Introduction
Hepatocellular carcinoma (HCC) causes nearly half a million deaths annually worldwide. Owing to the increasing incidence of hepatitis B virus (HBV) infection, HCC has become a fast-growing cancer in Asian countries, especially in China [1]. The incidence of HCC is expected to increase in the future. However, few therapies can improve the prognosis of patients with HCC [2]. Surgery is the primary curative treatment for HCC. However, no more than 50% of patients survive longer than five years after surgery, even when diagnosed and operated at an early stage [3]. Recently, molecular targeted therapy has brought a glimmer of hope for the treatment of HCC [4]. Owing to its favorable overall survival, the FDA approved the multi-kinase inhibitor sorafenib for the treatment of advanced HCC [5, 6]. As a targeted anticancer molecule, sorafenib has demonstrated only partial efficacy in advanced HCC. Therefore, it is mandatory to better understand the genes and signaling pathways involved in the tumorigenesis and progression of HCC and to identify more effective druggable targets for improving HCC management.
Over the past few years, Wnt pathway activation in HCC has been reported in several studies [7, 8]. This pathway has been indicated to play an important role in the clinicopathological features and prognosis of HCC [9]. According to a recent report, CTNNB1 is one of the most frequently mutated genes in HCC [10]. Mutation of CTNNB1, which is the key downstream effector of the pathway, appears to be the main cause of activation of the Wnt pathway [11]. β-catenin, a protein expressed by CTNNB1, integrates the intercellular E-cadherin–catenin adhesion system, the disruption of which has been observed in HCC [12]. β-catenin is normally located in the cytomembrane and is directly connected to E-cadherin, which in turn forms an adhesion complex. This adhesion complex, which can be degraded by phosphorylation or ubiquitination, can regulate cell-cell adhesion and maintain tissue architecture and function. In HCC, unbound β-catenin translocates to the nucleus and regulates the transcription of target genes relevant to cell proliferation and cell cycle progression. β-catenin accumulation in the cytoplasm and/or nucleus is thought to be closely associated with poor prognosis and deep invasion in HCC patients, independent of tumor stage [13]. Recently, mutations of CTNNB1 have been detected in human HCC, but the clinical implications of the CTNNB1 mutation are still unclear. Our meta-analysis showed that mutant CTNNB1 was associated with favorable clinical outcomes and survival in patients with HCC.
Materials and Methods
Study selection
PubMed, Web of Science, Elsevier, Cochrane Library, Ovid, and Embase databases were searched for articles published until January 20, 2023. The terms used in the search were “CTNNB1 or beta-catenin, or β-catenin” and “prognostic or prognosis or survival” and “hepatocellular carcinoma or HCC or liver cancer or liver tumor or hepatic cancer or liver tumor or liver neoplasms”. The reference lists of all retrieved articles were manually searched. Only studies published in English were included. Two reviewers (GLL and GQS) completed the systematic literature search and extracted the following parameters from each study: study population characteristics, number of participants, sex ratio, first author, and year of publication.
Criteria for inclusion and exclusion
Inclusion criterion:
Patients with HCC were diagnosed by pathology;
Information about CTNNB1 mutation, OS (Overall Survival), and other clinicopathological features were provided;
The CTNNB1 mutation was sequenced for exon 3, SSCP analysis of exon 3, Sanger sequencing, mass array, PCR, or other methods in primary HCC tissue
The study with the highest quality assessment was enrolled when more than one study was reported by one individual author;
Studies were published in English.
Exclusion criterion:
Articles not related to the clinic;
Overlapping publications;
Information about CTNNB1 mutation or OS or other clinicopathological features that were not clearly reported;
Abstracts, reviews, letters, editorials, and expert opinions;
Non-English publications.
Data extraction and literature quality assessment
Two reviewers (GLL and GQS) independently evaluated each study, and relevant characteristics were listed: (1) the first author and publication year; (2) population origin; (3) number of cases; (4) mean age, (5) gender, (6) the number of cases with CTNNB1 mutation; (7) level of evidence, (8) disease stage, (9) clinicopathological features, (10) methods of evaluating CTNNB1 mutation, and (11) OS data.
The quality of each study was assessed using the Newcastle-Ottawa scale (NOS), which evaluates various aspects of the methodology, including selection, comparability, and outcome [14]. The final scores ranged from 0 (lowest) to 9 (highest); the higher the value, the better the eligibility.
