Introduction
Alzheimer’s disease (AD) is an irreversible, progressive brain disorder characterized by memory, thinking and cognitive skills deterioration that results in behavioral problems among affected elderly individuals. While there are many environmental factors that may influence the risk of AD, it is estimated that nearly 70% of the variance in AD may be explained by genetic determinants [1, 2]. Although GWAS studies have identified over 20 disease-associated genomic loci AD-associated loci, the causal mechanisms implicated in the onset of AD remain unclear [3]. Traditional epidemiology studies reported several common risk factors for AD including obesity [4, 5], type 2 diabetes (T2D) and glycemic traits [6, 7], smoking [8], and lower education level [8]. However, these findings may be influenced by unmeasured confounding factors that may obscure the true relationship.
The Mendelian randomization (MR) approach [9] enables us to assess the potential causal effect of a risk factor on the outcome by using genetic instrumental variables (IVs). Previous two sample MR studies have demonstrated several potential causal risk factors for AD, such as lower education level and smoking [10]. Multivariable MR (mvMR) [11] is an extension of two sample MR approach that incorporates a set of pleiotropic SNPs [12] associated with several risk factors to simultaneously assess the causal effect of each risk factor on the outcome. In comparison with two sample MR, mvMR assumes the genetic IV is associated with at least one risk factor, although not necessarily all risk factors. Additionally, under the circumstances of horizontal pleiotropy, causal effect can be assessed even if none of the variants show specific associations with any individual risk factor [11]. Burgess et al. [13] successfully applied mvMR to estimate the causal effects of lipid fractions on cardiovascular artery disease. However, the current application of mvMR are not capable of feature ranking and selecting.
Recently, Zuber V et al. developed a novel approach [14] which combined multivariable MR with Bayesian model averaging (MR-BMA) that scales to high dimensional settings and can select biomarkers as causal risk factors for the disease of interest. Study demonstrated that the method can detect and prioritize true risk factors even when the multiple risk factors are highly correlated [14]. The MR-BMA approach has been successfully applied to prioritize the most likely causal metabolites for age-related macular degeneration [14].
Of all the previously reported risk factors for AD, it remains unclear which are causal, and which may play the most pivotal role in disease susceptibility. In the current study we intend to integrate the mvMR and MR-BMA approach to identify and prioritize the most likely causal risk factors for AD, the risk factors included in the current study are body mass index (BMI), type 2 diabetes (T2D), high-density lipoprotein cholesterol (HDL cholesterol), low-density lipoprotein cholesterol (LDL cholesterol), total cholesterol, fasting glucose (FG), fasting insulin (FI), currently tobacco smoking, and years of schooling.
Results
Genetic IVs selection and validation
Overall, we obtained 1235 LD-independent SNPs that achieved genome-wide significance for all the risk factors after implementing the pruning strategy previously described. Then those SNPs were extracted from AD dataset. After harmonizing the exposure and outcome datasets, there were 1159 SNPs remained for the MR analysis. The number of IVs included for each risk factor was demonstrated in Table 1, and detailed information for the characteristics of SNPs used for each risk factor was shown in Supplementary Table 1.
Table 1. Baseline information of exposures and outcomes used in our analysis.
| Exposure | Sample size | Population | PMID | Numbers of IVs | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Body mass index (BMI) | 461460 | European | - | 360 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Current tobacco smoking | 462434 | European | - | 27 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Fasting insulin | 108557 | European | 22885924 | 13 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Fasting glucose | 133010 | European | 22885924 | 31 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HDL cholesterol | 196476 | Mix* | 24097068 | 85 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LDL cholesterol | 196476 | Mix* | 24097068 | 72 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Total cholesterol | 196476 | Mix* | 24097068 | 78 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Type 2 diabetes | 659316 | European | 30054458 | 101 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Years of schooling | 1131881 | European | 30038396 | 248 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Notes: Mix population was composed of 188,578 European-ancestry individuals and 7,898 non-European ancestry individuals. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
mvMR estimates results
Our standard MR approach showed that genetically increased years of schooling (OR = 0.674, 95% confidence interval (CI): 0.571-0.796, P = 3.337E-06) and elevated HDL cholesterol (OR = 0.761, 95% CI: 0.697-0.830, P = 6.940E-10) were significantly associated with decreasing risk of AD. We found a significant association between total cholesterol and AD, the odds ratio per genetically predicted 1 SD higher total cholesterol level was 1.300 (1.196 to 1.412; P = 6.223E-10) (Table 2 and Figure 1). Elevated LDL cholesterol level was associated with increased ADD susceptibility (OR ranging from 1.097 to 1.296 per SD increment in genetically determined LDL cholesterol level, P < 3.564E-05). However, there was a suggestive association between FG and AD, one SD increase in FG was associated with 29.7% increase in AD risk (OR = 1.297, 95% CI: 1.013-1.661, P = 0.039). No association was observed between the other risk factors and AD, and for detailed information please find Figure 1 and Table 2.
