Introduction
Cervical cancer is a common malignant tumor that threatens the health and life of the female population. It has been reported that the morbidity and mortality of cervical cancer both rank fourth among malignant tumors worldwide [1]. In China, the incidence of cervical cancer is increasing year by year [2]. Although multiple therapies such as surgery, chemotherapy and radiotherapy, have been widely applied to treat cervical cancer patients, the overall efficiency and prognosis are still unsatisfactory, owing to the lack of systematic understanding of the underlying mechanism. Therefore, it is necessary to elucidate the exact mechanism to develop targeted drugs.
Plenty of evidence has manifested that long non-coding RNAs (lncRNAs), possessing no capacity to encode functional proteins, function as crucial regulators in pathogenesis of cervical cancer by epigenetically modulating targeted molecules [3, 4]. Generally, lncRNAs act as the oncogenes in development and progression of cervical cancer. LncRNA NEAT1, a well-known oncogenic lncRNA, contributes to the increased proliferative and invasive abilities of cervical cancer cells [5]. Recent research reveals that the overexpressed lncRNA SOX2OT facilitates malignant phenotypes of cervical cancer cells by evaluating the proliferative, migratory and invasive abilities [6]. However, some other lncRNAs exert inhibitory effects on cervical cancer cells. Wang et al. have reported that cervical cancer expresses lower level of lncRNA GAS5-AS1, and lncRNA GAS5-AS1 overexpression inhibits degradation of GAS5 by methyl adenosine modification to suppress development of cervical cancer [7]. In addition, decreased expression of lncRNA miR503HG has been shown in cervical cancer, and the in vitro assays indicate lncRNA miR503HG acts as a sponge of miR-155 to induce apoptosis and increase sensitivity of cervical cancer cells to cisplatin [8]. Therefore, lncRNAs function as the double-edged swords in pathogenesis of cervical cancer. Recently, researchers concentrate on the pro-tumor effects of lncRNA SNHG7 in malignant tumors [9]. In addition, overexpression of lncRNA SNHG7 exists in cervical cancer [10]. However, the exact mechanism of lncRNA SNHG7 in cervical cancer needs to be fully elucidated.
Notably, involvement of microRNAs (miRNAs) as downstream molecule of lncRNA SNHG7 is a key point to regulate biological function of cancer cells [9]. In breast cancer, the increased lncRNA SNHG7 could sponge miR-34a to induce epithelial-mesenchymal transition (EMT) and augment proliferative and invasive abilities [11]. In addition, lncRNA SNHG7 small interfering RNA (siRNA) abolished its inhibitory role in miR-140-5p to impair proliferation of nasopharyngeal carcinoma cells [12]. Currently, miR-122-5p was screened as a downstream molecule of lncRNA SNHG7 by using the online database (http://starbase.sysu.edu.cn/panCancer.php), and we aimed to explore contribution of lncRNA SNHG7/miR-122-5p in pathogenesis of cervical cancer.
Results
Aberrant expression of lncRNA SNHG7 and miR-122-5p existed in cervical cancer
Firstly, upregulation of lncRNA SNHG7 and downregulation of miR-122-5p were found in cervical cancer tissues compared with that in tissues adjacent to cervical cancer (Figure 1A and 1B). Moreover, cisplatin-resistant cervical cancer tissues displayed a dramatically higher expression of lncRNA SNHG7 and a lower expression of miR-122-5p compared with cervical cancer tissues (Figure 1C and 1D). We also found the negative relationship between the expression of lncRNA SNHG7 and the expression of miR-122-5p (Figure 1E). In addition, the clinicopathological data revealed that upregulation of lncRNA SNHG7 was positively correlated to the tumor size, lymphatic metastasis and the TNM staging (Table 1). Meanwhile, we identified lncRNA SNHG7 was elevated and miR-122-5p was reduced in cervical cancer cells lines (Figure 1F and 1G)., and we selected HeLa and SiHa in the subsequent analysis due to the expression of lncRNA SNHG7 and miR-122-5p. As was shown in Figure 1H, lncRNA SNHG7 was upregulated in HeLa and CR-HeLa cells, and CR-HeLa cells possessed a higher level of lncRNA SNHG7 than HeLa cells. In addition, the decreased miR-122-5p existed in HeLa and CR-HeLa cells, and CR-HeLa cells displayed a lower level of miR-122-5p in comparison with HeLa cells (Figure 1I).
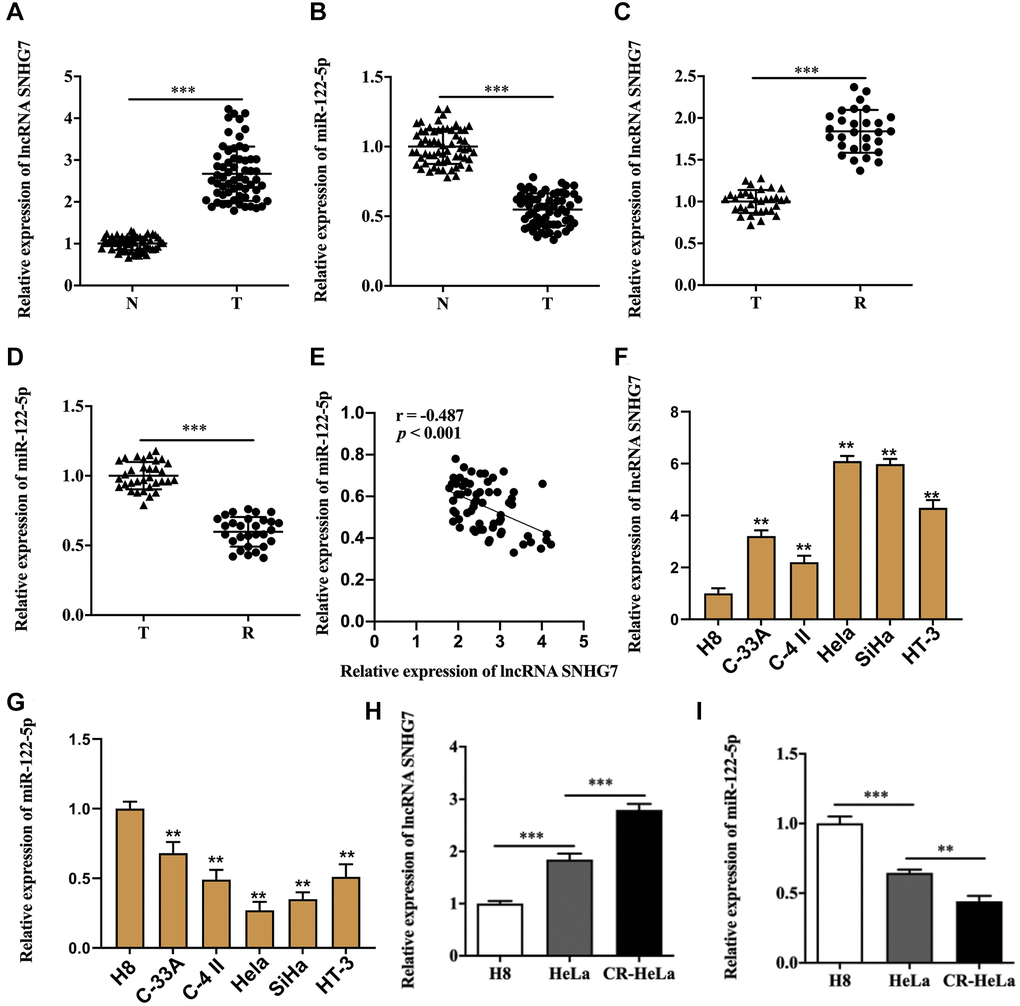
Figure 1. Aberrant expression of lncRNA SNHG7 and miR-122-5p existed in cervical cancer. (A and B) Relative expression of lncRNA SNHG7and miR-122-5p in cervical cancer tissues and non-tumor tissues adjacent to cervical cancer. (C and D) Relative expression of lncRNA SNHG7 and miR-122-5p in cervical cancer tissues and cisplatin-resistant cervical cancer tissues. (E) The correlation between lncRNA SNHG7 and miR-122-5p in clinical samples. (F and G) Relative expression of lncRNA SNHG7 and miR-122-5p in human cervical epithelial cell line H8 cells and human cervical cancer cell lines. (H and I) Relative expression of lncRNA SNHG7 and miR-122-5p in human cervical epithelial cell line H8 cells, human cervical cancer cell line HeLa cells and cisplatin-resistant HeLa cells. N, non-tumor tissues adjacent to cervical cancer; T, cervical cancer tissues; R, cisplatin-resistant cervical cancer tissues. CR-HeLa, cisplatin-resistant HeLa cells. **p < 0.01, ***p < 0.001.