Statistical analysis
Review Manager (RevMan) software (version 5.2; Cochrane Collaboration) was used for the meta-analysis. Odds ratios (OR) combined with 95%confidence intervals (CI) were analyzed to evaluate the association of CTNNB1 mutation with the prognosis and clinicopathological factors of HCCs. Pooled ORs and 95%CIs were used as the recommended summary statistics. A fixed- or random-effects model was used to calculate pooled effects. Funnel plots, which were used to examine the risk of potential publication bias, were constructed using Egger’s test and Begg’s test. Heterogeneity was evaluated by I2, and I2 statistics of ≥50%, defined as heterogeneity. Statistical significance was set at P <0.05.
Availability of data and materials
All data and materials were availability from the web.
Consent to publication
All co-authors consented to publish this paper.
Results
Selection of trials
The original search strategy retrieved 223 publications. After screening, 187 studies were excluded, and 36 papers were captured. Of these, 19 were excluded because of a lack of adequate data on CTNNB1 mutations and specific parameters. Thus, 17 studies with sufficient evaluation met the inclusion criteria and were retrieved for further evaluation. A flowchart of the strategies is presented in Figure 1.
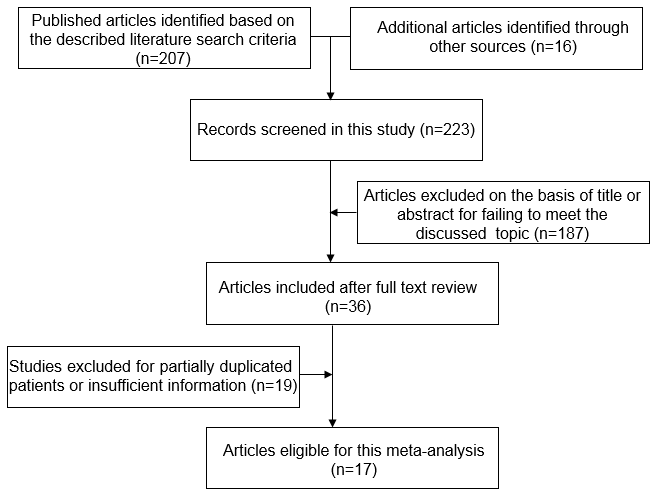
Figure 1. Flow chart of literature search strategies.
Study characteristics
The patient characteristics in each selected study are shown in Table 1. The total number of patients was 1828, with 319 CTNNB1 mutations. The mean incidence of CTNNB1 mutation was 17.5%. The information extracted from the selected studies included the gene type of CTNNB1, prognosis, disease stage, methods of evaluating CTNNB1 mutation, OS data, and clinicopathological features. The ORs and 95% CI between CTNNB1 mutation and OS are provided. All studies retrieved in this meta-analysis were performed properly, and the gene type of CTNNB1 was determined by sequencing of exon 3, SSCP analysis of exon 3, Sanger sequencing, mass array, PCR, or other methods without subjective interference. The primary mutation was found in exon 3.
Table 1. Characteristics of studies included in the meta-analysis.
