, p < 0.001.
Discussion
Results of the present study indicate that interventions which extend longevity in laboratory mice can also alter the distribution and the variance of age at death. The presence, the magnitude, and apparently also the direction of these changes depend on the sex of the animals and the nature of the interventions. In the large data set from the ITP studies, increased longevity was associated with reduced variance in males (p<0.001) but not in females. Moreover, distribution of the data suggested a possible trend for variance in female data changing in the opposite direction (Fig 1). In long-lived mice with GH-related mutations, subjected to CR, variance was not related to longevity in males, but seemed to increase with longevity in females (p=0.071).
Sexual dimorphism in the responses to longevity-extending interventions is not unexpected and our findings show that the relationship of the distribution of individual age at death data to the average life span of the population is also sex-dependent. Thus, the present results add to the growing body of evidence that various aspects of aging, including longevity, as well as responses to various anti-aging interventions cannot be predicted from data obtained in individuals of different sex [8].
The reciprocal relationship of human longevity and its variablitiy that Van Raalte, et al. [3] found in various data sets were from segments of the same population separated on the basis of income, educational attainment, and/or type of employment. The biological basis of the well-documented impact of socio-economic factors on human aging is believed to include differences in diet, health-related behaviors, and access to as well as quality of health care, although significance of the latter factors in Finland and other countries with excellent health care systems is presumably small or absent.
Since animals in each of the four examined cohorts were genetically heterogeneous and three of the four cohorts were relatively small, this study does not allow any firm conclusion on the possible differences between strains. Studies including large genetically distinct populations, different inbred strains and preferably also their crosses would be needed to address this issue.
What can be concluded from our analysis is that in studies of anti-aging interventions in laboratory animals the changes in variability of lifespan cannot be predicted from changes in longevity. It is interesting to speculate that this may be related to differences in the mechanisms responsible for lifespan extension by the various interventions.
In the data sets we analyzed, there were examples of differences in the proportion of early deaths in females and males as well as in long-lived (mutant, calorie restricted or drug-treated) vs control (wild type, fed ad libitum, or untreated) animals and the slopes of survival curves also differed in some cases. Skewness and kurtosis were as expected. The differences in the distribution of individual lifespans may have contributed to the observed differences in variation. For example, decrease in the occurrence of early deaths likely contributed to longevity of drug-treated males in the ITP studies [9]. This and other examples of the role of early deaths in determining the longevity of the whole cohort resemble demographic findings on the effects of socio-economic factors in human populations [3].
Analysis of distribution of individual age at death values suggests a differential impact of the examined anti-aging interventions at different segments of life history. This, in turn, is likely related to different mechanisms of their actions. Significant correlation of extended longevity with reduced variance of individual death in various human populations [3] and in male mice from the ITP cohort (Fig 1), may have been due, in part, to the “ceiling effect” as longevity approaches the value of maximal lifespan for the species. More likely, it reflects greater proportion of early deaths of various etiologies in groups with shorter median lifespan. Intriguingly, trajectories of several physiological processes mechanistically related to aging are more variable in short-lived than in exceptionally long-lived people [10].
Regardless of the mechanisms involved, our data suggest that analysis of changes in the variability of life-spans in response to various interventions can provide additional information on the nature of the observed effects.
Materials and Methods
We used data on the length of life of individual mice (age at death measured in days) from the ITP [4] and from studies conducted at Southern Illinois University (SIU) [5–7]. In the ITP, genetically heterogeneous (UM-HET3) female and male mice derived from crosses of four inbred strains were treated with various compounds starting in early adulthood and continuing until they died or were euthanized when judged to be moribund. These studies were conducted in parallel at three collaborating institutions. Both significant extension of longevity and absence of effects were reported in peer-reviewed literature [9,11–15]. A listing of the compounds, their actions and the percent increase in median longevity of treated animals are listed in Table 4 along with references to publications reporting longevity data.
