Introduction
Esophageal squamous cell carcinoma (ESCC) is one of the most common digestive tract tumors worldwide and is associated with high morbidity and mortality, especially in Asia [1]. Despite improvements in surgical techniques and perioperative management, the 5-year survival rate of ESCC is less than 30% [1]. Currently, the most frequently examined biomarkers for ESCC diagnosis are serum squamous cell carcinoma antigen (SCCA), carbohydrate antigen (CA) 19–9, and carcinoembryonic antigen (CEA). However, these biomarkers are relatively insensitive and inaccurate [1]. Novel effective biomarkers for ESCC are therefore urgently needed.
Long non-coding RNAs (lncRNAs) are longer than 200 bases in length and do not code for proteins. An increasing number of studies indicate that lncRNAs can affect cancer initiation, progression, and metastasis by modulating oncogenic and tumor-suppressing pathways [2]. Recently, several lncRNAs have been identified that are secreted via exosomes from tumor tissues into the plasma, and dysregulated expression of these lncRNAs in exosomes could potentially serve as a diagnostic or prognostic biomarker for many tumors, such as liver and lung cancer [3, 4]. However, few studies have comprehensively explored exosome lncRNA profiles in plasma from ESCC patients.
Here, we conducted a four-stage study to identify plasma exosome lncRNAs with diagnostic potential in ESCC. First, lncRNA expression profiles were characterized in plasma exosomes from ESCC patients using RNA-sequencing. Five lncRNAs, NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1, were found to be significantly increased in the ESCC patients. Expression of these five lncRNAs was then evaluated by qRT-PCR in a large cohort of ESCC patients, esophagitis patients, and healthy controls. Levels of all five lncRNAs were significantly increased in exosomes from ESCC patients and decreased in those patients after surgery. Our results therefore indicate that these five exosome lncRNAs might serve as useful biomarkers for ESCC diagnosis.
Results
Characterization of plasma exosomes
Because exosome purity affects the variability of exosome lncRNA analysis results, TEM, Western blotting, and nanoparticle tracking analysis were used to characterize exosomes isolated from the plasma of ESCC patients, esophagitis patients, and healthy controls. As shown in Figure 1A, exosomes were confirmed to have sizes falling within the typical range as well as normal rounded membrane-bound morphologies using TEM. Expression of the established exosome markers CD63, CD9, and CD81 was then characterized through Western immunoblotting analysis (Figure 1B). Finally, nanoparticle tracking analysis was used to confirm that the exosome diameters were approximately 100 nm as previously reported (Figure 1C) [5]. These results demonstrated that exosomes were successfully extracted from plasma.
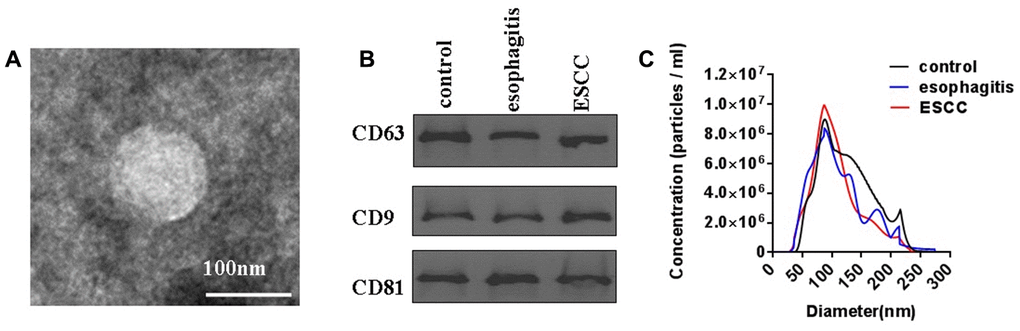
Figure 1. Characterization of plasma exosomes from ESCC patients, esophagitis patients, and healthy controls. (A) Shape and structure of plasma exosomes isolated by ultracentrifugation under TEM. (B) Western blots of exosome membrane markers CD63, CD9, and CD81. (C) Sizes of plasma exosomes from ESCC patients, esophagitis patients, and healthy controls were analyzed by NTA.
Identification of exosome lncRNA profiles using Illumina HiSeq 2500
In order to identify plasma exosome lncRNAs with diagnostic potential in ESCC, a four-stage study was conducted (Figure 2). Exosome lncRNAs in pooled plasma from ESCC patients, esophagitis patients, and healthy controls were profiled using Illumina HiSeq 2500 analysis; each pooled sample included plasma from ten patients. The heatmap shown in Figure 3 revealed distinct lncRNA expression patterns for plasma exosomes from ESCC patients, esophagitis patients, and healthy controls. LncRNAs with a median number of reads > 100 and with a ≥ 2-fold change (P < 0.05) between ESCC patients and esophagitis patients/healthy controls were selected as potential biomarkers. The following five lncRNAs were significantly increased in ESCC patients compared to esophagitis patients and healthy controls: NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 (Table 1).