| First author and year | Country /region | No. of patients | Mean age | Gender (M/F) | Mu/total | Level of evidence | Stage | Clinicopathological features | Method | ProvidedOS data | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ding [15] 2014 | China | 156 | 53.09±11.19 | NR | 15/156 | 5 | I–IV | D,T | Mass array | Yes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lin [16] 2010 | Taiwan | 160 | 57(14-88) | 122/38 | 22/128 | 5 | NR | NR | Direct sequencing of exon 3 | Yes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lu [17] 2014 | Taiwan | 115 | 56.3(23.6–83.1) | 97/18 | 21/115 | 5 | I–IV | T,M | Direct sequencing of exon 3 | Yes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mao [18] 2001 | Taiwan | 372 | NR | 293/162 | 36/372 | 5 | I–III | D,T | PCR | Yes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wong [19] 2001 | Hong Kong | 60 | 54(28-74) | 46/14 | 6/60 | 4 | I–IV | D | PCR | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Yuan [20] 2013 | Taiwan | 305 | 55.09 (15–88) | 239/66 | 32/214 | 4 | I–IV | T | direct sequencing of exon 3 | Yes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cavard [21] 2006 | France | 42 | NR | NR | 21/42 | 3 | NR | NR | sequencing | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cieply [22] 2009 | USA | 25 | NR | 19/6 | 9/25 | 3 | I–IV | T,M | Direct sequencing of exon 3 | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hsu [23] 2000 | Japan | 125 | 63(16-79) | 88/37 | 57/434 | 3 | I–IV | D,T | PCR | Yes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Huang [24] 1999 | Japan/ Switzerland | 16+6 | NR | NR | 9/22 | 3 | NR | D | DNA sequence | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kim [25] 2008 | Korea | 36 | 57.7(34-71) | 32/4 | 1/36 | 3 | I–IV | D,T,M | sequencing | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Puig [26] 2001 | France | 137 | NR | 110/27 | 32/137 | 3 | NR | D,M | Direct sequencing | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Li [27] 2011 | United States | 7 | 56.86 | ¾ | 5/7 | 3 | I–III | T | Sanger sequencing | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| China | 1 | 68 | 1/0 | 1/1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The Netherlands | 1 | 53 | 1/0 | 0/1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Park [28] 2005 | Korea | 92 | 51.6(26-89) | 75/17 | 13/32 | 3 | I–IV | D,T | SSCP analysis of exon 3 | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taniguchi [29] 2002 | USA | 73 | NR | 41/32 | 14/73 | 3 | NR | D | PCR | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tornesello [30] 2013 | Italian | 67 | NR | 53/14 | 10/67 | 3 | NR | D | DNA sequence electropherograms | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Rossi [31] 2007 | France | 32 | NR | NR | 15/32 | 3 | NR | NR | sequencing | NR | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| D, histologic differentiation degree; T, depth of tumor invasion; M, metastasis (include those with more than one Nodules); OS, overall survival; NR, not reported; M/F, male/female; Mu, CTNNB1 mutation. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Quality assessment
The methodological quality of the 17 studies was assessed using NOS. On the basis of the NOS, 4 studies scored 5 points [15–18], 2 studies scored 4 [19, 20], 11 studies scored 3 [21–31] in total of the 17 studies. Studies with a score ≥ 5 were defined as high-quality (Table 1).
Impact of CTNNB1 mutation on overall survival (OS)
Some of the included studies [15–18, 20, 23] provided the ORs and 95%CI directly or indirectly when discussing the correlation between CTNNB1 mutation and OS. This meta-analysis systematically assessed the association of CTNNB1 mutation with OS in patients with HCC at 1-year, 3-year and 5-year, respectively. It was demonstrated that CTNNB1 mutation significantly correlated with poor 1-, 3-and 5-year OS, as shown in Figure 2. And the pooled ORs were 0.52(n = 6 studies, 95% CI: 0.34–0.81, Z = 2.89, P =0.004, 0.28 (n =6 studies, 95% CI: 0.18–0.42, Z = 6.03, P<0.00001), -0.22(n = 6 studies, 95% CI: 0.37–0.06, Z = 2.78, P = 0.005) respectively. The statistical heterogeneity of the 1-, 3-and 5-year OS was 18%, 0%, and 81%, respectively. The above results suggest that CTNNB1 mutation is correlated with a favorable prognosis for HCC.
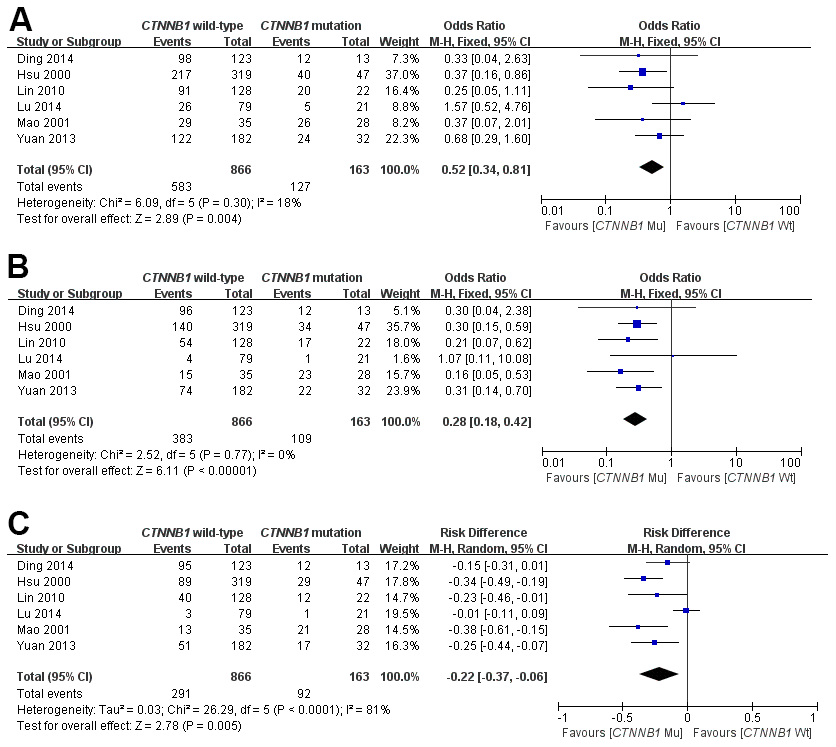
Figure 2. Forest plot of odds ratio for the association of CTNNB1 mutation with 1-year (A), 3-year (B) and 5-year (C) overall survival.