Table 1. The characteristics of patients with cervical cancer.
| Characteristic | lncRNA SNHG7 low | lncRNA SNHG7 high | p value |
| Age (year) | 0.961 | ||
| < 45 | 15 | 13 | |
| ≥ 45 | 18 | 16 | |
| Tumor diameter (cm) | 0.036 | ||
| < 3 | 19 | 9 | |
| ≥ 3 | 14 | 20 | |
| Lymphatic metastasis | 0.02 | ||
| No | 23 | 9 | |
| Yes | 13 | 20 | |
| TNM staging | 0.022 | ||
| I~II | 21 | 10 | |
| III~IV | 12 | 19 |
LncRNA SNHG7 acted upstream of miR-122-5p
ENCORI database suggested the potential binding sites between lncRNA SNHG7 and miR-122-5p (Figure 2A). Luciferase reporter assay showed that HeLa and SiHa cells co-transfected with wild type lncRNA SNHG7 and miR-122-5p mimics displayed the weakened luciferase activity compared with that co-transfected with wild type lncRNA SNHG7 and mimics NC (Figure 2A and Supplementary Figure 1A). To further confirm the sponge role of lncRNA SNHG7, we performed cell transfection with lncRNA SNHG7 overexpression vector or lncRNA SNHG7 siRNA into HeLa and SiHa cells to assess the efficacy, and the results were shown in Figure 2B and 2C, and Supplementary Figure 1B and 1C. We found that lncRNA SNHG7 overexpression suppressed the expression of miR-122-5p (Figure 2D and Supplementary Figure 1D). In contrast, lncRNA SNHG7 siRNA facilitated miR-122-5p expression (Figure 2E and Supplementary Figure 1E). The above findings indicated that miR-122-5p was sponged by lncRNA SNHG7.
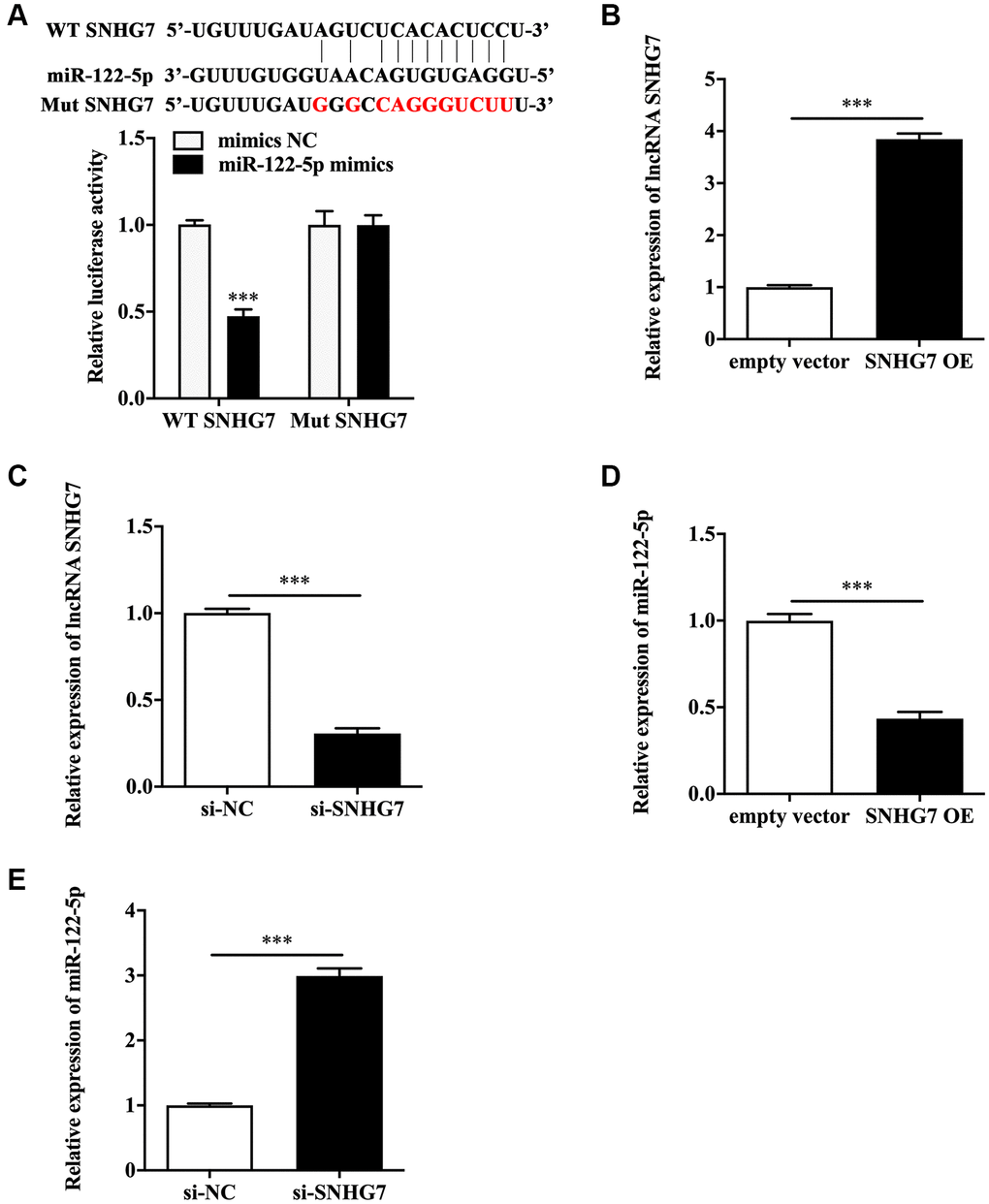
Figure 2. LncRNA SNHG7 acted upstream of miR-122-5p. (A) The schematic diagram of binding sites between lncRNA SNHG7 and miR-122-5p, and the binding capacity was evaluated by luciferase reporter assay. (B and C) Relative expression of lncRNA SNHG7 in HeLa cells with SNHG7 OE or si-SNHG7. (D and E) Relative expression of miR-122-5p in HeLa cells with SNHG7 OE or si-SNHG7. WT SNHG7, wild type lncRNA SNHG7; Mut SNHG7, mutant type lncRNA SNHG7; mimics NC, negative control mimics; SNHG7 OE, lncRNA SNHG7 overexpression; si-NC, negative control siRNA; si-SNHG7, lncRNA SNHG7 siRNA. ***p < 0.001.
MiR-122-5p antagonized the biological roles of lncRNA SNHG7 in promoting malignant behaviors of HeLa and SiHa cells
To figure out the biological effects of lncRNA SNHG7 on cervical cancer, we performed the following assays.