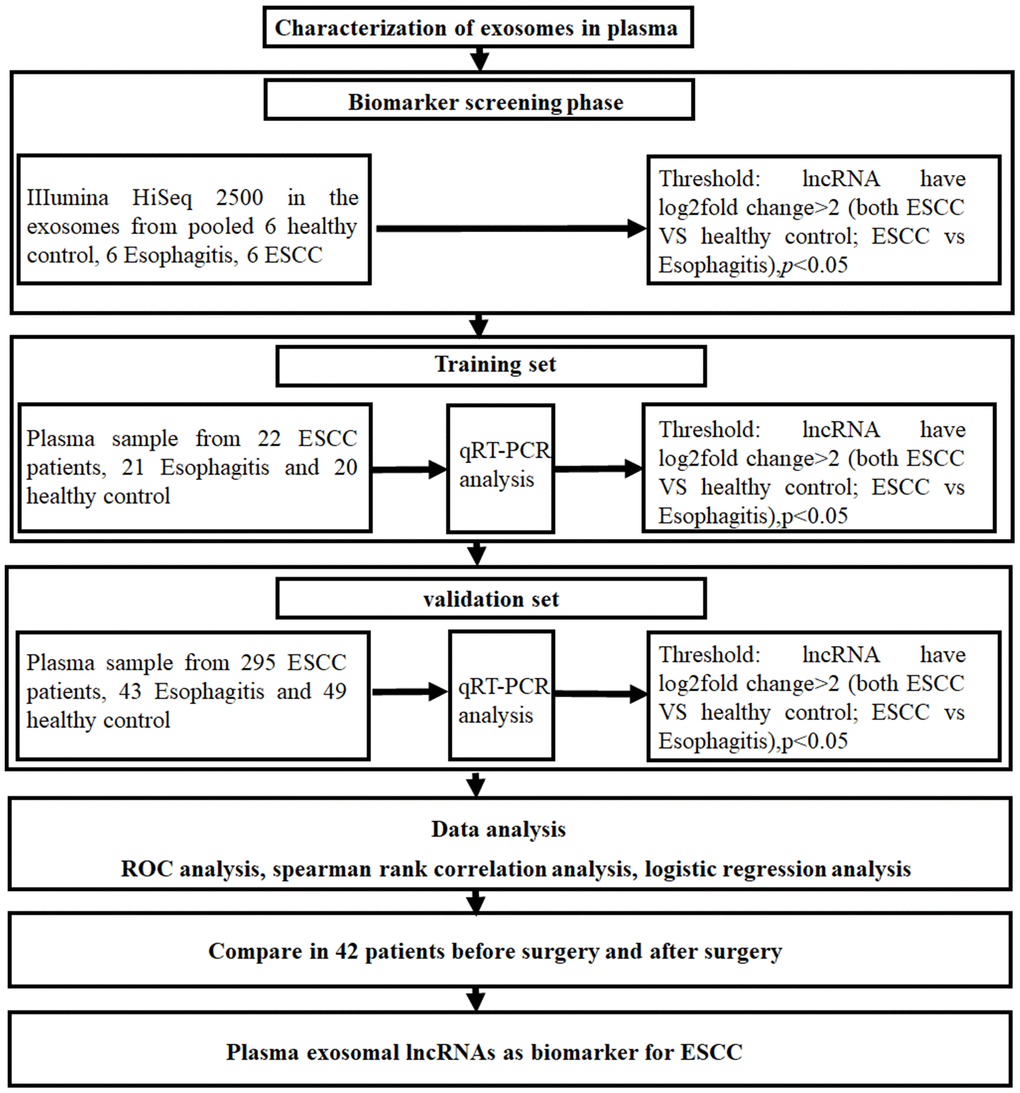
Figure 2. Experimental design flowchart.