Correlation of CTNNB1 mutation with clinicopathological parameters
This meta-analysis assessed the relationship between CTNNB1 mutations and clinicopathological parameters, including metastasis, vascular invasion, tumor size, differentiation, TNM stages, liver cirrhosis, and HBV/HCV infection (Table 2). Ten studies [15, 18, 19, 23–26, 28–30] evaluated the correlation between CTNNB1 mutations and differentiation (Figure 3). The pooled OR was 0.54 (95%CI: 0.36-0.81, Z=2.98, P=0.003). This result showed that there was a significant correlation between CTNNB1 mutation and differentiation. Nine studies [15, 17, 18, 20, 22, 23, 25, 27, 28] evaluated the correlation of CTNNB1 mutations with TNM stages (T3/T4 versus T1/T2) (Figure 4). The pooled OR was -0.25 (95%CI: -0.33--0.18, Z=6.6, P<0.00001). This result indicated that there was a significant correlation between CTNNB1 mutations and the TNM stages of HCC. Additionally, we also assessed the relationship between CTNNB1 mutation and liver cirrhosis of HCC.4 studies [19, 21, 22, 26] evaluated the correlation of CTNNB1 mutation with liver cirrhosis (Figure 5). The pooled OR was 0.21 (95%CI: 0.11-0.39, Z=4.94, P<0.00001). This result showed a significant correlation between CTNNB1 mutations and liver cirrhosis. We found that CTNNB1 mutation had a better effect on the above two clinicopathological features. For other parameters, such as metastasis (Supplementary Figure 1), vascular invasion (Supplementary Figure 2), and tumor size (Supplementary Figure 3), of HCC showing CTNNB1 mutation, the pooled ORs were1.25(n=4studies, 95%CI:0.93-1.66, Z=1.49, P=0.14),1.42 (n=3 studies, 95%CI:0.82-2.45, Z=1.26, P=0.21) and 1.24 (n=5 studies, 95%CI:0.37-4.11, Z=0.35, P=0.72), respectively, demonstrating that CTNNB1 mutation had no significant correlation with these parameters.
Table 2. Meta-analysis comparing HCC with CTNNB1 mutation and wild-type.
