Introduction
DNA methylation, as the most common epigenetic modification, plays a crucial role in tissue- and stage-specific gene regulation, genomic imprinting, and X-chromosome inactivation, and has shown to be essential for normal mammalian development [1]. Also, accumulating studies have proved that both global DNA hypomethylation and hypermethylation occur frequently in tumorigenesis [2]. The hypermethylation of CpG islands at the promoter regions is often associated with the inactivation of tumor suppressor genes (TSGs) [3]. In hematopoietic disorders, aberrant DNA hypermethylation is proved to be involved in leukemogenesis [4]. For example, Spencer et al demonstrated that CpG island hypermethylation mediated by DNMT3A was a consequence of acute myeloid leukemia (AML) progression [5]. In addition, aberrant DNA methylation was also regarded as a dominant mechanism in progression from myelodysplastic syndromes (MDS) to AML [6]. Furthermore, deregulated DNA methylation in MDS and AML has led to the approval for the clinical use of hypomethylating agents (HMAs) in both MDS and AML patients [7].
DNA methyltransferases (DNMTs) are the main key effectors of DNA methylation by catalyzing the transfer of methyl groups from S-adenosyl-lmethionine to the 5’-position of cytosine residing in the dinucleotide sequence cytosine-guanine [8]. The DNMTs include three major members (DNMT1, DNMT3A and DNMT3B), among which DNMT3A and DNMT3B catalyze cytosine methylation of mammalian genomic DNA to establish de novo DNA methylation patterns, whereas DNMT1 maintains a methylation state through DNA replication [9]. Recent studies have demonstrated that DNMTs play vital roles in the progression of hematologic malignancies, especially AML [10]. Trowbridge et al showed that haploinsufficiency of DNMT1 impaired leukemia stem cell (LSC) function through derepression of bivalent chromatin domains [11]. More importantly, high incidence of DNMT3A mutation was identified in AML and DNMT3A mutation correlated with poor prognosis in AML [12]. Functional studies showed that DNMT3A was essential for hematopoietic stem cell (HSC) differentiation and mutated DNMT3A initiated AML [13–14], suggesting DNMT3A acted as a tumor suppressor gene. Although DNMT3B was rarely mutated in AML [15], studies have proved that loss of DNMT3B accelerated MLL-AF9 leukemia progression and increased expression of DNMT3B in LSC delayed leukemogenesis [16–17]. A few studies have shown the expression of DNMTs and their clinical significance in AML, but the results remain to be discussed [18–21]. Herein, we systemically analyzed DNMTs expression and their relationship with clinic-pathological features and prognosis in patients with AML.
Results
DNMTs expression associated with AML among human cancer cell lines
Using the Cancer Cell Line Encyclopedia (CCLE) databases, we found that expression of DNMTs was highly expressed in AML cell lines among 40 types of human cancer cell lines (Figure 1A). Moreover, DNMTs expression was also closely associated with myeloid cell lines revealed by The Human Protein Atlas (HPA) databases (Figure 1B). The detailed comparison of DNMTs expression in 14 types of AML cell lines was assessed by using the European Bioinformatics Institute (EMBL-EBI) website, which was shown in Figure 1C.
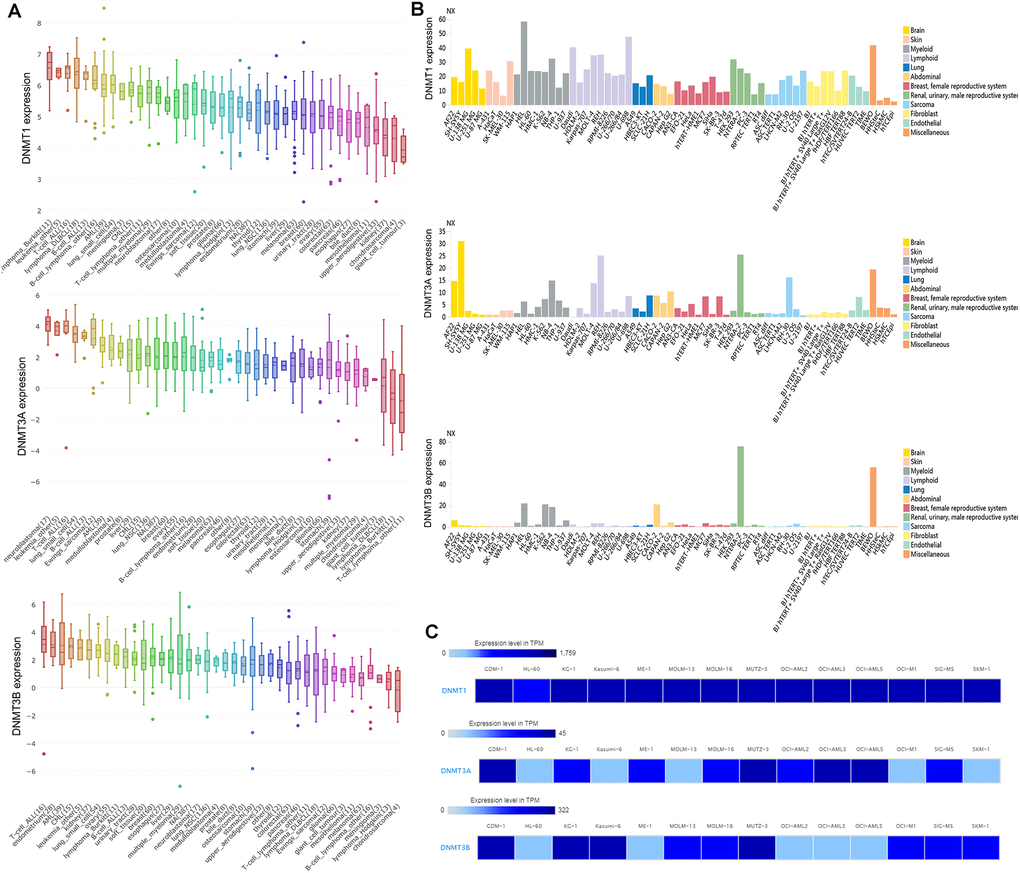
Figure 1. The expression of DNMTs in human cancer cell lines including AML cell lines. (A) The expression of DNMTs in leukemia cell lines, analyzing by Cancer Cell Line Encyclopedia (CCLE) dataset. (B) The expression of DNMTs in leukemia cell lines, analyzing by The Human Protein Atlas (HPA) dataset. (C) The expression of DNMTs in leukemia cell lines, analyzed by European Bioinformatics Institute (EMBL-EBI) dataset.