CCK-8 assay showed that lncRNA SNHG7 overexpression increased the IC50 value of HeLa and SiHa cells (Figure 3A and Supplementary Figure 2A). Colony formation assay revealed that lncRNA SNHG7 overexpression promoted the proliferative ability (Figure 3B and Supplementary Figure 2B). Wound healing assay depicted that lncRNA SNHG7 overexpression facilitated the migratory ability (Figure 3C and Supplementary Figure 2C). Moreover, transwell assay showed that lncRNA SNHG7 overexpression led to the increased invasive ability (Figure 3D and Supplementary Figure 2D). These results demonstrated lncRNA SNHG7 exerted a positive role in boosting chemoresistance, proliferation, migration and invasion of HeLa and SiHa cells. On the contrary, miR-122-5p mimics partially reversed the increased IC50 value (Figure 3A and Supplementary Figure 2A), and proliferative (Figure 3B and Supplementary Figure 2B), migratory (Figure 3C and Supplementary Figure 2C) and invasive (Figure 3D and Supplementary Figure 2D) abilities of HeLa and SiHa cells in the presence of lncRNA SNHG7 overexpression plasmid.
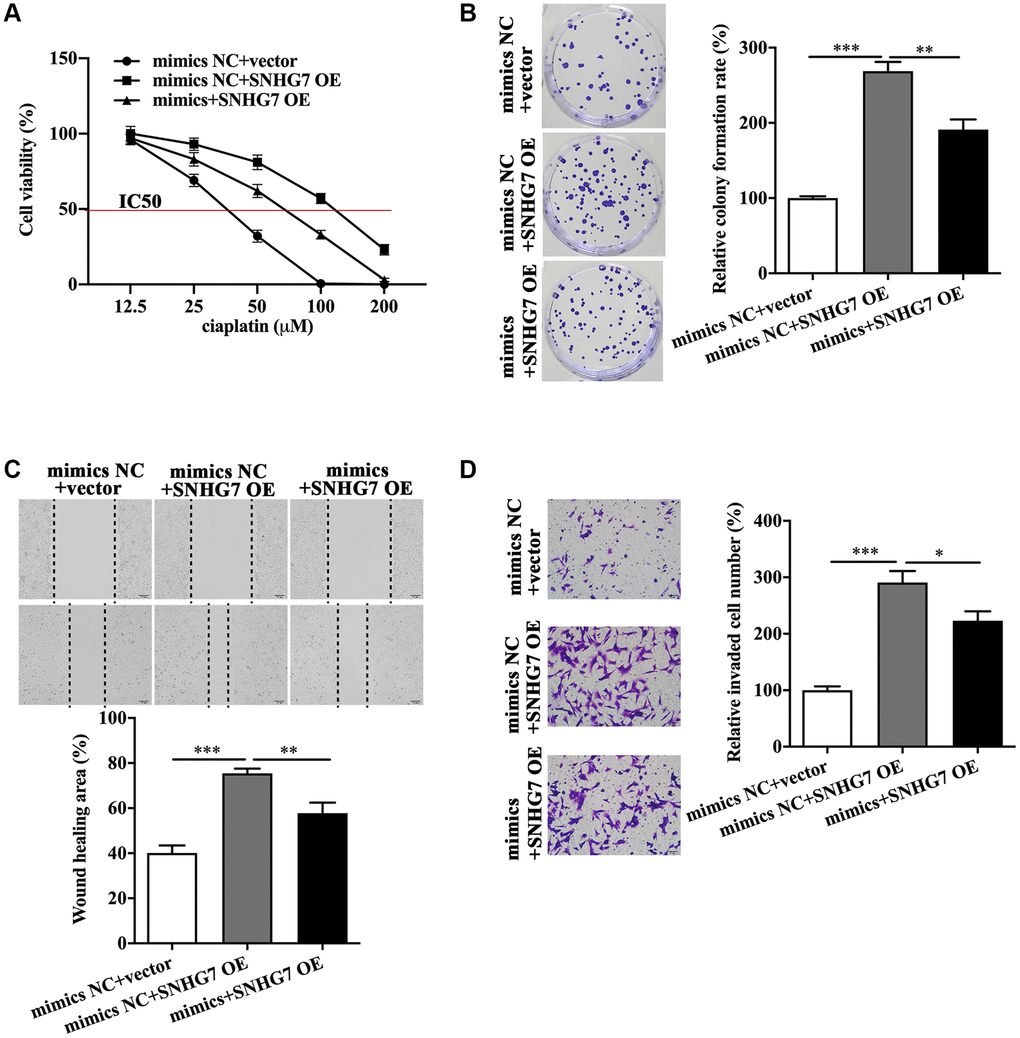
Figure 3. MiR-122-5p antagonized the biological roles of lncRNA SNHG7 in promoting malignant behaviors of HeLa cells. IC50 value (A), colony formation (B), migratory capacity (C), and invasive capacity (D) of HeLa cells in the presence or absence of SNHG7 OE and miR-122-5p mimics. mimics NC, negative control mimics; SNHG7 OE, lncRNA SNHG7 overexpression; mimics, miR-122-5p mimics; IC50, half maximal inhibitory concentration. *p < 0.05, **p < 0.01, ***p < 0.001.
FOXP1 was the downstream molecule of miR-122-5p
The miRBase and TargetScanHuman database were utilized to identify FOXP1 as the potential target of miR-122-5p (Figure 4A). Next, luciferase reporter assay showed that HeLa and SiHa cells co-transfected with wild type FOXP1 and miR-122-5p mimics exhibited a remarkably lower luciferase activity compared with that co-transfected with mutant type FOXP1 and mimics NC (Figure 4A and Supplementary Figure 3A). The miR-122-5p level was measured to confirm the efficiency of miR-122-5p mimics (Figure 4B and Supplementary Figure 3B) and miR-122-5p inhibitor (Figure 4C and Supplementary Figure 3C) in HeLa and SiHa cells. It was shown that miR-122-5p mimics inhibited FOXP1 (Figure 4D and Supplementary Figure 3D), but miR-122-5p inhibitor augmented FOXP1 mRNA level (Figure 4E and Supplementary Figure 3E). In addition, we found the upregulated FOXP1 in cervical cancer tissues (Figure 4F), and cisplatin-resistant cervical cancer tissues expressed more FOXP1 mRNA than cervical cancer tissues (Figure 4G). Western blot analysis showed that the protein level of FOXP1 was higher in cervical cancer tissues than that in non-tumor tissues adjacent to cervical cancer (Figure 4H). In addition, we found the expression of FOXP1 was negatively related to the expression of miR-122-5p, but positively related to the expression of lncRNA SNHG7 (Figure 4I and 4J). The qRT-PCR analysis showed that CR-HeLa cells expressed the highest mRNA level of FOXP1 among the three cell lines in our study (Figure 4K).