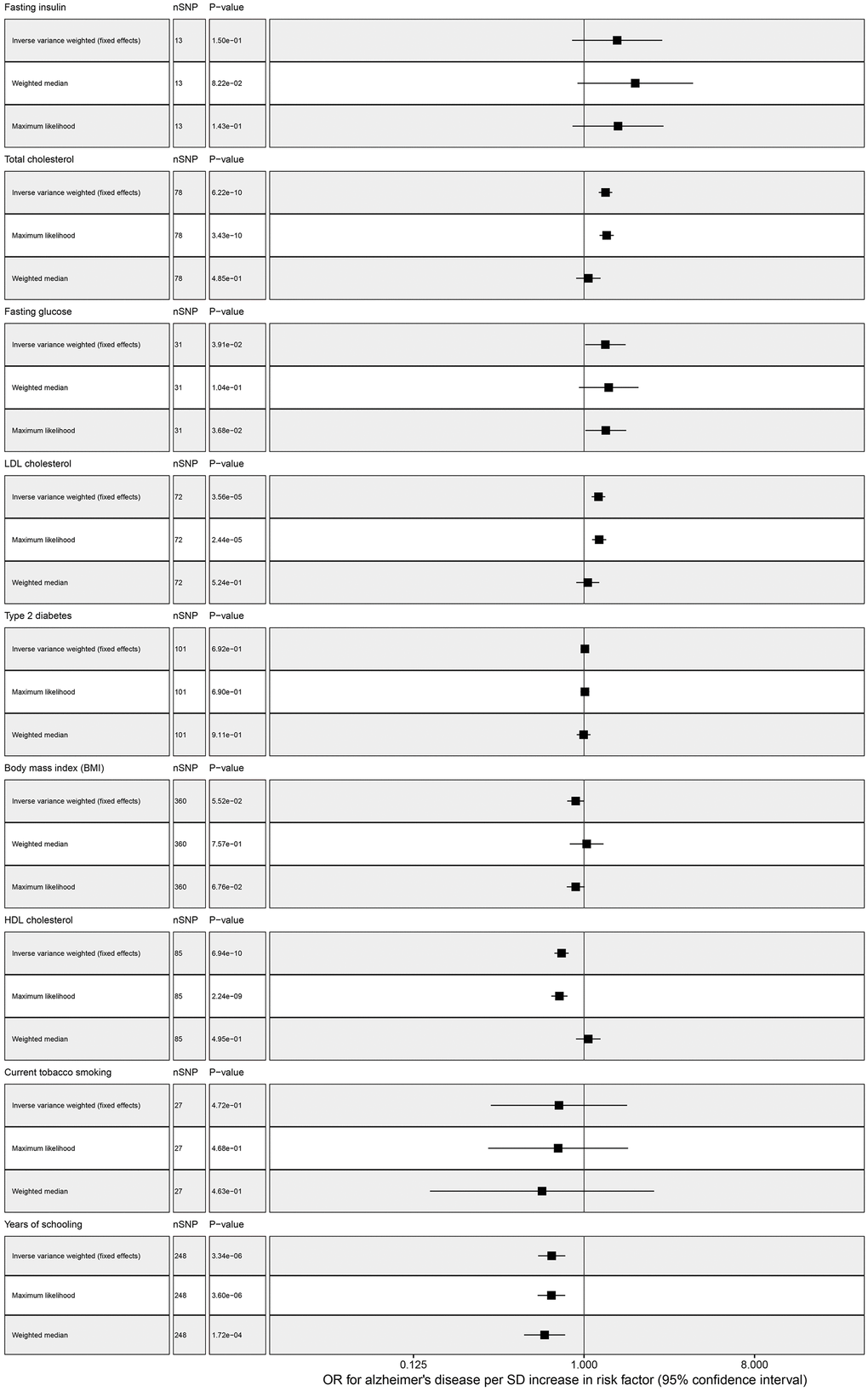
Figure 1. Multivariable MR analysis forest plot: effect of multiple risk factors on AD.