DNMTs expression associated with AML patients among human cancers
We further determined expression of DNMTs in AML patients by using Gene Expression Profiling Interactive Analysis (GEPIA) dataset. Aberrant expression of DNMT3A and DNMT3B was observed in AML among 33 types of human cancers, whereas DNMT1 did not show significant difference in AML (Figure 2A). DNMT3A expression was significantly increased in AML patients, whereas DNMT3B expression was markedly decreased in AML patients (Figure 2B). Moreover, DNMT1 expression was slightly associated with both DNMT3A and DNMT3B expression, whereas DNMT3A expression was positively correlated with DNMT3B expression in AML patients (Figure 2C).
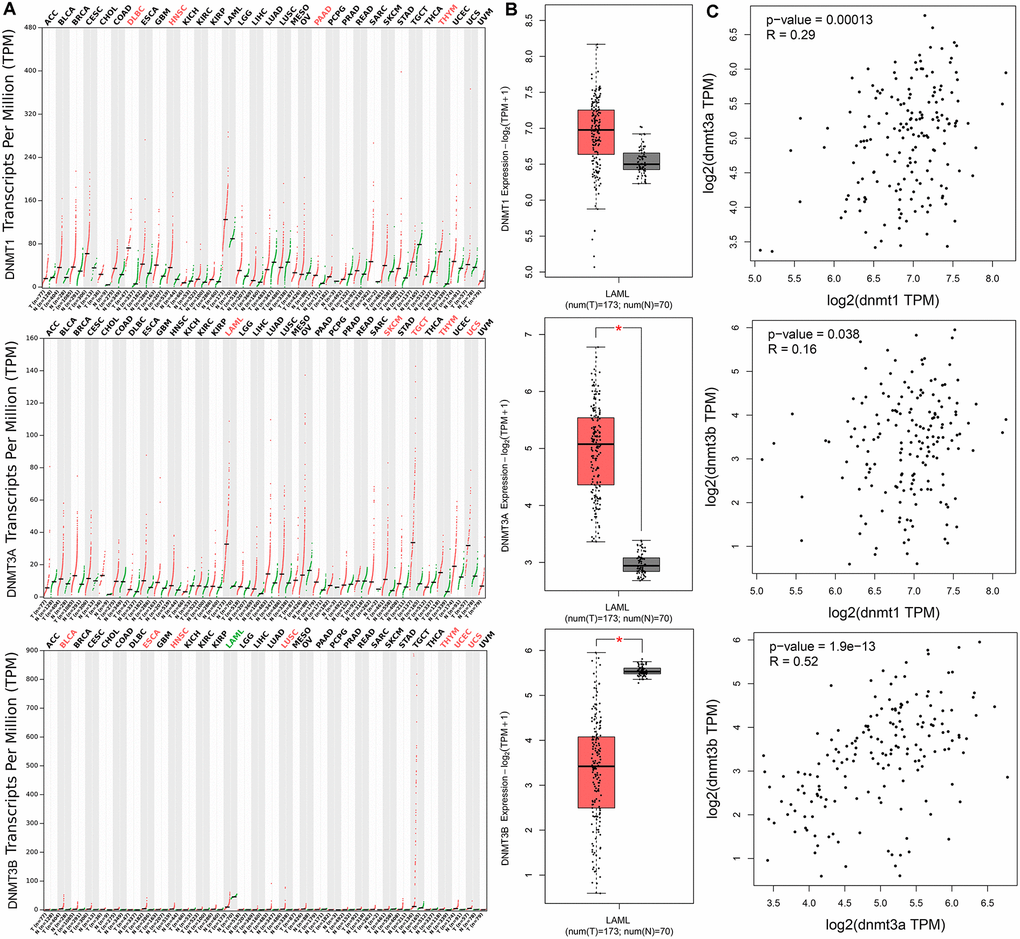
Figure 2. The expression of DNMTs in human cancers including AML patients. (A) The expression of DNMTs in pan-cancer analyzed by Gene Expression Profiling Interactive Analysis (GEPIA) web (http://gepia.cancer-pku.cn/). (B) The expression of DNMTs in AML analyzed by Gene Expression Profiling Interactive Analysis (GEPIA) web (http://gepia.cancer-pku.cn/). (C) The correction between DNMTs in AML analyzed by Gene Expression Profiling Interactive Analysis (GEPIA) web (http://gepia.cancer-pku.cn/).
Association between DNMT3A/3B expression and clinical characteristics in AML
Since aberrant expression of DNMT3A and DNMT3B was identified in AML, we further explore their clinical significance in patients with AML. Clinical implication of DNMT3A and DNMT3B was obtained by the comparison of clinical/laboratory characteristics of the AML patients between two groups divided based on median level of DNMT3A and DNMT3B transcript (Table 1). Interestingly, DNMT3A overexpression was associated younger age and lower white blood cells (WBCs) (P=0.008 and 0.063, respectively), higher peripheral blood (PB) blasts (P=0.006). Among the distribution of French-American-British (FAB) and cytogenetic subtypes, DNMT3A overexpression was associated with higher frequency of FAB-M0/M2, t(8;21), and -7/del(7) (P=0.009, 0.028, 0.007, and 0.007, respectively), whereas lower frequency of FAB-M4/M5 and normal karyotype (P=0.004, 0.001, and 0.000, respectively). In addition, DNMT3B underexpression was correlated with higher WBCs and lower PB blasts (P=0.041 and 0.006, respectively). Among the distribution of FAB and cytogenetic subtypes, DNMT3B underexpression was associated with higher frequency of FAB-M4/M5 and inv(16) (P=0.000, 0.000, and 0.001, respectively), but lower frequency of FAB-M1 and complex karyotype (P=0.000 and 0.000, respectively).
Table 1. Correlation of DNMT3A/B expression with clinic-pathologic characteristics in AML.