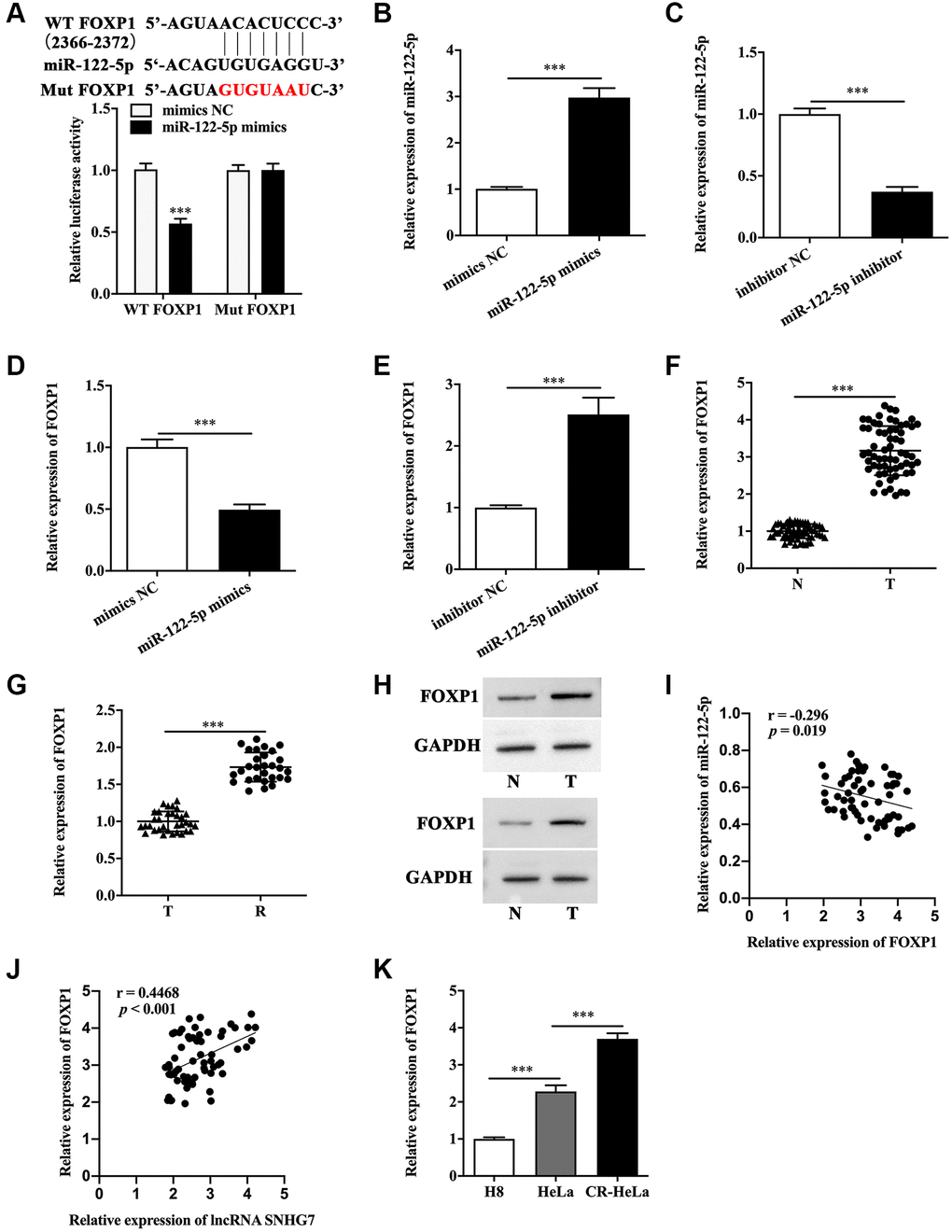
Figure 4. FOXP1 was the downstream molecule of miR-122-5p. (A) The schematic diagram of binding sites between FOXP1 and miR-122-5p, and the binding capacity was evaluated by luciferase reporter assay. (B and C) Relative expression of miR-122-5p in HeLa cells with miR-122-5p mimics transfection or miR-122-5p inhibitor transfection. (D and E) Relative expression of FOXP1 in HeLa cells with miR-122-5p mimics transfection or miR-122-5p inhibitor transfection. (F and G) Relative mRNA level of FOXP1 in non-tumor tissues adjacent to cervical cancer, cervical cancer tissues and cisplatin-resistant cervical cancer tissues. (H) The protein level of FOXP1 in non-tumor tissues adjacent to cervical cancer and cervical cancer tissues. (I) The correlation between FOXP1 and miR-122-5p or lncRNA SNHG7. (J and K) Relative mRNA level of FOXP1 in human cervical epithelial cell line H8 cells, human cervical cancer cell line HeLa cells and cisplatin-resistant HeLa cells. WT FOXP1, wild type FOXP1; Mut FOXP1, mutant type FOXP1; mimics NC, negative control mimics; inhibitor NC, negative control inhibitor; N, non-tumor tissues adjacent to cervical cancer; T, cervical cancer tissues; R, cisplatin-resistant cervical cancer tissues. CR-HeLa, cisplatin-resistant HeLa cells. ***p < 0.001.
FOXP1 overexpression rescued the biological roles of lncRNA SNHG7 knockdown in HeLa and SiHa cells
As FOXP1 was the direct downstream molecule of miR-122-5p, it was speculated that FOXP1 might function downstream of lncRNA SNHG7. The qRT-PCR analysis showed that lncRNA SNHG7 overexpression promoted the expression of FOXP1 (Figure 5A and Supplementary Figure 4A), which was inhibited by lncRNA SNHG7 siRNA (Figure 5B and Supplementary Figure 4B). In parallel, the western blot analysis showed the similar results (Figure 5C and Supplementary Figure 4C). We found that lncRNA SNHG7 siRNA-induced decreased IC50 value was reversed by FOXP1 overexpression (Figure 5D and Supplementary Figure 4D). It was demonstrated that lncRNA SNHG7 siRNA inhibited proliferative, migratory and invasive abilities (Figure 5E, 5F and 5G). Notably, FOXP1 overexpression reversed the inhibitory effects of lncRNA SNHG7 knockdown on malignant behaviors of HeLa and SiHa cells (Figure 5E, 5F and 5G, and Supplementary Figure 4E, 4F, 4G).
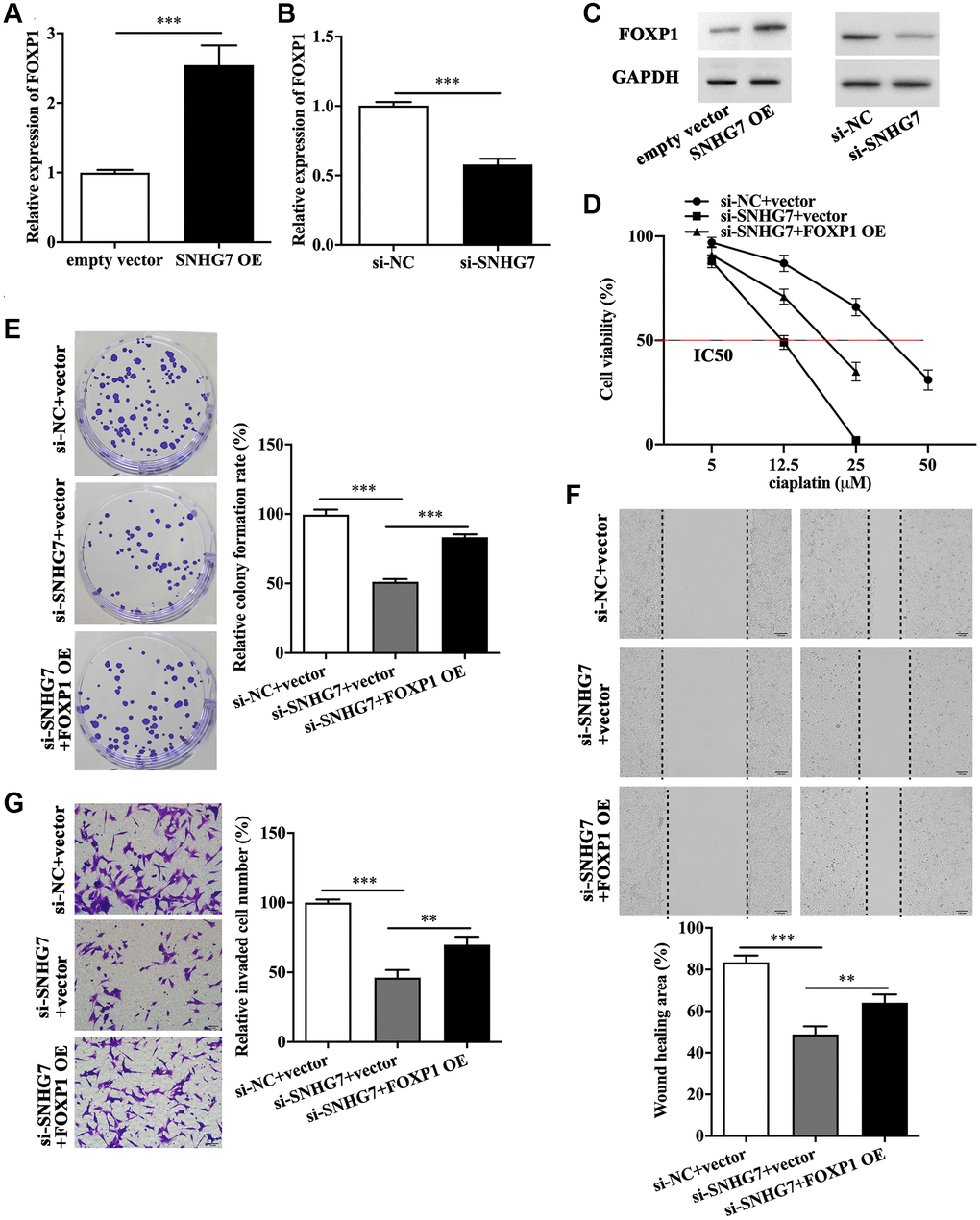
Figure 5. FOXP1 overexpression rescued the biological roles of lncRNA SNHG7 knockdown in HeLa cells. (A–C) Relative expression of FOXP1 in HeLa cells with SNHG7 OE and si-SNHG7 transfection. IC50 value (D), colony formation (E), migratory capacity (F), and invasive capacity (G) of HeLa cells in the presence or absence of si-SNHG7 and FOXP1 OE. SNHG7 OE, lncRNA SNHG7 overexpression; si-NC, negative control siRNA; si-SNHG7, lncRNA SNHG7 siRNA; FOXP1 OE, FOXP1 overexpression; IC50, half maximal inhibitory concentration. **p < 0.01, ***p < 0.001.