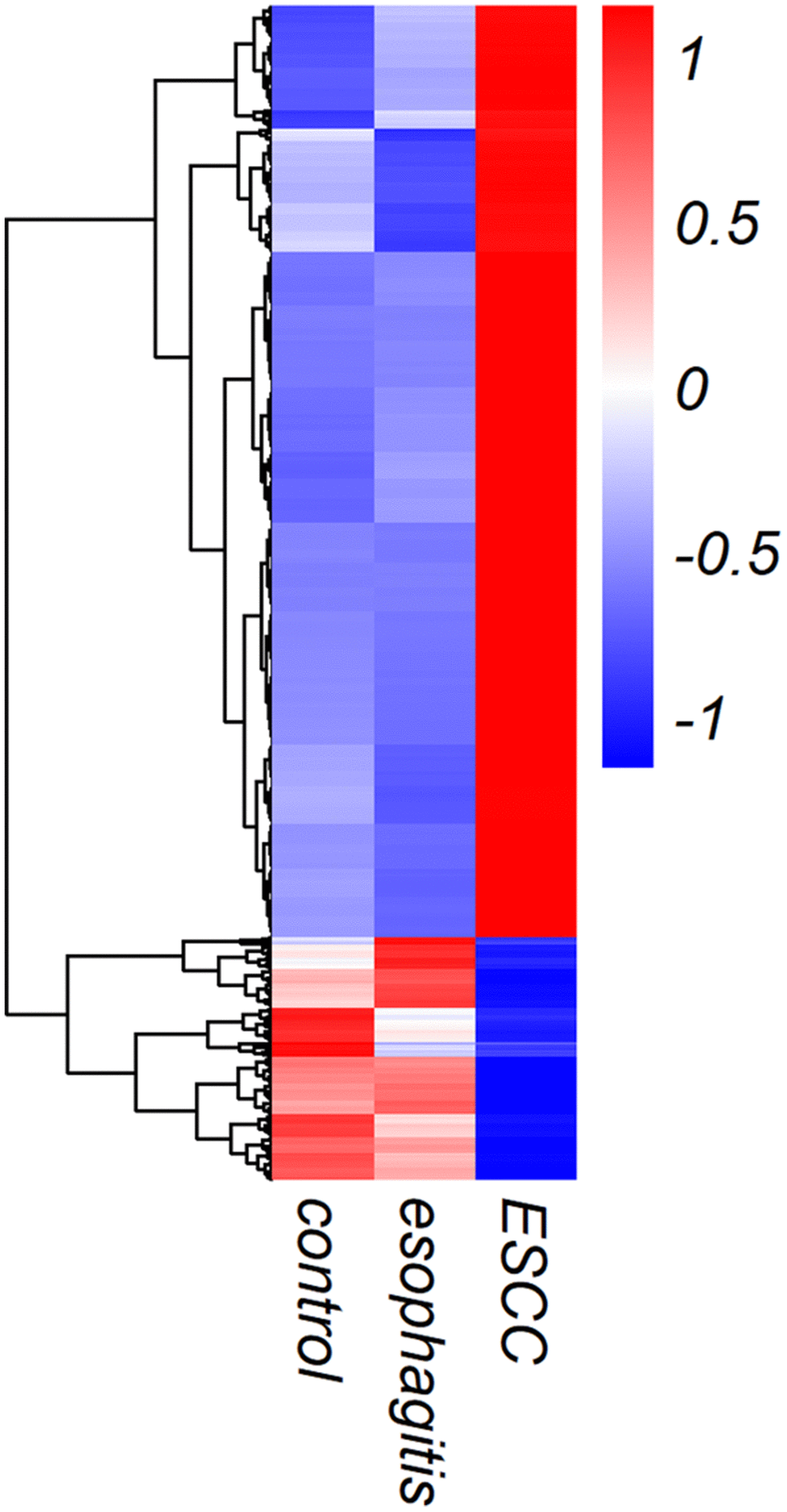
Figure 3. Heat map showing lncRNA expression in exosomes from ESCC patients, esophagitis patients, and healthy controls.
Table 1. LncRNAs with reads > 100, log2(fold change) ≥ 2, and P < 0.05 in Illumina HiSeq 2500 biomarker screening.
| Transcript | Gene | Control | Esophagitis | ESCC | log2(fold change) | p-value | log2(fold change) | p-value |
| (ESCC vs. control) | (ESCC vs. control) | (ESCC vs. esophagitis) | (ESCC vs. esophagitis) | |||||
| NR_039819 | pre-MIR4672 | 0.3807 | 0.4221 | 7.0044 | 4.2013 | 0.0237 | 4.0527 | 0.0176 |
| NR_036133 | pre-MIR323B | 0.7824 | 0.2191 | 12.2192 | 3.9652 | 0.0019 | 5.8015 | 0.0012 |
| NR_003353 | SNORD115-38 | 0.8659 | 1.1275 | 10.7715 | 3.6369 | 0.0201 | 3.2561 | 0.0137 |
| ENST00000442416.1 | LINC02832 | 0.8827 | 1.1183 | 7.2648 | 3.0409 | 0.0000 | 2.6997 | 0.0001 |
| ENST00000416100.1 | AC093734.1 | 1.7574 | 1.5737 | 7.1267 | 2.0198 | 0.0064 | 2.1791 | 0.0034 |
qRT-PCR confirmed increased plasma exosome lncRNA levels in ESCC
qRT-PCR using the Illumina HiSeq 2500 was performed to confirm increased expression the five potential lncRNA biomarker candidates identified above in plasma exosomes of ESCC patients (n=22) compared to esophagitis patients (n=21) and healthy volunteers (n=20) in the training set. As expected, plasma exosome NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 expression was at least 2-fold higher in ESCC patients than in esophagitis patients and healthy volunteers (P < 0.05). Next, we evaluated these five lncRNAs in a larger cohort consisting of 295 ESCC patients, 43 esophagitis patients, and 49 healthy volunteers. Consistent with previous results, NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 expression was higher in plasma exosomes from ESCC patients than in the other groups (P < 0.001, Figure 4A–4E). A logistic regression model analysis of the combined diagnostic value of these five lncRNAs revealed an AUC of 0.9995 (P < 0.001) (Figure 4F). Taken together, these results suggest that these five lncRNAs could serve as useful and stable diagnostic biomarkers for ESCC.