| Patient’s parameters | DNMT3A expression | DNMT3B expression | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Low (n=87) | High (n=86) | P | Low (n=87) | High (n=86) | P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sex, male/female | 47/40 | 45/41 | 0.879 | 47/40 | 45/41 | 0.879 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median age, years (range) | 60 (18-88) | 54 (21-81) | 0.008 | 58 (18-88) | 57 (21-81) | 0.922 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median WBC, ×109/L (range) | 25.9 (0.8-137.2) | 11.5 (0.4-297.4) | 0.063 | 22.2 (1-137.2) | 10.7 (0.4-297.4) | 0.041 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median PB blasts, % (range) | 17 (0-97) | 48 (0-98) | 0.006 | 25 (0-94) | 49 (0-98) | 0.006 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Median BM blasts, % (range) | 75 (30-98) | 72 (32-100) | 0.788 | 73 (30-99) | 72 (30-100) | 0.951 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FAB classifications | 0.000 | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M0 | 3 | 13 | 0.009 | 5 | 11 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M1 | 19 | 25 | 11 | 33 | 0.000 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M2 | 13 | 25 | 0.028 | 17 | 21 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M3 | 6 | 10 | 7 | 9 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M4 | 25 | 9 | 0.004 | 28 | 6 | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M5 | 16 | 2 | 0.001 | 17 | 1 | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M6 | 1 | 1 | 0 | 2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M7 | 3 | 0 | 1 | 2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| No data | 1 | 1 | 1 | 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cytogenetics | 0.000 | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| normal | 55 | 25 | 0.000 | 42 | 38 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| t(15;17) | 6 | 9 | 7 | 8 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| t(8;21) | 0 | 7 | 0.007 | 6 | 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| inv(16) | 2 | 8 | 10 | 0 | 0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| +8 | 4 | 4 | 3 | 5 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| del(5) | 1 | 0 | 0 | 1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| -7/del(7) | 0 | 7 | 0.007 | 2 | 5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 11q23 | 2 | 1 | 3 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| others | 5 | 9 | 9 | 5 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| complex | 11 | 14 | 4 | 21 | 0.000 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| No data | 1 | 2 | 1 | 2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene mutation | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FLT3 (+/-) | 28/59 | 21/65 | 0.312 | 22/65 | 27/59 | 0.402 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NPM1 (+/-) | 37/50 | 11/75 | 0.000 | 24/63 | 24/62 | 1.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DNMT3A (+/-) | 30/57 | 12/74 | 0.002 | 20/67 | 22/64 | 0.725 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IDH2 (+/-) | 11/76 | 6/80 | 0.307 | 8/79 | 9/77 | 0.804 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IDH1 (+/-) | 11/76 | 5/81 | 0.188 | 4/83 | 12/74 | 0.038 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TET2 (+/-) | 5/82 | 10/76 | 0.188 | 8/79 | 7/79 | 1.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| RUNX1 (+/-) | 8/79 | 7/79 | 1.000 | 9/78 | 6/80 | 0.590 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TP53 (+/-) | 8/79 | 6/80 | 0.782 | 2/85 | 12/74 | 0.005 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NRAS (+/-) | 8/79 | 4/82 | 0.370 | 7/80 | 5/81 | 0.766 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CEBPA (+/-) | 4/83 | 9/77 | 0.162 | 6/81 | 7/79 | 0.782 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| WT1 (+/-) | 4/83 | 6/80 | 0.535 | 4/83 | 6/80 | 0.535 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PTPN11 (+/-) | 5/82 | 3/83 | 0.720 | 4/83 | 4/82 | 1.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KIT (+/-) | 1/86 | 6/80 | 0.064 | 5/82 | 2/84 | 0.443 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| U2AF1 (+/-) | 4/83 | 3/83 | 1.000 | 3/84 | 4/82 | 0.720 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KRAS (+/-) | 4/83 | 3/83 | 1.000 | 4/83 | 3/83 | 1.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMC1A (+/-) | 4/83 | 3/83 | 1.000 | 5/82 | 2/84 | 0.443 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMC3 (+/-) | 4/83 | 3/83 | 1.000 | 3/84 | 4/82 | 0.720 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PHF6 (+/-) | 3/84 | 2/84 | 1.000 | 3/84 | 2/84 | 1.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| STAG2 (+/-) | 2/85 | 3/83 | 0.682 | 2/85 | 3/83 | 0.682 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| RAD21 (+/-) | 2/85 | 2/84 | 1.000 | 3/84 | 1/85 | 0.621 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| AML: acute myeloid leukemia; WBC: white blood cells; PB: peripheral blood; BM: bone marrow; FAB: French-American-British. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Correlation between DNMT3A/3B expression and gene mutations in AML
Among the common gene mutations in AML, patients with DNMT3A overexpression showed lower NPM1 and DNMT3A mutation rates (P=0.000 and 0.002, respectively), whereas cases with DNMT3B underexpression presented lower frequency of IDH1 and TP53 mutation (P=0.038 and 0.005, respectively). In addition, we further compared the expression of DNMT3A and DNMT3B in mutation and wild-type groups of these genes (Figure 3).
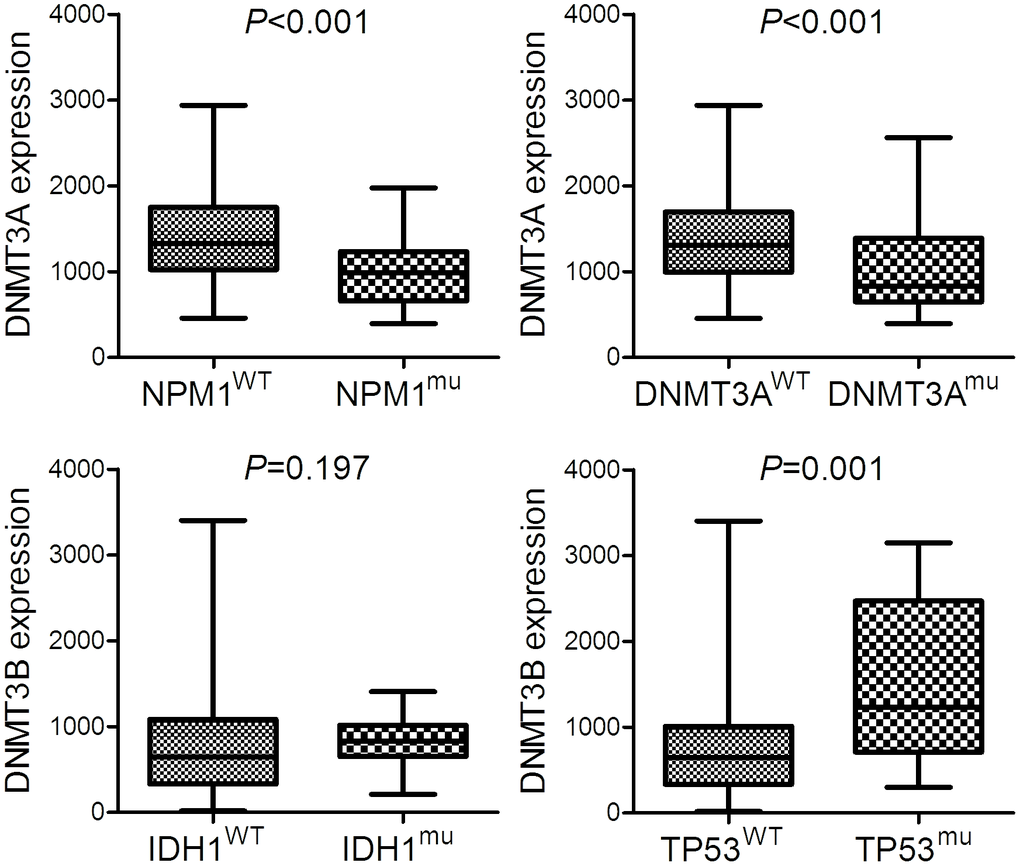
Figure 3. The expression of DNMT3A and DNMT3B in AML patients with different molecular signature. The expression of DNMT3A in AML patients with and without NPM1 mutation as well as AML patients with and without DNMT3A mutation. The expression of DNMT3B in AML patients with and without IDH1 mutation as well as AML patients with and without TP53 mutation.