Table 4. Percent increase in median lifespan due to pharmacological agents administered in the Interventions Testing Program.
| Drug | Effects, mode of action | Δ Median lifespan | Citation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Females | Males | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Acarbose | Inhibitor of intestinal alpha-glocosidase | 5 | 22 | Harrison et al., 2014 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Aspirin | anti-inflammatory, antithrombotic and antioxidant | ns | 8 | Stong et al., 2008 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 17 alpha-estradiol | neuroprotection | ns | 12 | Harrison et al., 2014 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Dietary glycine | anti-inflammatory | 4 | 1 | Brind et al., 2018 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NDGA | Antioxidant, anti-inflammatory | ns | 8-10 | Harrison et al., 2014 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protandim (mixture of plant extracts) | Nrf activation; increase of antioxidant defenses | ns | 7 | Strong et al., 2016 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Rapamycin | Inhibitor of mTORC1 | 18 | 10 | Miller et al., 2011 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Footnote: For some of the above listed drugs, additional studies were conducted using different doses and or starting treatment at a different age which resulted in somewhat different % extension of median life-span. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The effects of pharmacological anti-aging interventions in the ITP studies were analyzed using the data from treated mice with documented life extension and matching controls. These groups include animals treated with acarbose [9], aspirin [11], 17-α-estradiol [9], dietary glycine [15], nordihydroguaiaretic acid [9,11], ProtandimTM (mixture of plant extracts) [14], or rapamycin [16]. Longevity variance was examined in relation to treatment and sex while taking into account differences in testing laboratories [4].
A second dataset was created with individual longevity data of mice from three separate experiments on the interaction of calorie restriction (CR) with one of three life-extending mutations [5–7]. These experiments used mice with suppression of growth hormone (GH) signaling by a mutation interfering with differentiation of three types of anterior pituitary cells leading to hypopituitarism including GH deficiency [17,18], deletion of GH releasing hormone (GHRH) gene leading to isolated GH deficiency [19], or deletion of the GH receptor (GHR) gene leading to GH resistance [20]. Approximately half of the animals in each genotype/sex group were fed ad libitum (AL), while the remainder were subjected to calorie restriction (CR) starting in early adulthood by feeding them 60 -70% of the amount of food consumed during the preceding week by age, sex, and genotype matched AL mice. The employed mutations were maintained on a heterogeneous genetic background which was different for each mutation and different from the genetic make-up of the ITP mice. Each of the three mutations used in these studies increases longevity of both females and males while the effects of CR on longevity were mutation- and sex-dependent [5–7]. Percent increase of median longevity in each genotype/sex/diet group is listed in Table 5.
Table 5. Interactive effects of calorie restriction and genetic suppression of growth hormone signaling on median longevity in each genotype/sex/diet group.
| Strain | Diet | Female | Male | Action | Citation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ames N | CR | 20 | 26 | Hypopituitarism including GH deficiency | Bartke et al., 2001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ames dwarf | AL | 44 | 20 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ames dwarf | CR | 65 | 72 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GHR N | CR | 28 | 19 | deletion of the GH receptor (GHR) gene leading to GH resistance | Bonkowski et al., 2006 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GHR KO | AL | 30 | 31 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GHR KO | CR | 25 | 32 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GHRH N | CR | 18 | 29 | deletion of GH releasing hormone (GHRH) gene leading to isolated GH deficiency | Sun et al., 2013 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GHRH KO | AL | 44 | 51 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GHRH KO | CR | 74 | 54 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| There are 24 groups: three strains (Ames, GHR, GHRH); two sexes; two diets (CR and AL); and two genotypes (Normal and dwarf or knock-out). Normal (N) indicates normal siblings of mutant mice. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Longevity variance was examined in relation to different groups (genotype, diet, and sex combinations). Interventions that were ineffective (such as CR in GHR -/- mice) were included to account for the possibility that CR could alter variance without altering the median.
We performed all statistical analysis using R software 3.4.4 [21]. For studying the relationship between longevity and variance in longevity we calculated the median age at death of the females and males of all available test groups. We chose median instead of mean because we could not assume that the variance in longevity was normally distributed. For the variance in longevity we calculated the MAD, i.e., the Median Absolute Deviation, a robust metric fit for non-normal distributions [22]. Median, MAD, skewness, and kurtosis were all calculated with the function describeBy() of the package psych [23]. For testing the relationships between longevity and variance, skewness and kurtosis, the function lmer() of the package lme4 [24] for linear mixed-effect models were applied, combined with the drop1() function for the Likelihood Ratio Test (LRT). For testing the difference in survival between females and males, the Cox test for mixed effects was used, i.e., the function foxme() of the package foxme [25]. As random effect variable, the research site was included in all analyses of the ITP data and the strain in all analyses of the SIU (mutations/CR) data. The graphs were constructed using scatterplot() and densityPlot(), both of the package car [26]. We preferred density plots over Kaplan-Meier survival plots, because they show more clearly where the differences between females and males occurred.