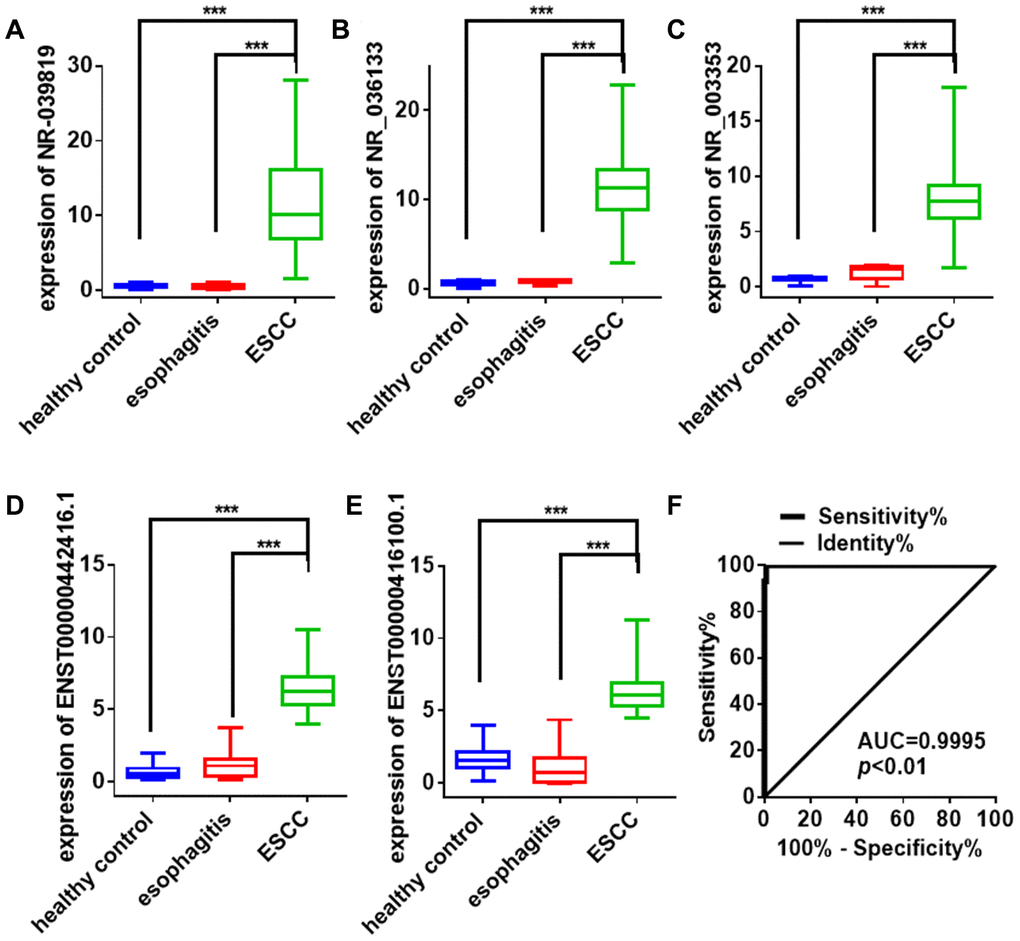
Figure 4. Exosome lncRNA expression signature for ESCC diagnosis. (A–E) Differences in plasma exosome lncRNAs levels in ESCC patients, esophagitis patients, and healthy controls. Each point represents the mean of triplicate samples. Each P-value was calculated with a nonparametric Mann-Whitney test. ***P<0.001. (F) ROC curve analysis of the combined lncRNA panel in exosomes from ESCC patients and non-cancer controls (including healthy controls and esophagitis patients). Each value is the mean±SD; ***P <0.001.
Comparison of plasma exosome lncRNAs before and after surgery
To investigate whether the five plasma exosome lncRNAs identified above could be used to monitor treatment efficacy in ESCC patients, we collected plasma samples from 42 stage I ESCC patients before and one month after surgery. Plasma exosome NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 levels decreased significantly one month after surgery compared to levels before surgery (Figure 5).