Prognostic value of DNMTs expression in AML
We next evaluate the prognostic effect of DNMTs expression on survival in AML. By Kaplan-Meier analysis, only DNMT3A overexpression was associated with longer overall survival (OS) and leukemia-free survival (LFS) in whole-cohort AML (P=0.001 and 0.003, respectively, Figure 4). In order to confirm the independent prognostic value of DNMT3A expression on both OS and LFS, we performed Cox regression analysis adjusting for prognosis-related factors. By Cox regression multivariate analysis, DNMT3A expression could act as an independent prognostic biomarker for OS in whole-cohort AML (Table 2). However, no significant differences were observed in either DNMT1 or DNMT3B groups (Figure 4).
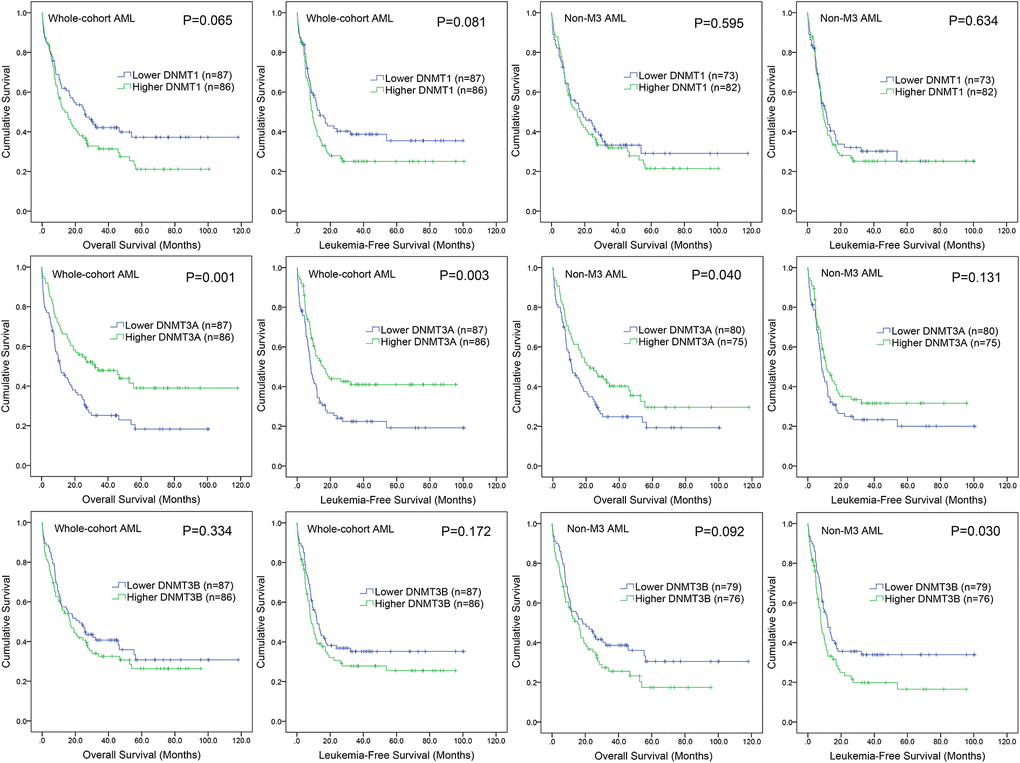
Figure 4. The impact of DNMTs expression on survival of AML patients. Kaplan–Meier survival curves of DNMTs expression on overall survival and leukemia-free survival in both chemotherapy and hematopoietic stem cell transplantation (HSCT) groups.
Table 2. Cox regression analyses of variables for overall survival and leukemia-free survival in AML.
| Variables | Overall survival | Leukemia-free survival | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hazard ratio (95% CI) | P | Hazard ratio (95% CI) | P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DNMT3A expression | 0.628 (0.429-0.920) | 0.017 | 0.696 (0.476-1.020) | 0.063 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age | 1.038 (1.022-1.053) | 0.000 | 1.034 (1.019-1.049) | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| WBC | 1.008 (1.003-1.012) | 0.001 | 1.008 (1.004-1.012) | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular risk | 2.148 (1.537-3.000) | 0.000 | 1.901 (1.382-2.614) | 0.000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FLT3 mutation | 1.686 (1.082-2.627) | 0.021 | 1.719 (1.100-2.687) | 0.017 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CEBPA mutation | 1.685 (0.799-3.553) | 0.171 | 1.732 (0.814-3.687) | 0.154 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NPM1 mutation | 0.742 (0.425-1.297) | 0.295 | 0.810 (0.471-1.394) | 0.447 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DNMT3A mutation | 1.309 (0.812-2.110) | 0.269 | 1.134 (0.717-1.793) | 0.592 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| RUNX1 mutation | 1.940 (1.288-2.924) | 0.002 | 1.660 (1.104-2.498) | 0.015 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TET2 mutation | 0.767 (0.386-1.524) | 0.448 | 0.824 (0.414-1.639) | 0.581 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TP53 mutation | 2.900 (1.483-5.669) | 0.002 | 2.616 (1.350-5.068) | 0.004 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IDH1 mutation | 0.702 (0.337-1.463) | 0.344 | 0.751 (0.344-1.639) | 0.472 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IDH2 mutation | 0.644 (0.338-1.226) | 0.180 | 0.649 (0.344-1.225) | 0.183 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ASXL1 mutation | 1.779 (0.503-6.289) | 0.372 | 1.813 (0.509-6.459) | 0.359 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| AML: acute myeloid leukemia; CI: confidence interval; WBC: white blood cells. Variables in multivariate analysis including DNMT3A expression (low vs. high), age, WBC, karyotype (favorable vs. intermediate vs. poor), and gene mutations (mutant vs. wild-type). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
DNMT3A expression may guide treatment choice between chemotherapy and HSCT
Because lower DNMT3A expression predicted poor clinical outcome in AML, we intended to investigate whether patients with lower DNMT3A expression could benefit from HSCT. We compared OS and LFS between patients with and without HSCT in both lower and higher DNMT3A expression groups. In lower DNMT3A expression groups, patients who received HSCT showed significantly longer OS and LFS than patients who were not treated with HSCT among both total AML (Figure 5). However, in higher DNMT3A expression groups, there were no significant differences in OS and LFS between two groups (Figure 5). Taken together, AML patients with lower DNMT3A expression could benefit from HSCT, whereas those with higher DNMT3A expression did not. Therefore, we deduced that DNMT3A expression pattern may guide treatment choice between chemotherapy and HSCT.