Table 2. Multivariable MR analysis for risk factors and AD.
| Exposure | Methods | nsnp | OR | OR_lci95 | OR_uci95 | P_value |
| Years of schooling | IVW fixed effects | 248 | 0.674 | 0.571 | 0.796 | 3.337E-06 |
| Years of schooling | Maximum likelihood | 248 | 0.672 | 0.568 | 0.795 | 3.597E-06 |
| Years of schooling | Weighted median | 248 | 0.619 | 0.481 | 0.795 | 1.814E-04 |
| T2D | Weighted median | 101 | 0.995 | 0.915 | 1.082 | 9.106E-01 |
| T2D | IVW fixed effects | 101 | 1.011 | 0.959 | 1.066 | 6.921E-01 |
| T2D | Maximum likelihood | 101 | 1.011 | 0.958 | 1.067 | 6.900E-01 |
| Total cholesterol | IVW fixed effects | 78 | 1.300 | 1.196 | 1.412 | 6.223E-10 |
| Total cholesterol | Maximum likelihood | 78 | 1.318 | 1.209 | 1.436 | 3.430E-10 |
| Total cholesterol | Weighted median | 78 | 1.055 | 0.912 | 1.220 | 4.742E-01 |
| LDL cholesterol | IVW fixed effects | 72 | 1.193 | 1.097 | 1.296 | 3.564E-05 |
| LDL cholesterol | Maximum likelihood | 72 | 1.203 | 1.104 | 1.311 | 2.439E-05 |
| LDL cholesterol | Weighted median | 72 | 1.047 | 0.912 | 1.201 | 5.151E-01 |
| HDL cholesterol | IVW fixed effects | 85 | 0.761 | 0.697 | 0.830 | 6.940E-10 |
| HDL cholesterol | Maximum likelihood | 85 | 0.741 | 0.671 | 0.817 | 2.243E-09 |
| HDL cholesterol | Weighted median | 85 | 1.053 | 0.908 | 1.222 | 4.913E-01 |
| Fasting insulin | IVW fixed effects | 13 | 1.498 | 0.864 | 2.596 | 1.497E-01 |
| Fasting insulin | Maximum likelihood | 13 | 1.514 | 0.869 | 2.637 | 1.431E-01 |
| Fasting insulin | Weighted median | 13 | 1.868 | 0.921 | 3.787 | 8.318E-02 |
| Fasting glucose | IVW fixed effects | 31 | 1.297 | 1.013 | 1.661 | 3.908E-02 |
| Fasting glucose | Maximum likelihood | 31 | 1.304 | 1.016 | 1.673 | 3.679E-02 |
| Fasting glucose | Weighted median | 31 | 1.351 | 0.929 | 1.965 | 1.153E-01 |
| Current tobacco smoking | IVW fixed effects | 27 | 0.737 | 0.321 | 1.692 | 4.717E-01 |
| Current tobacco smoking | Maximum likelihood | 27 | 0.730 | 0.311 | 1.711 | 4.685E-01 |
| Current tobacco smoking | Weighted median | 27 | 0.599 | 0.161 | 2.228 | 4.449E-01 |
| BMI | IVW fixed effects | 360 | 0.904 | 0.815 | 1.002 | 5.524E-02 |
| BMI | Maximum likelihood | 360 | 0.904 | 0.811 | 1.007 | 6.764E-02 |
| BMI | Weighted median | 360 | 1.033 | 0.842 | 1.267 | 7.555E-01 |
Sensitivity analysis
Consistent with standard IVW results, MLM and weighted median results also showed a significant association between years of schooling and AD (Table 2). In the analysis of weighted median, genetically determined increasing HDL cholesterol level was associated with decreased AD risk (Table 2). Similar significant association was observed between total cholesterol with AD by MLM approach. Similar to MR main results, MLM approach also found a suggestive association between FG and AD. We still did detect any association between the rest risk factors and AD. Furthermore, MR Egger test suggested that there was no pleiotropic effect among the selected IVs for each risk factor (Table 3).