LncRNA SNHG7 knockdown suppressed cervical cancer growth in vivo
Finally, we performed animal experiments to investigate the effect of lncRNA SNHG7 in vivo. The efficacy of lncRNA SNHG7 shRNA transfection was evaluated by qRT-PCR (Figure 6A). As was represented in Figure 6B and 6C, the lumps isolated from mice injected with lncRNA SNHG7 shRNA-transfected HeLa cells presented the smaller size and lighter weight than that from the control group. The qRT-PCR analysis indicated that the lumps isolated from mice injected with lncRNA SNHG7 shRNA-transfected HeLa cells expressed less lncRNA SNHG7 (Figure 6D) and FOXP1 (Figure 6F), but more miR-122-5p (Figure 6E). Moreover, the protein level of FOXP1 was lower in the lumps isolated from mice injected with lncRNA SNHG7 shRNA-transfected HeLa cells (Figure 6G).
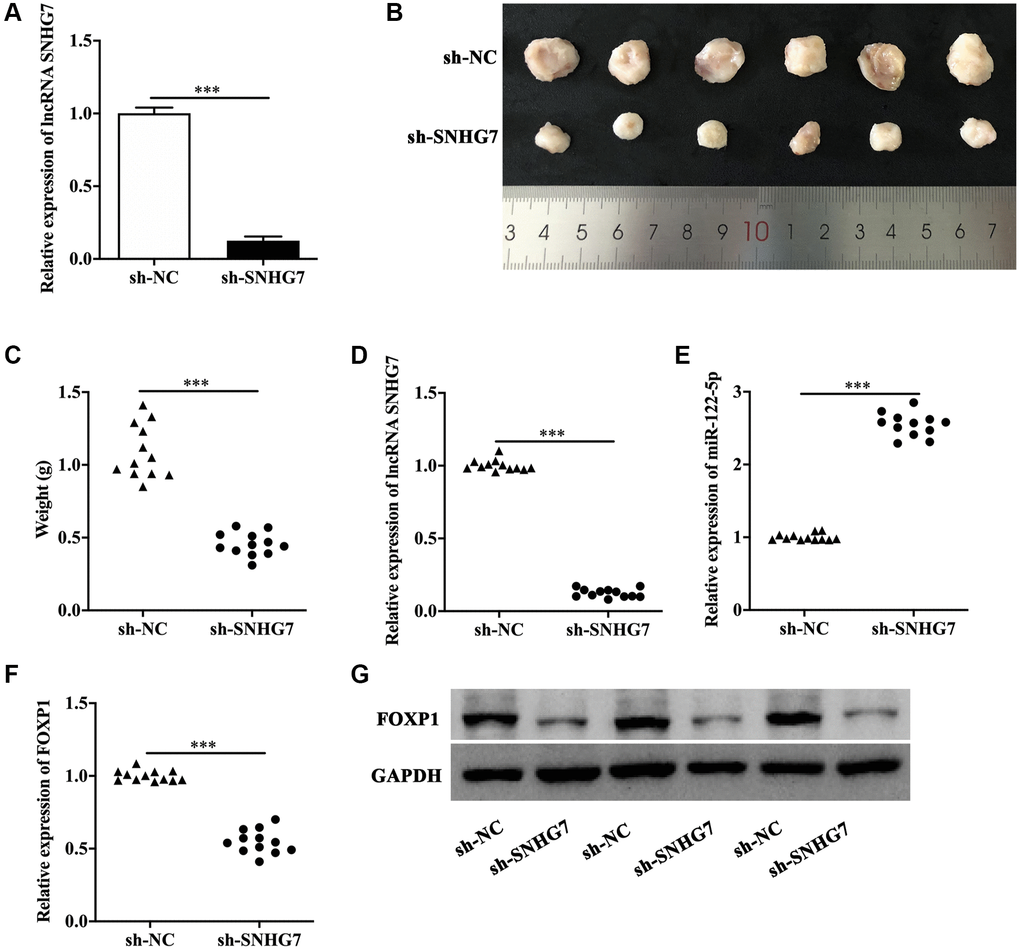
Figure 6. LncRNA SNHG7 knockdown suppressed cervical cancer growth in vivo. (A) Relative expression of lncRNA SNHG7 in HeLa cells transfected with sh-SNHG7. (B) The size of the lumps isolated from mice injected with negative control shRNA- or lncRNA SNHG7 shRNA-transfected HeLa cells. (C) The weight of the lumps isolated from mice injected with sh-NC- or sh-SNHG7-transfected HeLa cells. (D–G) Relative expression of lncRNA SNHG7, miR-122-5p, FOXP1 mRNA and FOXP1 protein in lumps isolated from mice injected with sh-NC- or sh-SNHG7-transfected HeLa cells. sh-NC, negative control shRNA; sh-SNHG7, lncRNA SNHG7 shRNA. ***p < 0.001.
Discussion
LncRNA SNHG7, located at Chromosome 9q34.3, belongs to the lncRNA SNHGs family, all of which are tightly correlated to the development and progression of a variety of malignant tumors [13, 14]. Functional exploration about the effect of lncRNA SNHG7 on tumor cells is originally reported in lung cancer, and this research reveals that inhibiting the upregulated lncRNA SNHG7 impairs proliferative, migratory and invasive abilities of lung cancer cells, which may result from lncRNA SNHG7 knockdown-induced downregulation of Fas apoptotic inhibitory molecule 2 [15]. A recent study has reported that prostate cancer cells display a remarkably higher lncRNA SNHG7, and silencing lncRNA SNHG7 reduces N-cadherin and increases E-cadherin to suppress EMT by directly sponging miR-324-3p [16]. In hepatocellular carcinoma, lncRNA SNHG7 is screened as a candidate that is involved in the metastasis and recurrence of hepatocellular carcinoma cells by using bioinformatic analysis [17]. The in vitro experiments demonstrate that lncRNA SNHG7-mediated upregulation of ribosomal protein L4 contributes to the enhancive malignant phenotypes of hepatocellular carcinoma cells [18]. In addition, lncRNA SNHG7 functions as a oncogene in pathogenesis of breast cancer, melanoma and glioma [11, 19, 20]. Herein, our research showed the increased lncRNA SNHG7 existed in cervical cancer tissues and cell lines, which coincides with the previous reporter [10]. Phenomenally, we found lncRNA SNHG7 reinforced chemoresistance, proliferation, migration and invasion of cervical cancer. Given that the crucial role of lncRNA SNHG7 in promoting the malignant progression of cervical cancer, targeting lncRNA SNHG7 may serve as a promising therapeutic strategy for the treatment of cervical cancer. Meanwhile, the correlation of lncRNA SNHG7 with the clinical characteristics of cervical cancer should be identified by more investigations. Based on the critical function of lncRNA SNHG7 and the clinical correlation with cervical cancer, the diagnostic and prognostic significance of lncRNA SNHG7 will be identified and lncRNA SNHG7 may serve as a potential biomarker for cervical cancer.