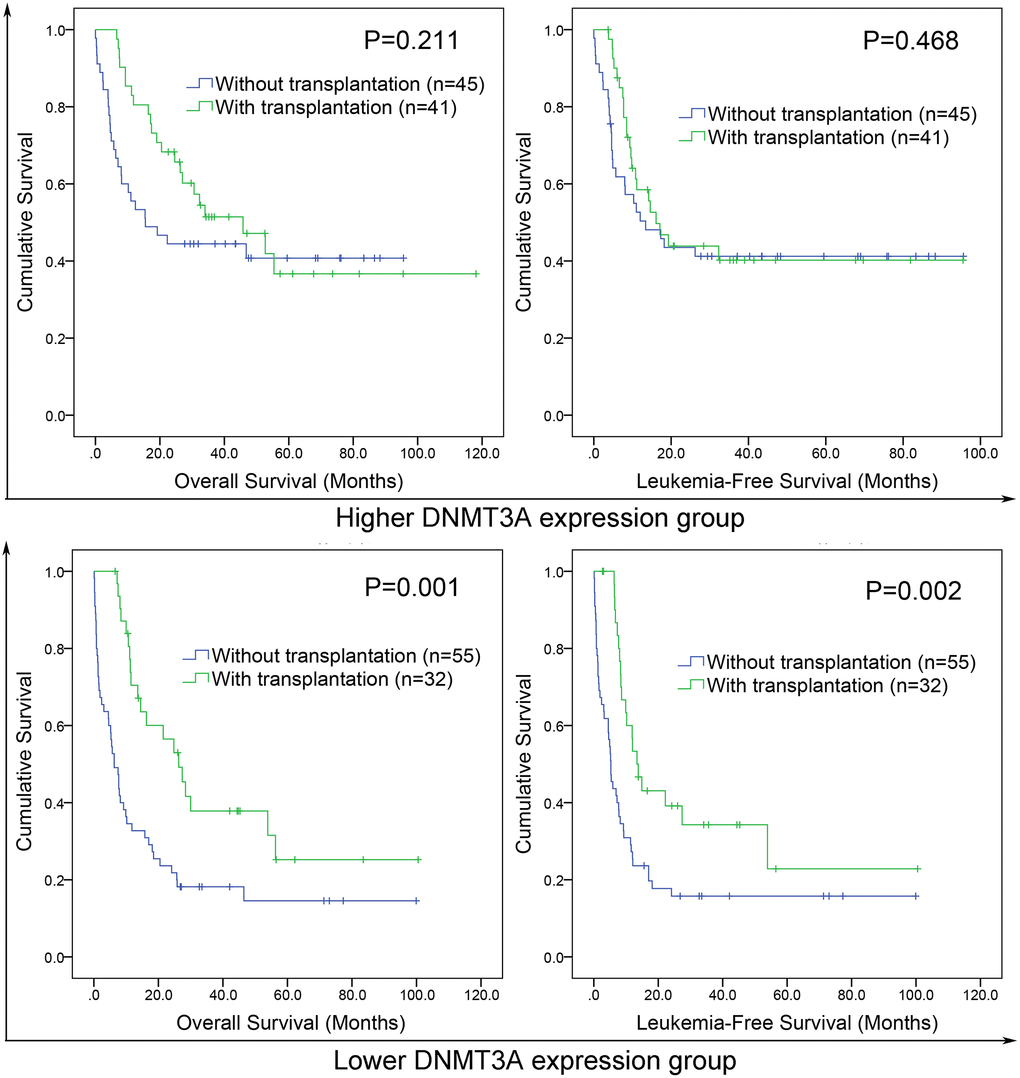
Figure 5. The effect of hematopoietic stem cell transplantation (HSCT) on survival of AML patients among different DNMT3A expression groups. Kaplan–Meier survival curves of overall survival and leukemia-free survival in low and high DNMT3A expression group.
Discussion
In this study, we systemically analyzed DNMTs expression and their relationship with clinic-pathological features and prognosis in patients with AML. We found that DNMT3A expression was increased in AML, whereas DNMT3B expression was decreased in AML. Although previous study showed DNMTs overexpression and its negative prognostic effects in AML [18–21], recent researches revealed that DNMT3B expression was decreased in AML blasts, whereas DNMT1 and DNMT3A expression showed no significant differences [16, 33]. Additionally, the potential roles of DNMT3A and DNMT3B in AML remained poorly defined. Peters et al previously showed the tumor suppressor functions of DNMT3A and DNMT3B in the prevention of malignant mouse lymphopoiesis, but not in the development of myeloid malignancies [34]. However, ectopic DNMT3B expression was reported to delay leukemogenesis [17], and the loss of DNMT3B accelerated MLL-AF9 leukemia progression [16]. These studies demonstrated that DNMT3B played a crucial role in AML development, but may not act as a cancer-related driver gene during leukemogenesis. As for DNMT3A, it was indicated that DNMT3A loss progressively impaired HSC differentiation [13]. Notably, loss of DNMT3A and endogenous KRASG12D/+ cooperated to regulate hematopoietic stem and progenitor cell functions in leukemogenesis [35]. In our study, we also observed the significant associations of DNMT3A expression with other molecular events such as NPM1 and DNMT3A mutations. These studies suggested that DNMT3A generally not worked independently in the development of AML, and it may cooperate with other molecular events.
The prognostic value of DNMT3A mutation in AML has been systemically revealed. Increasing studies showed that DNMT3A mutations were independently associated with poor outcome in AML patients with an intermediate-risk cytogenetic profile or CN-AML [12, 36]. Moreover, loss-of-function of DNMT3A caused by mutations or underexpression predicted response to the HMAs decitabine treatment in AML [37]. In this study, low DNMT3A expression was observed to act as an independent prognostic biomarker in AML and also helpful for the selecting treatment choice between chemotherapy and HSCT. Interestingly, although two recent reports showed that high DNMT3B expression was a poor prognostic biomarker in AML [19, 20], we did not observe the association of aberrant DNMT3B expression with AML survival. The differences may be caused by the specific cell population selection and different ethnics. Obviously, prospective studies are needed to confirm and expand our results before DNMT3A expression pattern can be used routinely as a potential prognostic biomarker guiding treatment choice for newly diagnosed AML.
Despite that the role of DNMT3A in regulation of DNA methylation is well-known, the potential mechanism regulating DNMT3A was poorly investigated. Jost et al reported that aberrant DNA hypermethylation within the DNMT3A gene was frequently observed in AML, and was associated with downregulation of DNMT3A mRNA transcript 2 [38]. Moreover, DNMT3A was also identified as a direct target of a number of microRNAs such as miR-30a-3p, miR-133a-3p, miR-450, miR-29a/b/c, and miR-129-5p [39–43]. From our study, we observed the direct association of two microRNAs miR-29b and miR-429 with DNMT3A in AML. Although several studies showed the direct link between DNMT3A and miR-29b in other human cancers, little studies showed the direct correlation between miR-429 and DNMT3A in any type of human cancers. Interestingly, our pervious study disclosed that miR-429 expression was decreased in AML [32], which presented the opposite expression pattern of DNMT3A in AML. Accordingly, further studies are required to confirm the direct associations of DNMT3A with miR-429 by luciferase assay.