Table 3. Pleiotropic test for the selected instrumental variables.
| Exposure | Egger_intercept | se | pvalue |
| Type 2 diabetes | -8.337E-03 | 0.006 | 0.186 |
| Years of schooling | 4.781E-03 | 0.005 | 0.327 |
| HDL cholesterol | 2.795E-03 | 0.019 | 0.885 |
| LDL cholesterol | -1.158E-02 | 0.009 | 0.215 |
| Total cholesterol | -1.663E-02 | 0.011 | 0.131 |
| Fasting glucose | 5.334E-04 | 0.009 | 0.955 |
| Fasting insulin | -4.983E-03 | 0.026 | 0.851 |
| Body mass index (BMI) | 9.387E-03 | 0.007 | 0.202 |
| Current tobacco smoking | 6.280E-03 | 0.019 | 0.746 |
For the bi-directional MR analysis, 16 LD independent SNPs that reached genome-wide significance were selected as IVs for AD, then those SNPs were extracted from the nine outcomes individually. After data harmonization, number of valid IVs left for each outcome was demonstrated in Table 4. The results showed significant association between AD and BMI (OR ranging from 0.980 to 0.995, P = 0.002), and borderline association between AD and T2D (OR = 1.047, 95 CI: 1.000-1.096, P = 0.049, Table 4). And MR Egger intercept suggested no existence of pleiotropic among selected IVs. Besides, as a complementary approach, Steiger test results also showed that the variances explained in the exposuresa were larger than that in the outcome (AD), and the causal direction turned out to be TRUE (Table 5).
Table 4. Bi-directional MR results: Alzheimer’s disease as exposure.
| Outcome | Methods | nsnp | OR | OR_lci95 | OR_uci95 | pvalue |
| BMI | IVW fixed effects | 13 | 0.988 | 0.980 | 0.995 | 0.002 |
| BMI | Maximum likelihood | 13 | 0.987 | 0.979 | 0.995 | 0.001 |
| BMI | Weighted median | 13 | 0.988 | 0.974 | 1.002 | 0.093 |
| Current tobacco smoking | IVW fixed effects | 13 | 1.000 | 0.995 | 1.004 | 0.864 |
| Current tobacco smoking | Maximum likelihood | 13 | 1.000 | 0.995 | 1.004 | 0.861 |
| Current tobacco smoking | Weighted median | 13 | 1.000 | 0.993 | 1.006 | 0.889 |
| HDL cholesterol | IVW fixed effects | 7 | 0.984 | 0.952 | 1.016 | 0.324 |
| HDL cholesterol | Maximum likelihood | 7 | 0.983 | 0.950 | 1.016 | 0.312 |
| HDL cholesterol | Weighted median | 7 | 0.983 | 0.940 | 1.028 | 0.454 |
| LDL cholesterol | IVW fixed effects | 7 | 1.001 | 0.966 | 1.038 | 0.949 |
| LDL cholesterol | Maximum likelihood | 7 | 1.001 | 0.966 | 1.038 | 0.948 |
| LDL cholesterol | Weighted median | 7 | 1.014 | 0.968 | 1.062 | 0.567 |
| Total cholesterol | IVW fixed effects | 7 | 0.988 | 0.954 | 1.023 | 0.489 |
| Total cholesterol | Maximum likelihood | 7 | 0.988 | 0.953 | 1.023 | 0.489 |
| Total cholesterol | Weighted median | 7 | 0.987 | 0.945 | 1.031 | 0.557 |
| Type 2 diabetes | IVW fixed effects | 7 | 1.047 | 1.000 | 1.096 | 0.049 |
| Type 2 diabetes | Maximum likelihood | 7 | 1.049 | 1.001 | 1.099 | 0.044 |
| Type 2 diabetes | Weighted median | 7 | 1.011 | 0.948 | 1.078 | 0.735 |
| Years of schooling | IVW fixed effects | 13 | 0.999 | 0.992 | 1.006 | 0.828 |
| Years of schooling | Maximum likelihood | 13 | 0.999 | 0.992 | 1.006 | 0.821 |
| Years of schooling | Weighted median | 13 | 1.008 | 0.999 | 1.017 | 0.090 |
Table 5. MR Steiger directionality test results.