Functioning as the sponge of miRNAs is a critical characteristic of lncRNAs, and miRNAs exert the regulatory effect downstream of lncRNAs [21]. We screened miR-122-5p as the direct target of lncRNA SNHG7. It can be concluded that miR-122-5p is a specific microRNA for liver diseases [22]. Recently, A group of evidence suggests that miR-122-5p is engaged in pathogenesis of malignant tumors. In hepatocellular carcinoma cells, miR-122-5p overexpression distinctly impedes the abnormal growth and metastasis of the tumor in vitro and in vivo [23]. Xu et al. has demonstrated the inhibitory effects of miR-122-5p on proliferative and invasive abilities of bile duct carcinoma cells [24]. In addition, miR-122-5p inhibits development of pancreatic ductal adenocarcinoma, breast cancer and gastric cancer [25–27]. Moreover, the reduced miR-122-5p in serum of patients with clear cell renal cell carcinoma predicts a poor prognosis [28]. We found cervical cancer exhibited the downregulated miR-122-5p, and miR-122-5p overexpression antagonized the effects of lncRNA SNHG7 on cervical cancer cells. Importantly, FOXP1 was screened to be a direct downstream molecule of miR-122-5p in our research. Meanwhile, in the present study, we identified that miR-122-5p served as a potential tumor suppression in cervical cancer. As the miRNA-based anti-tumor strategies have been developed for many years, the delivery and target system for miR-122-5p in the treatment of cervical cancer need to be explored in future. Due to that miRNAs have presented the diagnostic and prognostic values in cancers, the clinical expression and significance of miR-122-5p should be identified by further studies.
As a key transcription factor, FOXP1 participates in regulation of biological process of tumor cells [29]. In diffuse large B-cell lymphoma, FOXP1 downregulation promotes sensitivity of the tumor cells to doxorubicin [30]. Additionally, the overexpressed FOXP1 increases resistance of ovarian cancer cells to cisplatin and radio-resistance of hepatocellular carcinoma cells [31], [32]. Beyond that, FOXP1 accelerates growth of non-small cell lung cancer [33]. Li et al. has reported that miR-374b-5p-induced inhibition of malignant phenotypes of ovarian cancer cells is abolished by FOXP1 overexpression [34]. In cervical cancer, FOXP1 strengthens the growth and radio-resistance by counteracting the inhibitory role of miR-449b-5p [35]. It has been also shown that FOXP1 reduces the amount of tumor infiltrating lymphocytes to protect breast cancer [36]. Therefore, FOXP1 could be an oncogene by regulating biological function of tumor cells as well as lymphocytes. In our work, FOXP1 desensitized cervical cancer cells to cisplatin and exerted the pro-tumor effects on cervical cancer cells. Meanwhile, it has been reported that FOXP1 regulates skeletal aging by modulating mesenchymal stem cell senescence and commitment [37]. In this study, we identified that lncRNA SNHG7 induced FOXP1 expression by targeting miR-122-5p. The potential function of lncRNA SNHG7/miR-122-5p/FOXP1 axis in the regulation of aging and senescence deserves to be explored in future investigations.
Taken together, our research unveiled the engagement of lncRNA SNHG7/miR-122-5p/FOXP1 axis enabled cervical cancer cells to gain the potentiated chemoresistance, proliferation, migration and invasion in vitro and to grow in vivo, highlighting a critical role of lncRNA SNHG7 in the pathogenesis of cervical cancer.
Methods
Ethics statement
Investigation was conducted in accordance with the ethical standards and according to the Declaration of Helsinki. The experiments involving clinical tissue samples and mice were approved by Ethics Committee of Jinan Maternal and Child Health Hospital.
Clinical samples
The clinical tissue samples from patients with cervical cancer (n = 62) containing a group of patients with cisplatin-resistant cervical cancer (n = 29), who underwent surgical treatment, were obtained from 2016 to 2018 at Beijing Jishuitan Hospital. The characteristics of the participants were shown in Table 1. The investigators had informed the participants of the purposes and methods of usage of the tissues, and the participants agreed to signed the informed consent. All of the procedures were authorized by Jinan Maternal and Child Health Hospital.
Tumor-bearing mice
A total of twelve female nude mice were obtained from Beijing Jishuitan Hospital, and housed in cages at temperature between 22°C–26°C under the condition of 12 h light-dark cycles, and randomly grouped: control group and experimental group. Mice in control group were injected with HeLa cells with negative control short hairpin RNA (sh-NC) transfection, and mice in experimental group were injected with HeLa cells with short hairpin RNA against lncRNA SNHG7 (sh-SNHG7) transfection. To obtain a stable lncRNA SNHG7 knockdown cell line, we treated the transfected cells with Puromycin (2 mg/mL, Thermo Fisher Scientific, USA) before injecting the cells in the experimental mice. After 4 weeks, the mice were anesthetized with pentobarbital sodium (5 mg/100 g body weight) intraperitoneally, and killed by decapitation. The lumps were isolated carefully to be weighed. The procedures were in accordance with guidelines of Animal Use and Care Committee of Jinan Maternal and Child Health Hospital. The animal experiments were carried out in accordance with the requests of Animal Use and Care Committee of Jinan Maternal and Child Health Hospital.
Cell culture
DMEM (ThermoFisherScientific, USA) containing 10% FBS (Gibco, USA) was used to culture the human cervical epithelial immortalized cell line H8 and human cervical cancer cell line C-33A, C-4 II, Hela, SiHa, and HT-3 at 37°C. The cisplatin-resistant HeLa cell line (CR-HeLa) was established by using cisplatin (Selleck, USA), and cultured in the same condition.
Cell transfection
The lncRNA SNHG7 overexpression vector pEXP-RB-Mam (SNHG7 OE) and FOXP1 overexpression vector (FOXP1 OE) were purchased from RIBOBIO (China). Small interfering RNA against lncRNA SNHG7 (si-SNHG7), miR-122-5p mimics, miR-122-5p inhibitor and sh-SNHG7 were purchased from GenePharma (China). Lipofectamine 3000 (Invitrogen, USA) was adopted to conduct cell transfection. The relevant sequences were displayed in Table 2.
Table 2. Sequences of primers, siRNAs, mimics, and inhibitor used in this study.
| Gene | Sequence (5′–3′) |
| U6 | Forward: TGCGGGTGCTCGCTTCGGCAGC Reverse: CCAGTGCAGGGTCCGAGGT |
| GAPDH | Forward: AGACCACAGTCCATGCCATC Reverse: CAGGGCCCTTTTTCTGAGCC |
| lncRNA SNHG7 | Forward: CGATACCATTGAACACGCTGC Reverse: GGTTGAGGGTCCCAGTG |
| miR-122-5p | Forward: TGCGGTTGGAGTGTGACAATGG Reverse: TGGTGTCGTGGAGTCG |
| FOXP1 | Forward: CGGTACTCAGACAAATACAA Reverse: GRCGGAAGTAAGCAAACA |
| si-SNHG7 | GCUGGAAUAAAGAGUAACAUU |
| si-NC | AAUUCUCCGAACGUGUCACGU |
| miR-122-5p mimic | AACGCCAUUAUCACACUAAAUA |
| mimic NC | UUCUCCGAACGUGUCACUGUU |
| miR-122-5p inhibitor | CAAACACCAUUGUCACACUCCA |
| inhibitor NC | CAGUACUUUUGUGUAGUACAA |
Detection of the gene expression
TRIzol (Ambion, USA) was utilized to extract the total RNA. SuperScript III (Invitrogen, USA) was used to obtain cDNA. SYBR Premix Ex Taq II (Takara, Japan) was utilized to perform quantitative real-time PCR (qRT-PCR). U6 and GAPDH were utilized as the internal control. The relative expression of the indicated genes was calculated by using the comparative Ct (2−ΔΔCT) method. The primer sequences were shown in Table 2. All of the primers were synthesized and purchased from Sangon (China).