In summary, although we analyzed the expression and prognosis analysis of DNMTs expression only by public databases, our study demonstrated that DNMT3A and DNMT3B showed significant expression differences in AML. Moreover, DNMT3A expression acted as a potential prognostic biomarker and may guide treatment choice between chemotherapy and HSCT in AML.
Materials and Methods
CCLE, HPA, and EMBL-EBI dataset
DNMTs expression in human cancer cell lines were assessed by the Cancer Cell Line Encyclopedia (CCLE) dataset (https://www.broadinstitute.org/ccle) [44] and also evaluated by The Human Protein Atlas (HPA) dataset (https://www.proteinatlas.org/) [45]. Moreover, DNMTs expression in AML cell lines was verified by the European Bioinformatics Institute (EMBL-EBI) dataset (https://www.ebi.ac.uk) [46].
GEPIA dataset
DNMTs expression in AML patients and normal controls was analyzed by the Gene Expression Profiling Interactive Analysis (GEPIA) web (http://gepia.cancer-pku.cn/), whose data obtained from The Cancer Genome Atlas (TCGA) and the Genotype-Tissue Expression (GTEx) projects [47].
Patients from TCGA datasets
A total of 173 AML patients with available DNMTs expression data from TCGA were identified and included in this study [15]. Clinical and molecular characteristics were collected including age, sex, white blood cell (WBC) counts, peripheral blood (PB) blasts, bone marrow (BM) blasts, French-American-British (FAB) subtypes and the frequencies of genetic mutations. After induction chemotherapy, consolidation treatment included chemotherapy (100 patients received) and hematopoietic stem cell transplantation (HSCT) (73 patients received).
Bioinformatics analyses
The details for the identification of microRNAs targeting DNMT3A were reported as our previous study [48, 49].
Statistical analyses
SPSS 22.0 were used for statistical analyses and figures creation. Mann-Whitney’s U test was used for the comparison of continuous variables, whereas Pearson Chi-square analysis or Fisher exact test was applied for the comparison of categorical variables. The prognostic effect of DNMTs expression was evaluated using leukemia-free survival (LFS) and overall survival (OS) analyzed though Kaplan-Meier analysis and Cox regression analysis. The two-tailed P value < 0.05 in all statistical analyses was defined as statistically significant.
Ethical approval and consent to participate
The present study approved by the Ethics Committee and Institutional Review Board of the Affiliated People’s Hospital of Jiangsu University. Written informed consents were obtained from all enrolled individuals prior to their participation.
Author Contributions
Jing-dong Zhou conceived and designed the experiments, Ting-juan Zhang, Liu-chao Zhang, Zi-jun Xu and Jing-dong Zhou analyzed the data, all authors read and approved the final manuscript.
Conflicts of Interest
The authors declare that they have no conficts of interest.
Funding
The work was supported by National Natural Science foundation of China (81900166, 81970118), Zhenjiang Clinical Research Center of Hematology (SS2018009). Very thanks for Prof. Jun Qian providing technical and financial support.
References
- 1. Li E, Zhang Y. DNA methylation in mammals. Cold Spring Harb Perspect Biol. 2014; 6:a019133. https://doi.org/10.1101/cshperspect.a019133 [PubMed]
- 2. Kulis M, Esteller M. DNA methylation and cancer. Adv Genet. 2010; 70:27–56. https://doi.org/10.1016/B978-0-12-380866-0.60002-2 [PubMed]
- 3. Kim SY, Han YK, Song JM, Lee CH, Kang K, Yi JM, Park HR. Aberrantly hypermethylated tumor suppressor genes were identified in oral squamous cell carcinoma (OSCC). Clin Epigenetics. 2019; 11:116. https://doi.org/10.1186/s13148-019-0715-0 [PubMed]
- 4. Schoofs T, Müller-Tidow C. DNA methylation as a pathogenic event and as a therapeutic target in AML. Cancer Treat Rev. 2011 (Suppl 1); 37:S13–18. https://doi.org/10.1016/j.ctrv.2011.04.013 [PubMed]
- 5. Spencer DH, Russler-Germain DA, Ketkar S, Helton NM, Lamprecht TL, Fulton RS, Fronick CC, O’Laughlin M, Heath SE, Shinawi M, Westervelt P, Payton JE, Wartman LD, et al. CpG island hypermethylation mediated by DNMT3A is a consequence of AML progression. Cell. 2017; 168:801–16.e13. https://doi.org/10.1016/j.cell.2017.01.021 [PubMed]
- 6. Jiang Y, Dunbar A, Gondek LP, Mohan S, Rataul M, O’Keefe C, Sekeres M, Saunthararajah Y, Maciejewski JP. Aberrant DNA methylation is a dominant mechanism in MDS progression to AML. Blood. 2009; 113:1315–25. https://doi.org/10.1182/blood-2008-06-163246 [PubMed]
- 7. Daver N, Boddu P, Garcia-Manero G, Yadav SS, Sharma P, Allison J, Kantarjian H. Hypomethylating agents in combination with immune checkpoint inhibitors in acute myeloid leukemia and myelodysplastic syndromes. Leukemia. 2018; 32:1094–105. https://doi.org/10.1038/s41375-018-0070-8 [PubMed]
- 8. Yang L, Rau R, Goodell MA. DNMT3A in haematological Malignancies. Nat Rev Cancer. 2015; 15:152–65. https://doi.org/10.1038/nrc3895 [PubMed]
- 9. Okano M, Xie S, Li E. Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet. 1998; 19:219–20. https://doi.org/10.1038/890 [PubMed]
- 10. Brunetti L, Gundry MC, Goodell MA. DNMT3A in leukemia. Cold Spring Harb Perspect Med. 2017; 7:a030320. https://doi.org/10.1101/cshperspect.a030320 [PubMed]