| Exposure | snp_r2.exposure | snp_r2.outcome | Correct_causal_direction | Steiger_pvalue |
| Type 2 diabetes | 0.124 | 0.003 | TRUE | 0 |
| Years of schooling | 0.019 | 0.007 | TRUE | 2.09E-34 |
| HDL cholesterol | 0.053 | 0.020 | TRUE | 1.96E-79 |
| LDL cholesterol | 0.058 | 0.008 | TRUE | 4.38E-212 |
| Total cholesterol | 0.064 | 0.013 | TRUE | 2.09E-191 |
| Fasting glucose | 0.032 | 0.001 | TRUE | 2.13E-182 |
| Fasting insulin | 0.005 | 0.0002 | TRUE | 1.81E-26 |
| Body mass index (BMI) | 0.058 | 0.027 | TRUE | 7.22E-71 |
| Current tobacco smoking | 0.003 | 0.001 | TRUE | 9.74E-06 |
MR-BMA estimates results
All the risk factors were then prioritized and ranked by their MIP. The top five risk factors for AD were current tobacco smoking (MIP = 0.289), BMI (MIP = 0.210), FI (MIP = 0.164), FG (MIP = 0.163), years of schooling (MIP = 0.161), and which were confirmed to be included in the best five individual models with their PP values of 0.207, 0.138, 0.107, 0.112 and 0.111 respectively (Table 6). The MR-BMA were partially consistent with mvMR results, MR-BMA method prioritized two of the significant risk factors (FG and years of schooling) in mvMR result.
Table 6. Ranking of risk factors for AD. A) According to their marginal inclusion probability (MIP).
| Risk factors combination | Marginal inclusion probability (MIP) | Model averaged causal estimate | |
| 1 | Current tobacco smoking | 0.289 | 8.582E-02 |
| 2 | BMI | 0.210 | -4.085E-02 |
| 3 | Fasting insulin | 0.164 | 2.502E-02 |
| 4 | Fasting glucose | 0.163 | 3.087E-02 |
| 5 | Years of schooling | 0.161 | 1.195E-02 |
| 6 | LDL cholesterol | 0.093 | 9.312E-03 |
| 7 | Total cholesterol | 0.062 | 4.005E-03 |
| 8 | HDL cholesterol | 0.036 | -5.559E-04 |
| 9 | T2D | 0.024 | 3.122E-04 |
B). The best ten individual models according to their posterior probability (PP).
| Individual models | Risk factors combination | Posterior probability |
| 9 | Current tobacco smoking | 0.207 |
| 8 | BMI | 0.138 |
| 6 | Fasting glucose | 0.112 |
| 2 | Years of schooling | 0.111 |
| 7 | Fasting insulin | 0.107 |
| 4 | LDL cholesterol | 0.062 |
| 5 | Total cholesterol | 0.041 |
| 3 | HDL cholesterol | 0.024 |
| 8,9 | BMI, current tobacco smoking | 0.018 |
| 1 | T2D | 0.015 |
Discussion
In the present study, by performing mvMR and MR-BMA analysis together using summary statistics for AD and multiple risk factors, we successfully identified five causal risk factor (years of schooling, total cholesterol, HDL cholesterol, LDL cholesterol and FG) for AD and we also prioritized and ranked two of these five risk factors (FG and years of schooling) for AD, which might provide us novel insights into determine the causal risk factors for complex traits and diseases.
Our results are consistent with both previous traditional observational studies and two sample MR results which provide established evidence that educational attainment was associated with a reduced risk of AD [21–23]. Concentrations of genetically determined total cholesterol and LDL cholesterol showed positive associations with risks of AD, which is consistent with the known causal effect of them on AD risk from previous two sample MR study [24], and similar to our results, elevated HDL cholesterol level also showed causal association with decreased AD risk [24]. The suggestive association between FG and higher AD risk is also consistent with previous study that FG showed suggestive association with AD risk in sensitivity analysis [24]. No observed causal associations between fasting insulin, BMI and AD risk are also consistent with previous two sample MR result [24].
Although our mvMR did not find any causal association between current tobacco smoking with risk of AD, current tobacco smoking became the top first risk factor in the MR-BMA model, which is partially in accordance with previous MR study [24].
However, the observed associations differ from previous study that no evidence of causal association between lipid profiles and AD risk after excluding the potential pleiotropic SNPs. In the current study, the independent SNPs (r2 < 0.001) we included suggest no evidence of pleiotropic effect and our sensitivity analysis support the causal associations detected by the main MR analysis. One of the most interesting findings from the present study is the absence causal association of BMI and AD risk, however the bi-directional MR found causal association between AD and decreased BMI. This finding supports the hypothesis of reverse causation (the negative confounding effect of AD related weight loss) might be an explanation for the obesity paradox on AD risk [4].