Detection of the protein expression
The cells were lysed with RIPA (Beyotime, China) followed by quantification by using BCA Protein Assay Kit (Thermo Fisher Scientific, USA). SDS-PAGE (Beyotime, China) was utilized to test the protein samples. Primary antibodies (Anti-FOXP1, 1:2000, Abcam, USA; Anti-GAPDH, 1:2000, Beyotime, China) were used at 4°C overnight. The protein bands were detected by using ChemiDoc XRS+ (Bio-Rad, USA).
Luciferase reporter assay
ENCORI database was used to obtain the binding sites between lncRNA SNHG7 and miR-122-5p, and miRBase and TargetScanHuman database were used to obtain the binding sites between FOXP1 and miR-122-5p. Wild type lncRNA SNHG 7, mutant lncRNA SNHG 7, wild type FOXP1 and mutant FOXP1 were cloned into pmirGLO vector followed by transfection into HeLa cells in the presence of miR-122-5p mimics or negative control mimics (mimics NC). Bio-Glo™ Luciferase Assay System (Promega, USA) was utilized to evaluate the luciferase activity.
Measurement of cell viability
Cell Counting Kit-8 (CCK-8, Beyotime, China) was adopted to detect cell viability. After transfection, 6000 cells were seeded into each well in a 96-well plate for indicated time, and cultured in 110 ml culture medium containing 10 ml CCK-8 solution for 2 h at 37°C. Optical density (OD) 450 was the indicator of cell viability. Additionally, different concentration of cisplatin was used to treat the HeLa cells with indicated transfection to calculate the half maximal inhibitory concentration (IC50) when the cell viability reached 50%.
Colony formation assay
Proliferative capacity was evaluated by using colony formation assay. After transfection, HeLa cells were seeded into the 6-well plate at a concentration of 5000 cells per well. Two weeks later, formaldehyde was utilized to fix the cells followed by staining cells with crystal violet.
Wound healing assay
Migratory ability was indicated by performing wound healing assay. After transfection, HeLa cells were seeded into a 6-well plate for 24 h. The cells were scratched by A 10 ml tip was used to scratch the cells, and serum-free medium was utilized to culture the cells. At the time point of 0 h and 24 h, the scratch was captured. The migratory distance was measured to calculate the wound healing rate.
Measurement of invasive capacity
Invasive capacity was evaluated with transwell assay. HeLa cells with indicated treatment were seeded in Millicell Standing Cell Culture 24 well containing Matrigel (Millipore, USA) for 24 h in according to the manufacturer’s guidelines. Formaldehyde was utilized to fix the cells followed by staining cells with crystal violet.
Statistical analysis
The experiments were performed independently three times at least. Mean ± standard deviation (SD) depicted the data in our research, and the data were analyzed by GraphPad Prism 8 Software (GraphPad, USA). The statistical analysis method was t-test and Chi-Squared test, and p < 0.05 indicated statistical difference.
Supplementary Materials
Author Contributions
Yanmei Hou, Jie Li designed the study; Min Song, Junli Lu performed experiments; Hui Liu gathered and analysed data; Xiaohong Yu and Meng Xu wrote the manuscript.
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
References
- 1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492 [PubMed]
- 2. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–32. https://doi.org/10.3322/caac.21338 [PubMed]
- 3. Aalijahan H, Ghorbian S. Long non-coding RNAs and cervical cancer. Exp Mol Pathol. 2019; 106:7–16. https://doi.org/10.1016/j.yexmp.2018.11.010 [PubMed]
- 4. Tornesello ML, Faraonio R, Buonaguro L, Annunziata C, Starita N, Cerasuolo A, Pezzuto F, Tornesello AL, Buonaguro FM. The Role of microRNAs, Long Non-coding RNAs, and Circular RNAs in Cervical Cancer. Front Oncol. 2020; 10:150. https://doi.org/10.3389/fonc.2020.00150 [PubMed]
- 5. Guo HM, Yang SH, Zhao SZ, Li L, Yan MT, Fan MC. LncRNA NEAT1 regulates cervical carcinoma proliferation and invasion by targeting AKT/PI3K. Eur Rev Med Pharmacol Sci. 2018; 22:4090–97. https://doi.org/10.26355/eurrev_201807_15400 [PubMed]
- 6. Chang X, Zhang H, Yang Q, Pang L. LncRNA SOX2OT affects cervical cancer cell growth, migration and invasion by regulating SOX2. Cell Cycle. 2020; 19:1391–403. https://doi.org/10.1080/15384101.2020.1750812 [PubMed]
- 7. Wang X, Zhang J, Wang Y. Long noncoding RNA GAS5-AS1 suppresses growth and metastasis of cervical cancer by increasing GAS5 stability. Am J Transl Res. 2019; 11:4909–21. [PubMed]
- 8. Zhao S, Yu M, Wang L. LncRNA miR503HG Regulates the Drug Resistance of Recurrent Cervical Squamous Cell Carcinoma Cells by Regulating miR-155/Caspase-3. Cancer Manag Res. 2020; 12:1579–85. https://doi.org/10.2147/CMAR.S225489 [PubMed]
- 9. Zhou Y, Tian B, Tang J, Wu J, Wang H, Wu Z, Li X, Yang D, Zhang B, Xiao Y, Wang Y, Ma J, Wang W, Su M. SNHG7: A novel vital oncogenic lncRNA in human cancers. Biomed Pharmacother. 2020; 124:109921. https://doi.org/10.1016/j.biopha.2020.109921 [PubMed]
- 10. Zeng J, Ma YX, Liu ZH, Zeng YL. LncRNA SNHG7 contributes to cell proliferation, invasion and prognosis of cervical cancer. Eur Rev Med Pharmacol Sci. 2019; 23:9277–85. https://doi.org/10.26355/eurrev_201911_19420 [PubMed]
- 11. Sun X, Huang T, Liu Z, Sun M, Luo S. LncRNA SNHG7 contributes to tumorigenesis and progression in breast cancer by interacting with miR-34a through EMT initiation and the Notch-1 pathway. Eur J Pharmacol. 2019; 856:172407. https://doi.org/10.1016/j.ejphar.2019.172407 [PubMed]
- 12. Dai Y, Zhang X, Xing H, Zhang Y, Cao H, Sang J, Gao L, Wang L. Downregulated long non-coding RNA SNHG7 restricts proliferation and boosts apoptosis of nasopharyngeal carcinoma cells by elevating microRNA-140-5p to suppress GLI3 expression. Cell Cycle. 2020; 19:448–63. https://doi.org/10.1080/15384101.2020.1712033 [PubMed]
- 13. Yang H, Jiang Z, Wang S, Zhao Y, Song X, Xiao Y, Yang S. Long non-coding small nucleolar RNA host genes in digestive cancers. Cancer Med. 2019; 8:7693–704. https://doi.org/10.1002/cam4.2622 [PubMed]
- 14. Zimta AA, Tigu AB, Braicu C, Stefan C, Ionescu C, Berindan-Neagoe I. An Emerging Class of Long Non-coding RNA With Oncogenic Role Arises From the snoRNA Host Genes. Front Oncol. 2020; 10:389. https://doi.org/10.3389/fonc.2020.00389 [PubMed]