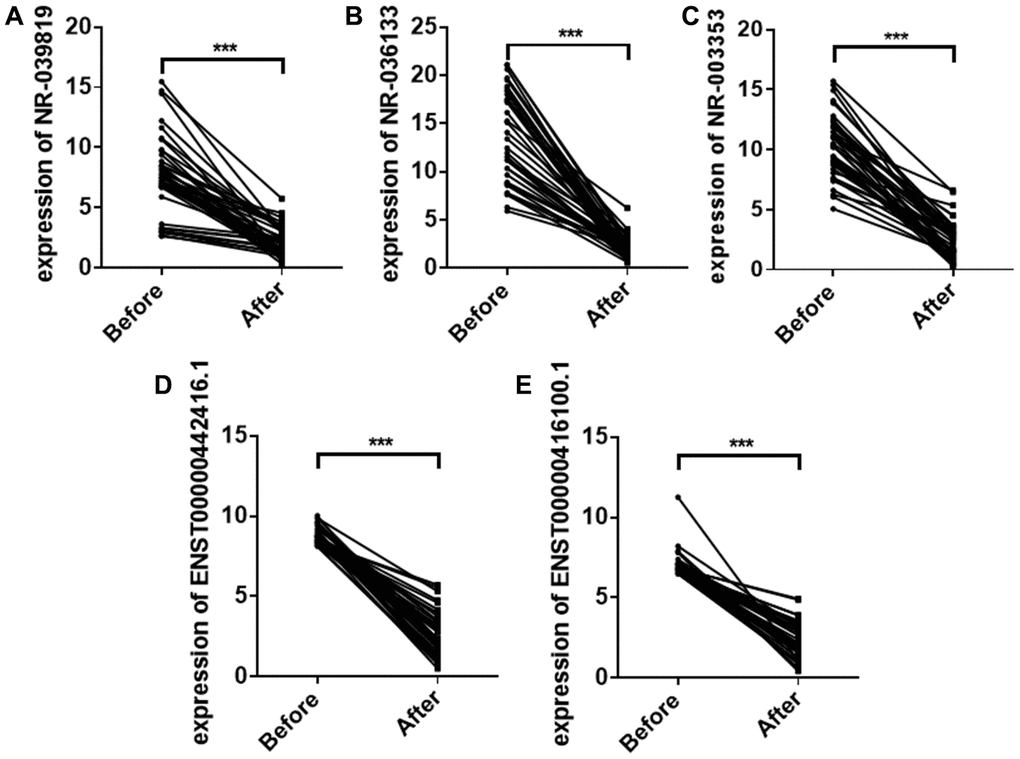
Figure 5. Plasma exosome NR_039819 (A) NR_036133 (B) NR_ 003353 (C) ENST00000442416.1(D) and ENST0000041 6100.1 (E) levels in ESCC patients before surgery and after surgery. Each point represents the mean of triplicate samples. Each P-value was calculated with a nonparametric Mann-Whitney test. ***P<0.001.
Discussion
Cell-free nucleic acids are detectable in plasma or serum exosomes from tumor patients and might therefore serve as novel biomarkers for tumor diagnosis [7]. Although numerous studies have focused on miRNAs in these exosomes as potential biomarkers for ESCC diagnosis and prognosis prediction, several recent studies have examined plasma exosome lncRNA levels in ESCC [8–11].
In this study, we screened plasma exosome lncRNAs in ESCC patients using the Illumina HiSeq 2500. LncRNAs with a mean number of reads greater than100 and a greater than 2-fold change in ESCC patients compared to esophagitis patients and healthy volunteers were further evaluated in two large independent patient cohorts by qRT-PCR. The results demonstrated that plasma exosome NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 levels were significantly higher in ESCC patients compared to esophagitis patients and healthy volunteers. Moreover, we also examined the expression of these five lncRNAs in exosomes from ESCC patient plasma before and after surgery. Consistent with our previous results, plasma exosome NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 expression decreased significantly after surgery. Together, these results provide strong evidence that plasma exosome lncRNAs could be used as novel diagnostic markers for ESCC.
LncRNAs, a relatively recently discovered type of long non-coding RNA, play critical roles in tumor development, differentiation, and formation [12]. Exosomes can mediate cellular communication in cancer by transmitting active molecules, including lncRNAs [12]. Two of the five lncRNAs identified in our study, NR_039819 and NR_036133, are precursors of MIR4672 and MIR323B. Several studies have demonstrated that exosomes can deliver miRNA precursors to target cells, allowing those precursors to reprogram the target cell transcriptome [13]. Additionally, we found that expression of these five lncRNAs was significantly increased in ESCC tumor tissues compared to distal normal tissues (Figure 6). Exosomes are known to mediate cellular communication by transmitting lncRNAs in ESCC [9, 14, 15]. For example, Li et al. confirmed that the exosomal lncRNA ZFAS1 could promote ESCC growth via the microRNA-124/STAT3 axis [9]. Exosomal FMR1-AS1 helped maintain dynamic equilibrium in cancer stem-like cells through the TLR7/NFκB/c-Myc signaling pathway [14]. Kang et al. demonstrated that exosomes facilitate transfer of the lncRNA PART1, which induces gefitinib resistance in esophageal squamous cell carcinoma via the miR-129/blc-2 axis [15]. Existing evidence therefore suggests that the five plasma exosome lncRNAs identified here might play important roles in the development and progression of ESCC. However, further studies are needed to confirm those roles.