There are several important strengths to note for the current study. Our MR analysis results may provide evidence of the causal role of five risk factors in the development of AD since the influence of traditional confounding factors in observational studies is minimized/eliminated. Since the alleles follow the principle of random distribution when forming gametes at meiosis, the causal effect of genotype on disease in MR studies will not be distorted by the confounding factors, a major limitation of traditional observational studies. By leveraging the summary statistics from the large available GWASs for multiple risk factors and AD, we were able to increase our discovery power. Furthermore, previous studies have shown that performing the MR analysis by using summary statistics data and by using individual-level data have similar efficiency [19]. Finally, the application of MR-BMA ranked the potential risk factors for AD, which could provide certain genetic evidence in disease prevention and curation.
However, there may also be some limitations. First, we included the mixed population data for lipid traits instead of European only, because the dataset for lipid we included in our analysis was the largest to date, and the individuals from other ethnicities only take up 4% (Table 1). Additionally, our MR results does not mean the potentially causal risk factors identified are playing truly causal roles in the AD susceptibility, we were trying to provide some novel insights into underlying mechanisms of the AD and hope to provide certain genetical evidence to disease prevention.
Conclusions
In conclusion, by combining mvMR and MR-BMA together, we successfully identified five potential causal risk factors for AD and we also ranked and prioritized two of them for AD, which might provide us novel insights into the causal mechanisms of AD. Our results demonstrate that by increasing the years of schooling and HDL cholesterol level, decreasing total cholesterol, LDL cholesterol and FG levels could decrease the risk of developing AD.
Materials and Methods
Genetic IVs selection and validation
Summary statistics for risk factor-associated SNPs were extracted from the large publicly available GWAS datasets to date performed by the corresponding Consortia in European populations (Table 1). For the implementation of mvMR, we selected SNPs that achieved genome-wide significance (p < 5 × 10−8) in the GWAS datasets as for each risk factor as IVs. Effect estimates of these risk factor-associated SNPs on the risk of AD were assessed using the summary statistics of 74,046 European individuals for AD from The International Genomics of Alzheimer's Project (IGAP) Consortium [15]. The European samples from the 1000 genomes project reference panel were adopted to estimate linkage disequilibrium (LD) between chosen SNPs. When target SNPs were not available in the outcome study, we used proxy SNPs that were in high LD (r2 > 0.8) with the SNPs of interest.
To ensure the SNPs used as IVs for risk factors are not in LD with each other, a vital assumption of MR, we calculated pairwise-LD between all our selected SNPs in the 1000 Genomes European reference sample using PLINK 1.90 [16]. For all pairs of SNPs determined to violate the independence assumption with r2 > 0.001 we retained only the SNP with the smaller association p-value. To ensure the effect of a SNP on the exposure and the effect of that SNP on the outcome correspond to the same allele, we harmonized the effect of these instrumental SNPs by using a function that ensures all corresponding risk factors and outcome (AD) alleles are on the same strand where possible. If they are not, then the function will flip alleles and use allele frequency to infer the strand of palindromic SNPs.
mvMR estimates
In the current study, standard inverse-variance-weighted (IVW) fixed-effects [17] analysis was used to estimate the causal effect of the multiple related risk factors on the BMD traits. After obtaining the selected instruments for each exposure, all exposures for those SNPs were then regressed against the outcome together, weighting for the inverse variance of the outcome to ensure the genetic instruments with more precise association receive more weight in our analysis.
Sensitivity analysis
As sensitivity analysis, weighted median estimator and maximum likelihood method (MLM) [18] were also performed to provide more robust MR estimates. MR-Egger approach [19] was also used to assess the potential pleiotropic effects among the selected IVs. A Bonferroni corrected threshold with P < 0.006 (0.05/9) was considered to be significant causal association, and 0.006 < P < 0.05 was considered suggestive evidence for causal association. Besides, to orient the causal relationship between them, we also performed the bi-directional MR analysis (P < 0.05) and MR Steiger directionality test [20].