- 15. She K, Huang J, Zhou H, Huang T, Chen G, He J. lncRNA-SNHG7 promotes the proliferation, migration and invasion and inhibits apoptosis of lung cancer cells by enhancing the FAIM2 expression. Oncol Rep. 2016; 36:2673–80. https://doi.org/10.3892/or.2016.5105 [PubMed]
- 16. Han Y, Hu H, Zhou J. Knockdown of LncRNA SNHG7 inhibited epithelial-mesenchymal transition in prostate cancer though miR-324-3p/WNT2B axis in vitro. Pathol Res Pract. 2019; 215:152537. https://doi.org/10.1016/j.prp.2019.152537 [PubMed]
- 17. Cui H, Zhang Y, Zhang Q, Chen W, Zhao H, Liang J. A comprehensive genome-wide analysis of long noncoding RNA expression profile in hepatocellular carcinoma. Cancer Med. 2017; 6:2932–41. https://doi.org/10.1002/cam4.1180 [PubMed]
- 18. Yang X, Sun L, Wang L, Yao B, Mo H, Yang W. LncRNA SNHG7 accelerates the proliferation, migration and invasion of hepatocellular carcinoma cells via regulating miR-122-5p and RPL4. Biomed Pharmacother. 2019; 118:109386. https://doi.org/10.1016/j.biopha.2019.109386 [PubMed]
- 19. Li H, Li T, Huang D, Zhang P. Long noncoding RNA SNHG17 induced by YY1 facilitates the glioma progression through targeting miR-506-3p/CTNNB1 axis to activate Wnt/β-catenin signaling pathway. Cancer Cell Int. 2020; 20:29. https://doi.org/10.1186/s12935-019-1088-3 [PubMed]
- 20. Zhang C, Zhu B, Li XB, Cao YQ, Yang JC, Li X, Liu YX, Wang YB. Long non-coding RNA SNHG7 promotes migration and invasion of melanoma via upregulating SOX4. Eur Rev Med Pharmacol Sci. 2019; 23:4828–34. https://doi.org/10.26355/eurrev_201906_18069 [PubMed]. Retraction in: Eur Rev Med Pharmacol Sci. 2020; 24:7553. https://doi.org/10.26355/eurrev_202007_22199 [PubMed]
- 21. Adams BD, Parsons C, Walker L, Zhang WC, Slack FJ. Targeting noncoding RNAs in disease. J Clin Invest. 2017; 127:761–71. https://doi.org/10.1172/JCI84424 [PubMed]
- 22. Matsuzaki J, Ochiya T. Extracellular microRNAs and oxidative stress in liver injury: a systematic mini review. J Clin Biochem Nutr. 2018; 63:6–11. https://doi.org/10.3164/jcbn.17-123 [PubMed]
- 23. Ma J, Li T, Han X, Yuan H. Knockdown of LncRNA ANRIL suppresses cell proliferation, metastasis, and invasion via regulating miR-122-5p expression in hepatocellular carcinoma. J Cancer Res Clin Oncol. 2018; 144:205–14. https://doi.org/10.1007/s00432-017-2543-y [PubMed]
- 24. Xu Z, Liu G, Zhang M, Zhang Z, Jia Y, Peng L, Zhu Y, Hu J, Huang R, Sun X. miR-122-5p Inhibits the Proliferation, Invasion and Growth of Bile Duct Carcinoma Cells by Targeting ALDOA. Cell Physiol Biochem. 2018; 48:2596–606. https://doi.org/10.1159/000492702 [PubMed]
- 25. Meng L, Chen Z, Jiang Z, Huang T, Hu J, Luo P, Zhang H, Huang M, Huang L, Chen Y, Lu M, Xu AM, Ying S. MiR-122-5p suppresses the proliferation, migration, and invasion of gastric cancer cells by targeting LYN. Acta Biochim Biophys Sin (Shanghai). 2020; 52:49–57. https://doi.org/10.1093/abbs/gmz141 [PubMed]
- 26. Dai C, Zhang Y, Xu Z, Jin M. MicroRNA-122-5p inhibits cell proliferation, migration and invasion by targeting CCNG1 in pancreatic ductal adenocarcinoma. Cancer Cell Int. 2020; 20:98. https://doi.org/10.1186/s12935-020-01185-z [PubMed]
- 27. Ergün S, Ulasli M, Igci YZ, Igci M, Kırkbes S, Borazan E, Balik A, Yumrutaş Ö, Camci C, Cakmak EA, Arslan A, Oztuzcu S. The association of the expression of miR-122-5p and its target ADAM10 with human breast cancer. Mol Biol Rep. 2015; 42:497–505. https://doi.org/10.1007/s11033-014-3793-2 [PubMed]
- 28. Heinemann FG, Tolkach Y, Deng M, Schmidt D, Perner S, Kristiansen G, Müller SC, Ellinger J. Serum miR-122-5p and miR-206 expression: non-invasive prognostic biomarkers for renal cell carcinoma. Clin Epigenetics. 2018; 10:11. https://doi.org/10.1186/s13148-018-0444-9 [PubMed]
- 29. Kim JH, Hwang J, Jung JH, Lee HJ, Lee DY, Kim SH. Molecular networks of FOXP family: dual biologic functions, interplay with other molecules and clinical implications in cancer progression. Mol Cancer. 2019; 18:180. https://doi.org/10.1186/s12943-019-1110-3 [PubMed]
- 30. Marques SC, Ranjbar B, Laursen MB, Falgreen S, Bilgrau AE, Bødker JS, Jørgensen LK, Primo MN, Schmitz A, Ettrup MS, Johnsen HE, Bøgsted M, Mikkelsen JG, Dybkær K. High miR-34a expression improves response to doxorubicin in diffuse large B-cell lymphoma. Exp Hematol. 2016; 44:238–46.e2. https://doi.org/10.1016/j.exphem.2015.12.007 [PubMed]
- 31. Hu Z, Cai M, Zhang Y, Tao L, Guo R. miR-29c-3p inhibits autophagy and cisplatin resistance in ovarian cancer by regulating FOXP1/ATG14 pathway. Cell Cycle. 2020; 19:193–206. https://doi.org/10.1080/15384101.2019.1704537 [PubMed]
- 32. Zhang Y, Huang B, Chen Z, Yang S. Knockdown of LINC00473 Enhances Radiosensitivity in Hepatocellular Carcinoma via Regulating the miR-345-5p/FOXP1 Axis. Onco Targets Ther. 2020; 13:173–83. https://doi.org/10.2147/OTT.S240113 [PubMed]
- 33. Zhou W, Li R. microRNA-605 inhibits the oncogenicity of non-small-cell lung cancer by directly targeting Forkhead Box P1. Onco Targets Ther. 2019; 12:3765–77. https://doi.org/10.2147/OTT.S193675 [PubMed]
- 34. Li H, Liang J, Qin F, Zhai Y. MiR-374b-5p-FOXP1 feedback loop regulates cell migration, epithelial-mesenchymal transition and chemosensitivity in ovarian cancer. Biochem Biophys Res Commun. 2018; 505:554–60. https://doi.org/10.1016/j.bbrc.2018.09.161 [PubMed]
- 35. Cheng L, Shi X, Huo D, Zhao Y, Zhang H. MiR-449b-5p regulates cell proliferation, migration and radioresistance in cervical cancer by interacting with the transcription suppressor FOXP1. Eur J Pharmacol. 2019; 856:172399. https://doi.org/10.1016/j.ejphar.2019.05.028 [PubMed]
- 36. De Silva P, Garaud S, Solinas C, de Wind A, Van den Eyden G, Jose V, Gu-Trantien C, Migliori E, Boisson A, Naveaux C, Duvillier H, Craciun L, Larsimont D, et al. FOXP1 negatively regulates tumor infiltrating lymphocyte migration in human breast cancer. EBioMedicine. 2019; 39:226–38. https://doi.org/10.1016/j.ebiom.2018.11.066 [PubMed]
- 37. Li H, Liu P, Xu S, Li Y, Dekker JD, Li B, Fan Y, Zhang Z, Hong Y, Yang G, Tang T, Ren Y, Tucker HO, et al. FOXP1 controls mesenchymal stem cell commitment and senescence during skeletal aging. J Clin Invest. 2017; 127:1241–53. https://doi.org/10.1172/jci89511 [PubMed]