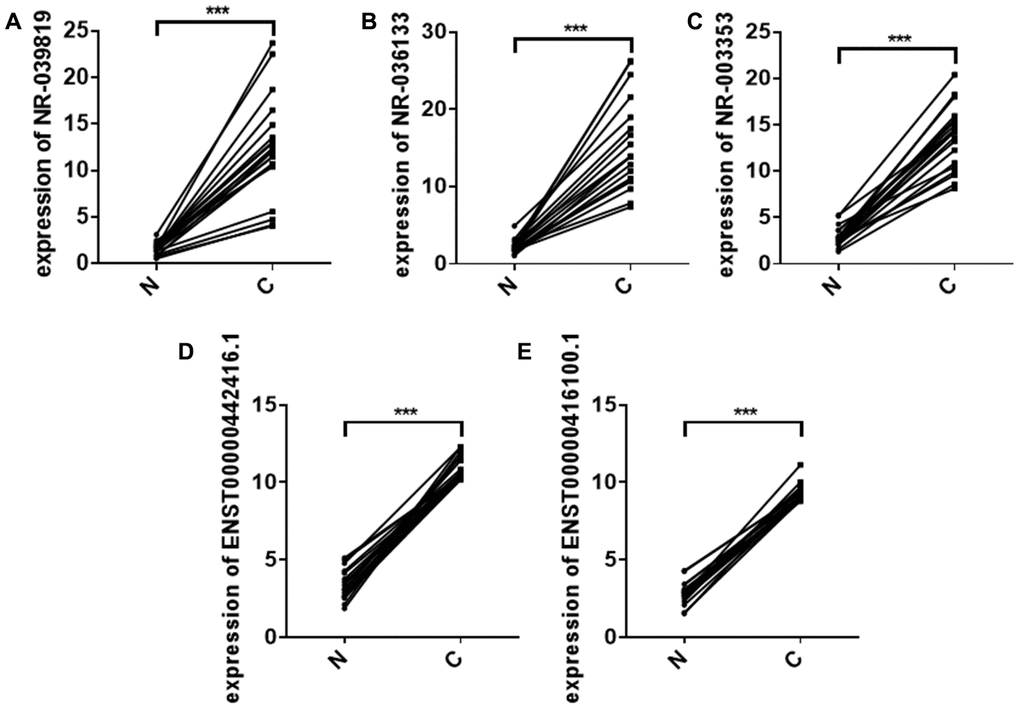
Figure 6. Relative NR_039819 (A) NR_036133 (B) NR_ 003353 (C) ENST00000442416.1 (D) and ENST0000041 6100.1 (E) levels in 20 paired non-cancerous (N) and ESCC cancer (C) tissues determined using qRT- PCR. ***P < 0.001.
In summary, our results suggest that increased expression of the plasma exosome lncRNAs NR_039819, NR_036133, NR_003353, ENST00000442416.1, and ENST00000416100.1 might serve as a useful tumor biomarker for ESCC diagnosis.
Materials and Methods
Patient characteristics
Plasma samples from 317 untreated ESCC patients, as well as 64 esophagitis patients and 69 healthy controls who were age-, sex-, and ethnicity-matched with the ESCC patients, were included in this study. Paired preoperative and postoperative plasma samples (n = 42) were collected from stage I ESCC patients before surgery and one month after resection. Twenty pairs of ESCC tissues and adjacent non-cancerous tissues were collected from patients who were diagnosed with ESCC at the Nanjing Drum Tower Hospital. The demographic and clinical characteristics of the patients and healthy volunteers are summarized in Table 2. Esophagogastroduodenoscopies were performed for all subjects (Figure 7). A lack of abnormal mucosa, ulcers, or abnormal protuberances in the esophagus was confirmed in healthy controls. Esophagitis patients had rough mucosa covered with white moss and with visible submucosal blood vessels. ESCC patients had rough and congested surrounding mucosa with rigid tube walls, poor peristalsis, narrow lumens, and intraluminal protrusions. Diagnoses for non-cancer esophagitis patients and ESCC patients were confirmed by histopathology. This study was approved by the Ethics Committee of Nanjing Drum Tower Hospital, and written informed consent was obtained from all patients. All experiments were performed in accordance with relevant guidelines and regulations. Protocols were designed and performed according to the principles of the Helsinki Declaration.