MR-BMA estimates
Following the mvMR analysis, MR-BMA was applied to prioritize the most causally related risk factors for AD. MR-BMA assumes that the true causal risk factors are very few and it considers the risk factor selection as a variable selection problem in the linear regression model. The approach considers all possible combinations of the risk factors and generates posterior probability (PP) for each specific model, where PP means the probability of including a specific risk factor in the model. Furthermore, MR-BMA adopts BMA which computes a marginal inclusion probability (MIP) for each risk factor, where MIP refers to the sum of the PP over all possible models where the risk factor is present. Then MR-BMA will compute the model-averaged causal estimate (MACE) for each risk factor by ranking all the risk factors according to the corresponding MIP. Finally, MR-BMA will prioritize the best model by the PP value for each individual model. All the analyses were implemented in R software environment.
Availability of data and materials
The datasets generated and/or analyzed during the current study are included in this published article and provided in Supplementary Table 1 of Supplementary Materials.
Supplementary Materials
Author Contributions
QZ as the first author performed data analysis and wrote the manuscript. FX, LW and WDZ contributed suggestions for manuscript revision and revised the manuscript. CQS and HWD conceived and initiated this project, provided advice on experimental design, oversaw the implementation of the statistical method, and revised/finalized the manuscript.
Acknowledgments
We thank all the consortia (included in Supplementary Table 1) for providing the GWAS datasets available to the public.
Conflicts of Interest
The authors Qiang Zhang, Fei Xu, Lianke Wang, Wei-Dong Zhang, Chang-Qing Sun, Hong-Wen Deng declare they have no conflicts of interest.
Funding
CQS was partially supported by Key Science and Technology Development of Henan Province (Grant NO: 192102310191). CQS was also supported by The Education Department of Henan Province (GRANT NO: 2019-YYZD-17). HWD was partially supported by grants from the NIH (R01-AR069055, U19-AG055373, R01-MH104680, R01-AR059781, and P20-GM109036) and Edward G. Schlieder Endowment fund at Tulane University. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Ballard C, Gauthier S, Corbett A, Brayne C, Aarsland D, Jones E. Alzheimer’s disease. Lancet. 2011; 377:1019–31. https://doi.org/10.1016/S0140-6736(10)61349-9 [PubMed]
- 2. Scheltens P, Blennow K, Breteler MM, de Strooper B, Frisoni GB, Salloway S, Van der Flier WM. Alzheimer’s disease. Lancet. 2016; 388:505–17. https://doi.org/10.1016/S0140-6736(15)01124-1 [PubMed]
- 3. Jansen IE, Savage JE, Watanabe K, Bryois J, Williams DM, Steinberg S, Sealock J, Karlsson IK, Hägg S, Athanasiu L, Voyle N, Proitsi P, Witoelar A, et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat Genet. 2019; 51:404–13. https://doi.org/10.1038/s41588-018-0311-9 [PubMed]
- 4. Pegueroles J, Jiménez A, Vilaplana E, Montal V, Carmona-Iragui M, Pané A, Alcolea D, Videla L, Casajoana A, Clarimón J, Ortega E, Vidal J, Blesa R, et al, and Alzheimer’s Disease Neuroimaging Initiative. Obesity and Alzheimer’s disease, does the obesity paradox really exist? a magnetic resonance imaging study. Oncotarget. 2018; 9:34691–98. https://doi.org/10.18632/oncotarget.26162 [PubMed]
- 5. Anstey KJ, Cherbuin N, Herath PM. Development of a new method for assessing global risk of Alzheimer’s disease for use in population health approaches to prevention. Prev Sci. 2013; 14:411–21. https://doi.org/10.1007/s11121-012-0313-2 [PubMed]
- 6. Jayaraman A, Pike CJ. Alzheimer’s disease and type 2 diabetes: multiple mechanisms contribute to interactions. Curr Diab Rep. 2014; 14:476. https://doi.org/10.1007/s11892-014-0476-2 [PubMed]
- 7. Vagelatos NT, Eslick GD. Type 2 diabetes as a risk factor for Alzheimer’s disease: the confounders, interactions, and neuropathology associated with this relationship. Epidemiol Rev. 2013; 35:152–60. https://doi.org/10.1093/epirev/mxs012 [PubMed]