- 11. Trowbridge JJ, Sinha AU, Zhu N, Li M, Armstrong SA, Orkin SH. Haploinsufficiency of Dnmt1 impairs leukemia stem cell function through derepression of bivalent chromatin domains. Genes Dev. 2012; 26:344–49. https://doi.org/10.1101/gad.184341.111 [PubMed]
- 12. Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE, Kandoth C, Payton JE, Baty J, Welch J, Harris CC, Lichti CF, Townsend RR, et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010; 363:2424–33. https://doi.org/10.1056/NEJMoa1005143 [PubMed]
- 13. Challen GA, Sun D, Jeong M, Luo M, Jelinek J, Berg JS, Bock C, Vasanthakumar A, Gu H, Xi Y, Liang S, Lu Y, Darlington GJ, et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat Genet. 2011; 44:23–31. https://doi.org/10.1038/ng.1009 [PubMed]
- 14. Dai YJ, Wang YY, Huang JY, Xia L, Shi XD, Xu J, Lu J, Su XB, Yang Y, Zhang WN, Wang PP, Wu SF, Huang T, et al. Conditional knockin of Dnmt3a R878H initiates acute myeloid leukemia with mTOR pathway involvement. Proc Natl Acad Sci USA. 2017; 114:5237–42. https://doi.org/10.1073/pnas.1703476114 [PubMed]
- 15. Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, Robertson A, Hoadley K, Triche TJ
Jr , Laird PW, Baty JD, Fulton LL, Fulton R, Heath SE, et al, and Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013; 368:2059–74. https://doi.org/10.1056/NEJMoa1301689 [PubMed] - 16. Zheng Y, Zhang H, Wang Y, Li X, Lu P, Dong F, Pang Y, Ma S, Cheng H, Hao S, Tang F, Yuan W, Zhang X, Cheng T. Loss of Dnmt3b accelerates MLL-AF9 leukemia progression. Leukemia. 2016; 30:2373–84. https://doi.org/10.1038/leu.2016.112 [PubMed]
- 17. Stanley RF, Steidl U. Ectopic DNMT3B expression delays leukemogenesis. Blood. 2016; 127:1525–26. https://doi.org/10.1182/blood-2016-01-692137 [PubMed]
- 18. Mizuno S, Chijiwa T, Okamura T, Akashi K, Fukumaki Y, Niho Y, Sasaki H. Expression of DNA methyltransferases DNMT1, 3A, and 3B in normal hematopoiesis and in acute and chronic myelogenous leukemia. Blood. 2001; 97:1172–79. https://doi.org/10.1182/blood.v97.5.1172 [PubMed]
- 19. Hayette S, Thomas X, Jallades L, Chabane K, Charlot C, Tigaud I, Gazzo S, Morisset S, Cornillet-Lefebvre P, Plesa A, Huet S, Renneville A, Salles G, et al. High DNA methyltransferase DNMT3B levels: a poor prognostic marker in acute myeloid leukemia. PLoS One. 2012; 7:e51527. https://doi.org/10.1371/journal.pone.0051527 [PubMed]
- 20. Niederwieser C, Kohlschmidt J, Volinia S, Whitman SP, Metzeler KH, Eisfeld AK, Maharry K, Yan P, Frankhouser D, Becker H, Schwind S, Carroll AJ, Nicolet D, et al. Prognostic and biologic significance of DNMT3B expression in older patients with cytogenetically normal primary acute myeloid leukemia. Leukemia. 2015; 29:567–75. https://doi.org/10.1038/leu.2014.267 [PubMed]
- 21. Lin N, Fu W, Zhao C, Li B, Yan X, Li Y. Biologico-clinical significance of DNMT3A variants expression in acute myeloid leukemia. Biochem Biophys Res Commun. 2017; 494:270–77. https://doi.org/10.1016/j.bbrc.2017.10.041 [PubMed]
- 22. Larmonie NS, Arentsen-Peters TC, Obulkasim A, Valerio D, Sonneveld E, Danen-van Oorschot AA, de Haas V, Reinhardt D, Zimmermann M, Trka J, Baruchel A, Pieters R, van den Heuvel-Eibrink MM, et al. MN1 overexpression is driven by loss of DNMT3B methylation activity in inv(16) pediatric AML. Oncogene. 2018; 37:107–15. https://doi.org/10.1038/onc.2017.293 [PubMed]
- 23. Morita K, Masamoto Y, Kataoka K, Koya J, Kagoya Y, Yashiroda H, Sato T, Murata S, Kurokawa M. BAALC potentiates oncogenic ERK pathway through interactions with MEKK1 and KLF4. Leukemia. 2015; 29:2248–56. https://doi.org/10.1038/leu.2015.137 [PubMed]
- 24. Zhang TJ, Zhou JD, Zhang W, Lin J, Ma JC, Wen XM, Yuan Q, Li XX, Xu ZJ, Qian J. H19 overexpression promotes leukemogenesis and predicts unfavorable prognosis in acute myeloid leukemia. Clin Epigenetics. 2018; 10:47. https://doi.org/10.1186/s13148-018-0486-z [PubMed]
- 25. Zhou JD, Li XX, Zhang TJ, Xu ZJ, Zhang ZH, Gu Y, Wen XM, Zhang W, Ji RB, Deng ZQ, Lin J, Qian J. MicroRNA-335/ ID4 dysregulation predicts clinical outcome and facilitates leukemogenesis by activating PI3K/Akt signaling pathway in acute myeloid leukemia. Aging (Albany NY). 2019; 11:3376–91. https://doi.org/10.18632/aging.101991 [PubMed]
- 26. Su YL, Wang X, Mann M, Adamus TP, Wang D, Moreira DF, Zhang Z, Ouyang C, He X, Zhang B, Swiderski PM, Forman SJ, Baltimore D, et al. Myeloid cell-targeted miR-146a mimic inhibits NF-κB-driven inflammation and leukemia progression in vivo. Blood. 2020; 135:167–180. https://doi.org/10.1182/blood.2019002045 [PubMed]
- 27. Mammoli F, Parenti S, Lomiento M, Gemelli C, Atene CG, Grande A, Corradini R, Manicardi A, Fantini S, Zanocco-Marani T, Ferrari S. Physiological expression of miR-130a during differentiation of CD34+ human hematopoietic stem cells results in the inhibition of monocyte differentiation. Exp Cell Res. 2019; 382:111445. https://doi.org/10.1016/j.yexcr.2019.05.026 [PubMed]
- 28. Raffel S, Trumpp A. miR-126 drives quiescence and self-renewal in leukemic stem cells. Cancer Cell. 2016; 29:133–35. https://doi.org/10.1016/j.ccell.2016.01.007 [PubMed]
- 29. Jiang X, Hu C, Arnovitz S, Bugno J, Yu M, Zuo Z, Chen P, Huang H, Ulrich B, Gurbuxani S, Weng H, Strong J, Wang Y, et al. miR-22 has a potent anti-tumour role with therapeutic potential in acute myeloid leukaemia. Nat Commun. 2016; 7:11452. https://doi.org/10.1038/ncomms11452 [PubMed]