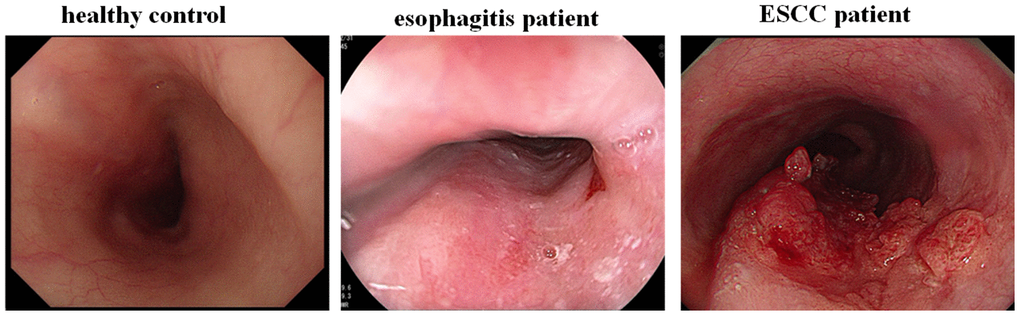
Figure 7. Endoscopic examination of ESCC patients, esophagitis patients and healthy controls.
Table 2. Demographic and clinical characteristics of ESCC patients, esophagitis patients, and healthy controls in the training and validation sets.
| Variable | Control (n=69) | Esophagitis (n=64) | ESCC (n=317) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age, yearsa | 44.6(16.54) | 45.3(18.25) | 44.4(18.43) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sex, n | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| male | 35 | 36 | 208 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Female | 34 | 28 | 109 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stage, n | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| I-II | 216 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| III-IV | 101 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| aAge data are presented as the mean (SD). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Plasma exosome isolation and RNA extraction
Exosomes were isolated from plasma using the Total Exosome Isolation Kit (from plasma) (Cat# 4484450, Invitrogen, USA) following the manufacturer’s protocol. Briefly, plasma samples were centrifuged at 1,000×g for 20 min, 3,000×g for 20 min, and 10,000×g for 20 min immediately after collection. The clarified plasma was then mixed with 0.5 volume of PBS and 0.3 volume of the Exosome Precipitation Reagent. The mixture was incubated at 4°C for 4 hours and then centrifuged at 10,000×g for 20 minutes. Subsequently, the pellet containing exosomes was collected and resuspended in 200 μL PBS for downstream analysis. The mirVana PARIS Kit (Ambion, Thermo Scientific, Shanghai, China) was used to extract total RNA from the exosomes.
Characterization of exosomes isolated from plasma via transmission electron microscopy (TEM), nanoparticle tracking analysis, and Western blotting
Transmission electron microscopy (TEM) of exosomes isolated from plasma was performed as previously described [5]. A JEM-1011 scanning transmission electron microscope (Hitachi, Tokyo, Japan) was used to obtain the microphotographs. Nanoparticle tracking analysis was performed to identify exosomes isolated from plasma using the ZetaView PMX 110 (Particle Metrix, Meerbusch, Germany) according to the guidelines of the International Society for Extracellular Vesicles and the manufacturer’s protocol [5]. Western blotting analysis was used to characterize expression of the exosome protein markers CD63, CD81, and CD9 as previously described [5]. CD63, CD81, and CD9 were detected using anti-CD63, anti-CD81, and CD9 rabbit polyclonal antibodies (Abcam, USA). Bound proteins were visualized using ECL Western blotting substrate (Thermo Fisher Scientific, MA, USA) and analyzed with ImageJ software.
Illumina HiSeq 2500 analysis
LncRNA expression profiles in plasma exosomes were evaluated using an Illumina HiSeq 2500 following the manufacturer’s protocol. LncRNA quantities were normalized and reported as reads per million (RPM). The EdgeR package [6] was used to analyze dysregulated lncRNAs and calculate associated p values. LncRNAs with a log2 (fold change) > 1 and p < 0.05 were classified as showing differential expression.
qRT-PCR for lncRNAs
The qRT-qPCR assay for exosome lncRNA isolated from 1 mL plasma as previous described was performed following the manufacturer's protocol (Applied Biosystems, Shanghai, China). All reactions were performed in triplicate, and the data were analyzed using the comparative Cq method. The primers used are listed in Table 3
Table 3. Primers used in qRT-PCR.
| Name | Forward (5'-3') | Reverse (5'-3') |
| NR_039819 | GCTGCTTCTCGCCTCTGT | GGCTGTTTCGTGCCTCTG |
| NR_036133 | TGGTACTCGGAGGGAGGTTG | GAGGTCGACCGTGTATTGGG |
| NR_003353 | GGGTCAATGATGAGAACCTTACA | GGGCCTCAGCGTAATCCTAT |
| ENST00000442416.1 | AGTAGTGCTCGTGAAGCGAT | ATCCCAAGGGTGATGTCAGG |
| ENST00000416100.1 | GCCACAGGATGGGAGCTTAT | GTGGCCGTCTTCTCTCTGTG |
Statistical analyses
Statistical analyses were performed using SPSS 18.0 software and graphs were generated using GraphPad Prism 6.0. The Mann-Whitney U test was used to test the statistical significance of differences in plasma exosome lncRNA expression. Spearman correlation analysis was performed to identify correlations between lncRNA and clinical indicators. P-values < 0.05 was regarded as statistically significant.