- 8. Xu W, Tan L, Wang HF, Jiang T, Tan MS, Tan L, Zhao QF, Li JQ, Wang J, Yu JT. Meta-analysis of modifiable risk factors for Alzheimer’s disease. J Neurol Neurosurg Psychiatry. 2015; 86:1299–306. https://doi.org/10.1136/jnnp-2015-310548 [PubMed]
- 9. Smith GD, Ebrahim S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int J Epidemiol. 2003; 32:1–22. https://doi.org/10.1093/ije/dyg070 [PubMed]
- 10. Larsson SC, Traylor M, Malik R, Dichgans M, Burgess S, Markus HS, and CoSTREAM Consortium, on behalf of the International Genomics of Alzheimer’s Project. Modifiable pathways in Alzheimer’s disease: Mendelian randomisation analysis. BMJ. 2017; 359:j5375. https://doi.org/10.1136/bmj.j5375 [PubMed]
- 11. Burgess S, Thompson SG. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am J Epidemiol. 2015; 181:251–60. https://doi.org/10.1093/aje/kwu283 [PubMed]
- 12. Hemani G, Bowden J, Davey Smith G. Evaluating the potential role of pleiotropy in Mendelian randomization studies. Hum Mol Genet. 2018; 27:R195–208. https://doi.org/10.1093/hmg/ddy163 [PubMed]
- 13. Burgess S, Freitag DF, Khan H, Gorman DN, Thompson SG. Using multivariable Mendelian randomization to disentangle the causal effects of lipid fractions. PLoS One. 2014; 9:e108891. https://doi.org/10.1371/journal.pone.0108891 [PubMed]
- 14. Zuber V, Colijn JM, Klaver C, Burgess S. Selecting causal risk factors from high-throughput experiments using multivariable Mendelian randomization. bioRxiv. 2019; 396333. https://doi.org/10.1101/396333
- 15. Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, DeStafano AL, Bis JC, Beecham GW, Grenier-Boley B, Russo G, Thorton-Wells TA, Jones N, et al, and European Alzheimer’s Disease Initiative (EADI), and Genetic and Environmental Risk in Alzheimer’s Disease, and Alzheimer’s Disease Genetic Consortium, and Cohorts for Heart and Aging Research in Genomic Epidemiology. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013; 45:1452–58. https://doi.org/10.1038/ng.2802 [PubMed]
- 16. Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007; 81:559–75. https://doi.org/10.1086/519795 [PubMed]
- 17. Bowden J, Davey Smith G, Haycock PC, Burgess S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016; 40:304–14. https://doi.org/10.1002/gepi.21965 [PubMed]
- 18. Burgess S, Butterworth A, Thompson SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013; 37:658–65. https://doi.org/10.1002/gepi.21758 [PubMed]
- 19. Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through egger regression. Int J Epidemiol. 2015; 44:512–25. https://doi.org/10.1093/ije/dyv080 [PubMed]
- 20. Hemani G, Tilling K, Davey Smith G. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet. 2017; 13:e1007081. https://doi.org/10.1371/journal.pgen.1007081 [PubMed]
- 21. Bandres-Ciga S, Faghri F. Unraveling the genetic complexity of Alzheimer disease with Mendelian randomization. Neurol Genet. 2019; 5:e313. https://doi.org/10.1212/NXG.0000000000000313 [PubMed]
- 22. Du L, Hu J. The effects of health education on knowledge about Alzheimer’s disease and health-promoting behaviours of older Chinese adults in a nursing home: a pilot study. Int J Nurs Pract. 2016; 22:31–42. https://doi.org/10.1111/ijn.12349 [PubMed]
- 23. Hendrie HC, Smith-Gamble V, Lane KA, Purnell C, Clark DO, Gao S. The association of early life factors and declining incidence rates of dementia in an elderly population of African Americans. J Gerontol B Psychol Sci Soc Sci. 2018; 73:S82–89. https://doi.org/10.1093/geronb/gbx143 [PubMed]
- 24. Østergaard SD, Mukherjee S, Sharp SJ, Proitsi P, Lotta LA, Day F, Perry JR, Boehme KL, Walter S, Kauwe JS, Gibbons LE, Larson EB, Powell JF, et al, and Alzheimer’s Disease Genetics Consortium, and GERAD1 Consortium, and EPIC-InterAct Consortium. Associations between potentially modifiable risk factors and Alzheimer disease: a Mendelian randomization study. PLoS Med. 2015; 12:e1001841. https://doi.org/10.1371/journal.pmed.1001841 [PubMed]