| Outcome of interest | No. of studies | Number of tissue samples | OR/WMD | 95% CI | P value | I2(%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Overall Survival [15–18, 20, 23] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 year | 6 | CTNNB1 Mu=163, CTNNB1 Wt =866 | 0.52 | 0.34-0.81 | 0.004 | 18 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 year | 6 | CTNNB1 Mu=163, CTNNB1 Wt =866 | 0.28 | 0.18-0.42 | <0.00001 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 5 year | 6 | CTNNB1 Mu=163, CTNNB1 Wt =866 | -0.22 | -0.37--0.06 | 0.005 | 81 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Differentiation grade [15, 18, 19, 23–26, 28–30,] | 10 | CTNNB1 Mu=184, CTNNB1 Wt =680 | 0.54 | 0.36-0.81 | 0.003 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TMN stage [15, 17, 18, 20, 22, 23, 25, 27, 28] | 9 | CTNNB1 Mu=174, CTNNB1 Wt =841 | -0.25 | -0.33--0.18 | <0.00001 | 39 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metastasis [17, 22, 25, 26] | 4 | CTNNB1 Mu=69, CTNNB1 Wt =242 | 1.25 | 0.93-1.66 | 0.14 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Vascular invasion [17, 22, 26] | 3 | CTNNB1 Mu=70, CTNNB1 Wt =223 | 1.42 | 0.82-2.45 | 0.21 | 43 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Liver cirrhosis [19, 21, 22, 26] | 4 | CTNNB1 Mu=76, CTNNB1 Wt =138 | 0.21 | 0.11-0.39 | <0.00001 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tumor size [19, 22, 25, 26, 28] | 5 | CTNNB1 Mu=67, CTNNB1 Wt =171 | 1.24 | 0.37-4.11 | 0.72 | 54 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HBV [15, 17–19, 21, 23, 25–28, 30, 31] | 12 | CTNNB1 Mu=213, CTNNB1 Wt =795 | 0.44 | 0.31-0.64 | <0.0001 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HCV [15, 17, 21, 22, 25, 27, 28, 30, 31] | 9 | CTNNB1 Mu=111, CTNNB1 Wt =356 | 1.70 | 0.93-3.11 | 0.09 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OR, odds ratio; WMD, weighted mean difference; CI, confidence interval; Mu, mutation; Wt, wild-type. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
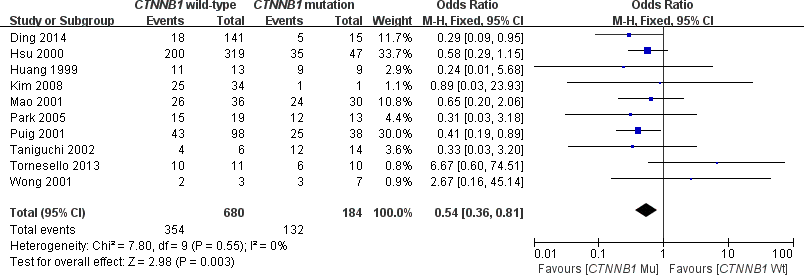
Figure 3. Forest plot of odds ratio for the association of CTNNB1 mutation with differentiation grade.
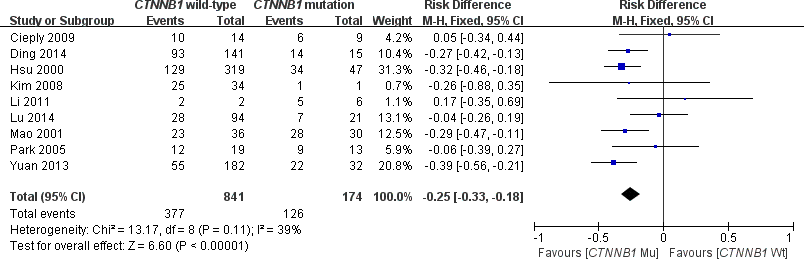
Figure 4. Forest plot of odds ratio for the association of CTNNB1 mutation with TNM stages.
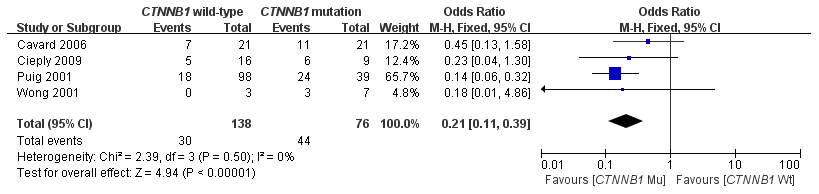
Figure 5. Forest plot of odds ratio for the association of CTNNB1 mutation with liver cirrhosis.
Correlation of CTNNB1 mutation with etiology
In this meta-analysis, the correlation between CTNNB1 mutations and etiology (HBV/HCV infection) was evaluated. As shown in Figure 6, 12 studies [15, 17, 18, 19, 21, 23, 25–28, 30, 31] assessed the relationship between CTNNB1 mutations and HBV. The combined ORs were 0.44(95%CI:0.31-0.64, Z=4.37, P<0.0001), with no significant statistical heterogeneity (I2=0%). This result indicates that a better effect was observed between CTNNB1 mutation and HBV infection. However, as shown in Supplementary Figure 4, 9 studies [15, 17, 21, 22, 25, 27, 28, 30, 31] assessed the relationship between CTNNB1 mutations and HCV. The combined ORs were 1.70(95%CI: 0.93-3.11, Z=1.72, P=0.09).