- 30. Gong JN, Yu J, Lin HS, Zhang XH, Yin XL, Xiao Z, Wang F, Wang XS, Su R, Shen C, Zhao HL, Ma YN, Zhang JW. The role, mechanism and potentially therapeutic application of microRNA-29 family in acute myeloid leukemia. Cell Death Differ. 2014; 21:100–12. https://doi.org/10.1038/cdd.2013.133 [PubMed]
- 31. Zhu B, Xi X, Liu Q, Cheng Y, Yang H. MiR-9 functions as a tumor suppressor in acute myeloid leukemia by targeting CX chemokine receptor 4. Am J Transl Res. 2019; 11:3384–97. [PubMed]
- 32. Zhou JD, Zhang LC, Zhang TJ, Gu Y, Wu DH, Zhang W, Ma JC, Wen XM, Guo H, Lin J, Qian J. Dysregulation of miR-200s clusters as potential prognostic biomarkers in acute myeloid leukemia. J Transl Med. 2018; 16:135. https://doi.org/10.1186/s12967-018-1494-7 [PubMed]
- 33. Schulze I, Rohde C, Scheller-Wendorff M, Bäumer N, Krause A, Herbst F, Riemke P, Hebestreit K, Tschanter P, Lin Q, Linhart H, Godley LA, Glimm H, et al. Increased DNA methylation of Dnmt3b targets impairs leukemogenesis. Blood. 2016; 127:1575–86. https://doi.org/10.1182/blood-2015-07-655928 [PubMed]
- 34. Peters SL, Hlady RA, Opavska J, Klinkebiel D, Pirruccello SJ, Talmon GA, Sharp JG, Wu L, Jaenisch R, Simpson MA, Karpf AR, Opavsky R. Tumor suppressor functions of Dnmt3a and Dnmt3b in the prevention of Malignant mouse lymphopoiesis. Leukemia. 2014; 28:1138–42. https://doi.org/10.1038/leu.2013.364 [PubMed]
- 35. Chang YI, You X, Kong G, Ranheim EA, Wang J, Du J, Liu Y, Zhou Y, Ryu MJ, Zhang J. Loss of Dnmt3a and endogenous kras(G12D/+) cooperate to regulate hematopoietic stem and progenitor cell functions in leukemogenesis. Leukemia. 2015; 29:1847–56. https://doi.org/10.1038/leu.2015.85 [PubMed]
- 36. Thol F, Damm F, Lüdeking A, Winschel C, Wagner K, Morgan M, Yun H, Göhring G, Schlegelberger B, Hoelzer D, Lübbert M, Kanz L, Fiedler W, et al. Incidence and prognostic influence of DNMT3A mutations in acute myeloid leukemia. J Clin Oncol. 2011; 29:2889–96. https://doi.org/10.1200/JCO.2011.35.4894 [PubMed]
- 37. Metzeler KH, Walker A, Geyer S, Garzon R, Klisovic RB, Bloomfield CD, Blum W, Marcucci G. DNMT3A mutations and response to the hypomethylating agent decitabine in acute myeloid leukemia. Leukemia. 2012; 26:1106–07. https://doi.org/10.1038/leu.2011.342 [PubMed]
- 38. Jost E, Lin Q, Weidner CI, Wilop S, Hoffmann M, Walenda T, Schemionek M, Herrmann O, Zenke M, Brümmendorf TH, Koschmieder S, Wagner W. Epimutations mimic genomic mutations of DNMT3A in acute myeloid leukemia. Leukemia. 2014; 28:1227–34. https://doi.org/10.1038/leu.2013.362 [PubMed]
- 39. Wei D, Yu G, Zhao Y. MicroRNA-30a-3p inhibits the progression of lung cancer via the PI3K/AKT by targeting DNA methyltransferase 3a. Onco Targets Ther. 2019; 12:7015–24. https://doi.org/10.2147/OTT.S213583 [PubMed]
- 40. Shi W, Tang T, Li X, Deng S, Li R, Wang Y, Wang Y, Xia T, Zhang Y, Zen K, Jin L, Pan Y. Methylation-mediated silencing of miR-133a-3p promotes breast cancer cell migration and stemness via miR-133a-3p/MAML1/DNMT3A positive feedback loop. J Exp Clin Cancer Res. 2019; 38:429. https://doi.org/10.1186/s13046-019-1400-z [PubMed]
- 41. Wang Y, Wang L, Yu X, Duan J. Overexpression of miR-450 affects the biological behavior of HepG2 cells by targeting DNMT3a. Onco Targets Ther. 2019; 12:5069–76. https://doi.org/10.2147/OTT.S203206 [PubMed]
- 42. Zhu Y, Zheng G, Wang H, Jia Y, Zhang Y, Tang Y, Li W, Fan Y, Zhang X, Liu Y, Liu S. Downregulated miR-29a/b/c during contact inhibition stage promote 3T3-L1 adipogenesis by targeting DNMT3A. PLoS One. 2017; 12:e0170636. https://doi.org/10.1371/journal.pone.0170636 [PubMed]
- 43. Gu X, Gong H, Shen L, Gu Q. MicroRNA-129-5p inhibits human glioma cell proliferation and induces cell cycle arrest by directly targeting DNMT3A. Am J Transl Res. 2018; 10:2834–47. [PubMed]
- 44. Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, Wilson CJ, Lehár J, Kryukov GV, Sonkin D, Reddy A, Liu M, Murray L, et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012; 483:603–07. https://doi.org/10.1038/nature11003 [PubMed]
- 45. Uhlén M, Björling E, Agaton C, Szigyarto CA, Amini B, Andersen E, Andersson AC, Angelidou P, Asplund A, Asplund C, Berglund L, Bergström K, Brumer H, et al. A human protein atlas for normal and cancer tissues based on antibody proteomics. Mol Cell Proteomics. 2005; 4:1920–32. https://doi.org/10.1074/mcp.M500279-MCP200 [PubMed]
- 46. Li W, Cowley A, Uludag M, Gur T, McWilliam H, Squizzato S, Park YM, Buso N, Lopez R. The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res. 2015; 43:W580–84. https://doi.org/10.1093/nar/gkv279 [PubMed]
- 47. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017; 45:W98–102. https://doi.org/10.1093/nar/gkx247 [PubMed]
- 48. Zhou JD, Zhang TJ, Xu ZJ, Gu Y, Ma JC, Li XX, Guo H, Wen XM, Zhang W, Yang L, Liu XH, Lin J, Qian J. BCL2 overexpression: clinical implication and biological insights in acute myeloid leukemia. Diagn Pathol. 2019; 14:68. https://doi.org/10.1186/s13000-019-0841-1 [PubMed]
- 49. Chu MQ, Zhang TJ, Xu ZJ, Gu Y, Ma JC, Zhang W, Wen XM, Lin J, Qian J, Zhou JD. EZH2 dysregulation: potential biomarkers predicting prognosis and guiding treatment choice in acute myeloid leukaemia. J Cell Mol Med. 2020; 24:1640–49. https://doi.org/10.1111/jcmm.14855 [PubMed]