Acknowledgments
We are grateful to Dr. Richard Miller for most helpful comments and suggestions and Dr. Scott Pletcher for his comments concerning interpretation of our findings. This work was supported by NIA via the ITP program and grants AG031736, AG019899, and AG038850, and by the American Diabetes Association, grant 1-19-IBS-126.
Conflicts of Interest
The authors declare no competing financial interests.
References
- 1. Cohen AA, Li Q, Milot E, Leroux M, Faucher S, Morissette-Thomas V, Legault V, Fried LP, Ferrucci L. Statistical distance as a measure of physiological dysregulation is largely robust to variation in its biomarker composition. PLoS One. 2015; 10:e0122541. https://doi.org/10.1371/journal.pone.0122541 [PubMed]
- 2. Arbeev KG, Ukraintseva SV, Bagley O, Zhbannikov IY, Cohen AA, Kulminski AM, Yashin AI. “Physiological Dysregulation” as a Promising Measure of Robustness and Resilience in Studies of Aging and a New Indicator of Preclinical Disease. J Gerontol A Biol Sci Med Sci. 2019; 74:462–68. https://doi.org/10.1093/gerona/gly136 [PubMed]
- 3. van Raalte AA, Sasson I, Martikainen P. The case for monitoring life-span inequality. Science. 2018; 362:1002–04. https://doi.org/10.1126/science.aau5811 [PubMed]
- 4. Miller RA, Harrison DE, Astle CM, Floyd RA, Flurkey K, Hensley KL, Javors MA, Leeuwenburgh C, Nelson JF, Ongini E, Nadon NL, Warner HR, Strong R. An Aging Interventions Testing Program: study design and interim report. Aging Cell. 2007; 6:565–75. https://doi.org/10.1111/j.1474-9726.2007.00311.x [PubMed]
- 5. Bartke A, Wright JC, Mattison JA, Ingram DK, Miller RA, Roth GS. Extending the lifespan of long-lived mice. Nature. 2001; 414:412. https://doi.org/10.1038/35106646 [PubMed]
- 6. Bonkowski MS, Rocha JS, Masternak MM, Al Regaiey KA, Bartke A. Targeted disruption of growth hormone receptor interferes with the beneficial actions of calorie restriction. Proc Natl Acad Sci USA. 2006; 103:7901–05. https://doi.org/10.1073/pnas.0600161103 [PubMed]
- 7. Sun LY, Spong A, Swindell WR, Fang Y, Hill C, Huber JA, Boehm JD, Westbrook R, Salvatori R, Bartke A. Growth hormone-releasing hormone disruption extends lifespan and regulates response to caloric restriction in mice. eLife. 2013; 2:e01098. https://doi.org/10.7554/eLife.01098 [PubMed]
- 8. Austad SN, Bartke A. Sex Differences in Longevity and in Responses to Anti-Aging Interventions: A Mini-Review. Gerontology. 2015; 62:40–46. https://doi.org/10.1159/000381472 [PubMed]
- 9. Harrison DE, Strong R, Allison DB, Ames BN, Astle CM, Atamna H, Fernandez E, Flurkey K, Javors MA, Nadon NL, Nelson JF, Pletcher S, Simpkins JW, et al. Acarbose, 17-α-estradiol, and nordihydroguaiaretic acid extend mouse lifespan preferentially in males. Aging Cell. 2014; 13:273–82. https://doi.org/10.1111/acel.12170 [PubMed]
- 10. Yashin AI, Arbeev KG, Wu D, Arbeeva LS, Kulminski A, Akushevich I, Culminskaya I, Stallard E, Ukraintseva SV. How lifespan associated genes modulate aging changes: lessons from analysis of longitudinal data. Front Genet. 2013; 4:3. https://doi.org/10.3389/fgene.2013.00003 [PubMed]
- 11. Strong R, Miller RA, Astle CM, Floyd RA, Flurkey K, Hensley KL, Javors MA, Leeuwenburgh C, Nelson JF, Ongini E, Nadon NL, Warner HR, Harrison DE. Nordihydroguaiaretic acid and aspirin increase lifespan of genetically heterogeneous male mice. Aging Cell. 2008; 7:641–50. https://doi.org/10.1111/j.1474-9726.2008.00414.x [PubMed]