Author Contributions
Tao Wang designed the experiments. Zichen Jiao, Ao Yu, Xiaofeng He, Minke Shi, and Weiwei Rong performed the experiments and analyzed results. Tao Wang wrote the manuscript. Ke Zen provided critical reading and constructive discussion of the manuscript.
Conflicts of Interest
The authors declare that they have no conflicts of interest regarding the publication of this paper.
References
- 1. van Hagen P, Hulshof MC, van Lanschot JJ, Steyerberg EW, van Berge Henegouwen MI, Wijnhoven BP, Richel DJ, Nieuwenhuijzen GA, Hospers GA, Bonenkamp JJ, Cuesta MA, Blaisse RJ, Busch OR, et al, and CROSS Group. Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med. 2012; 366:2074–84. https://doi.org/10.1056/NEJMoa1112088 [PubMed]
- 2. Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012; 22:1775–89. https://doi.org/10.1101/gr.132159.111 [PubMed]
- 3. Feng Q, Zhang H, Yao D, Chen WD, Wang YD. Emerging role of non-coding RNAs in esophageal squamous cell carcinoma. Int J Mol Sci. 2019; 21:258. https://doi.org/10.3390/ijms21010258 [PubMed]
- 4. Dhanoa JK, Sethi RS, Verma R, Arora JS, Mukhopadhyay CS. Long non-coding RNA: its evolutionary relics and biological implications in mammals: a review. J Anim Sci Technol. 2018; 60:25. https://doi.org/10.1186/s40781-018-0183-7 [PubMed]
- 5. Yao B, Qu S, Hu R, Gao W, Jin S, Liu M, Zhao Q. A panel of miRNAs derived from plasma extracellular vesicles as novel diagnostic biomarkers of lung adenocarcinoma. FEBS Open Bio. 2019; 9:2149–58. https://doi.org/10.1002/2211-5463.12753 [PubMed]
- 6. Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010; 26:139–40. https://doi.org/10.1093/bioinformatics/btp616 [PubMed]
- 7. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011; 11:426–37. https://doi.org/10.1038/nrc3066 [PubMed]
- 8. Tian L, Yang L, Zheng W, Hu Y, Ding P, Wang Z, Zheng D, Fu L, Chen B, Xiao T, Wang Y, Chen F, Liu J, et al. RNA sequencing of exosomes revealed differentially expressed long noncoding RNAs in early-stage esophageal squamous cell carcinoma and benign esophagitis. Epigenomics. 2020; 12:525–41. https://doi.org/10.2217/epi-2019-0371 [PubMed]
- 9. Li Z, Qin X, Bian W, Li Y, Shan B, Yao Z, Li S. Exosomal lncRNA ZFAS1 regulates esophageal squamous cell carcinoma cell proliferation, invasion, migration and apoptosis via microRNA-124/STAT3 axis. J Exp Clin Cancer Res. 2019; 38:477. https://doi.org/10.1186/s13046-019-1473-8 [PubMed]
- 10. Zhao A, Guo L, Xu J, Zheng L, Guo Z, Ling Z, Wang L, Mao W. Identification and validation of circulating exosomes-based liquid biopsy for esophageal cancer. Cancer Med. 2019; 8:3566–74. https://doi.org/10.1002/cam4.2224 [PubMed]
- 11. Lin Y, Dong H, Deng W, Lin W, Li K, Xiong X, Guo Y, Zhou F, Ma C, Chen Y, Ren H, Yang H, Dai N, et al. Evaluation of salivary exosomal chimeric GOLM1-NAA35 RNA as a potential biomarker in esophageal carcinoma. Clin Cancer Res. 2019; 25:3035–45. https://doi.org/10.1158/1078-0432.CCR-18-3169 [PubMed]
- 12. Mashouri L, Yousefi H, Aref AR, Ahadi AM, Molaei F, Alahari SK. Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer. 2019; 18:75. https://doi.org/10.1186/s12943-019-0991-5 [PubMed]
- 13. Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, Qiu L, Vitkin E, Perelman LT, Melo CA, Lucci A, Ivan C, Calin GA, Kalluri R. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014; 26:707–21. https://doi.org/10.1016/j.ccell.2014.09.005 [PubMed]
- 14. Li W, Zhang L, Guo B, Deng J, Wu S, Li F, Wang Y, Lu J, Zhou Y. Exosomal FMR1-AS1 facilitates maintaining cancer stem-like cell dynamic equilibrium via TLR7/NFκB/c-myc signaling in female esophageal carcinoma. Mol Cancer. 2019; 18:22. https://doi.org/10.1186/s12943-019-0949-7 [PubMed]
- 15. Kang M, Ren M, Li Y, Fu Y, Deng M, Li C. Exosome-mediated transfer of lncRNA PART1 induces gefitinib resistance in esophageal squamous cell carcinoma via functioning as a competing endogenous RNA. J Exp Clin Cancer Res. 2018; 37:171. https://doi.org/10.1186/s13046-018-0845-9 [PubMed]