- 12. Miller RA, Harrison DE, Astle CM, Baur JA, Boyd AR, de Cabo R, Fernandez E, Flurkey K, Javors MA, Nelson JF, Orihuela CJ, Pletcher S, Sharp ZD, et al. Rapamycin, but not resveratrol or simvastatin, extends life span of genetically heterogeneous mice. J Gerontol A Biol Sci Med Sci. 2011; 66:191–201. https://doi.org/10.1093/gerona/glq178 [PubMed]
- 13. Strong R, Miller RA, Astle CM, Baur JA, de Cabo R, Fernandez E, Guo W, Javors M, Kirkland JL, Nelson JF, Sinclair DA, Teter B, Williams D, et al. Evaluation of resveratrol, green tea extract, curcumin, oxaloacetic acid, and medium-chain triglyceride oil on life span of genetically heterogeneous mice. J Gerontol A Biol Sci Med Sci. 2013; 68:6–16. https://doi.org/10.1093/gerona/gls070 [PubMed]
- 14. Strong R, Miller RA, Antebi A, Astle CM, Bogue M, Denzel MS, Fernandez E, Flurkey K, Hamilton KL, Lamming DW, Javors MA, de Magalhães JP, Martinez PA, et al. Longer lifespan in male mice treated with a weakly estrogenic agonist, an antioxidant, an α-glucosidase inhibitor or a Nrf2-inducer. Aging Cell. 2016; 15:872–84. https://doi.org/10.1111/acel.12496 [PubMed]
- 15. Brind J, Miller RA, Strong R, Harrison DE, Macchiarini F. Dietary Glycine Supplementation Extends Lifespan of Genetically Heterogeneous Mice. FASEB J. 2018; 32:533.
- 16. Harrison DE, Strong R, Sharp ZD, Nelson JF, Astle CM, Flurkey K, Nadon NL, Wilkinson JE, Frenkel K, Carter CS, Pahor M, Javors MA, Fernandez E, Miller RA. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature. 2009; 460:392–95. https://doi.org/10.1038/nature08221 [PubMed]
- 17. Sornson MW, Wu W, Dasen JS, Flynn SE, Norman DJ, O’Connell SM, Gukovsky I, Carrière C, Ryan AK, Miller AP, Zuo L, Gleiberman AS, Andersen B, et al. Pituitary lineage determination by the Prophet of Pit-1 homeodomain factor defective in Ames dwarfism. Nature. 1996; 384:327–33. https://doi.org/10.1038/384327a0 [PubMed]
- 18. Bartke A. (2006). Life extension in the dwarf mouse. In: Conn PM, ed. Handbook of models for the study of human aging. Elsevier Academic Press, pp. 403-14.
- 19. Alba M, Fintini D, Salvatori R. Effects of recombinant mouse growth hormone treatment on growth and body composition in GHRH knock out mice. Growth Horm IGF Res. 2005; 15:275–82. https://doi.org/10.1016/j.ghir.2005.05.001 [PubMed]
- 20. Zhou Y, Xu BC, Maheshwari HG, He L, Reed M, Lozykowski M, Okada S, Cataldo L, Coschigamo K, Wagner TE, Baumann G, Kopchick JJ. A mammalian model for Laron syndrome produced by targeted disruption of the mouse growth hormone receptor/binding protein gene (the Laron mouse). Proc Natl Acad Sci USA. 1997; 94:13215–20. https://doi.org/10.1073/pnas.94.24.13215 [PubMed]
- 21. R Core Team. (2018). R: A language and environment for statistical computing. R Foundation for Statistical Computing. (Vienna, Austria: https://www.R-project.org/).
- 22. Leys C, Ley C, Klein O, Bernard P, Licata L. Detecting outliers: do not use standard deviation around the mean, use absolute deviation around the median. J Exp Soc Psychol. 2013; 49:764–66. https://doi.org/10.1016/j.jesp.2013.03.013
- 23. Revelle W. (2018). Procedures for Personality and Psychological Research. Northwestern University, Evanston, Illinois. https://CRAN.R-project.org/package=psych Version = 1.8.12.
- 24. Bates D, Machler M, Bolker BM, Walker SC. Fitting Linear Mixed-Effects Models Using lme4. J Stat Softw. 2015; 67:1–48. https://doi.org/10.18637/jss.v067.i01
- 25. Therneau T. (2018). Mixed Effects Cox Models. Mayo Clinic. http://cran.stat.nus.edu.sg/web/packages/coxme/vignettes/coxme.pdf.
- 26. Fox J, Weisberg S. (2011). An R Companion to Applied Regression. SAGE Publications.