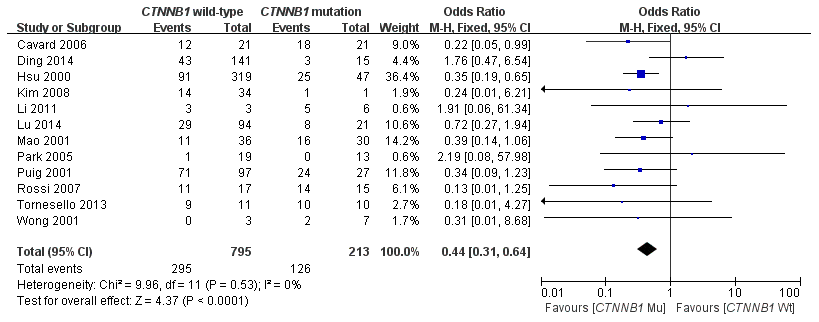
Figure 6. Forest plot of odds ratio for the association of CTNNB1 mutation with etiology (HBV).
Publication bias
For the studies included in this meta-analysis, Begg’s test indicated that there was no significant publication bias after assessing the funnel plot (Supplementary Figures 5–15).
Discussion
Human HCCs with activation of the Wnt/β-catenin pathway demonstrate unique gene expression patterns and pathological features. Activated Wnt/β-catenin synergizes with multiple signaling cascades to drive HCC formation, and it functions through its downstream effectors [32]. The aberrant Wnt pathway in HCC, has been well-studied and proven to be involved in the prognosis of HCC [33]. β-catenin is a key downstream effector of the Wnt signaling pathway and plays a crucial role. β-catenin has been the focus of attention as an attractive therapeutic target [9]. According to our previous observations, cytoplasmic and/or nuclear expression of β-catenin could serve as a potential predictor for the progression and prognosis of patients with HCC and act as a novel target for the developed therapies [13]. CTNNB1, the coding gene of β-catenin, is one of the most frequently mutated genes in HCC [10, 34]. Hsu et al. first reported a relationship between β-catenin mutations and prognosis in patients with HCC [23]. Based on the resected HCC of patients, their report showed that CTNNB1 mutations were associated with tumor differentiation, HBV infection, and clinical prognosis. However, according to a study by Lu et al., there is no prognostic significance of CTNNB1 mutation in patients with HCC [17]. Moreover, no difference in CTNNB1 mutation rate was observed between patients with HCC with or without HBV infection. Tumor mutational burden (TMB) was verified to be closely associated with immune checkpoint inhibitors, but it is unclear whether gene mutation has an effect on immunotherapy of HCC. Mo et al. firstly revealed the underlying association between CTNNB1 mutation and immunotherapy, and they speculated that CTNNB1 mutation may modulate NK cells by affecting CD96 [35]. To date, however, tremendous work has been done to investigate the association between CTNNB1 mutations and clinicopathological characteristics and prognosis of patients with HCC; however, no conclusive results have been achieved. Wang et al. reported that the CTNNB1 mutation was correlated with a favorable prognosis in HCC in a meta-analysis [36]. Of note, the studies enrolled in their meta-analysis failed to provide adequate information about the relationship between CTNNB1 mutations and clinical prognosis. The current study focused on CTNNB1 mutations in HCC in a clinical setting. We showed that CTNNB1 mutation has a robust effect on the clinical and prognosis of patients with HCC.
A number of studies have implicated an alteration in CTNNB1, including gene mutation and protein overexpression, in HCC [34, 37]. CTNNB1 plays a crucial role in hepatocyte adhesion and Wnt signaling pathway [38]. CTNNB1 is mainly located in the cell membrane. By binding to the lymphoid enhancer factor (LEF)/T-cell factor (TCF) family of DNA binding proteins, CTNNB1 enters the nucleus and regulates transcription of target genes such as c-mycor cyclin D1, resulting in proliferation and metastasis of liver tumor cells [39]. Xiao X et al. demonstrated that CTNNB1 mutation in HCC led to a decrease in chemokine expression and subsequent suppression of immune cell infiltration [40]. Recently, somatic mutations in CTNNB1 have been demonstrated not only in animal models but also in human HCC [10]. Comprehensive analysis of clinical samples has identified immunological and molecular classification of HCC, and the CTNNB1-mutated subtype exhibits distinctive characteristics of immunosuppressive tumor microenvironment [41]. However, the clinical implications of CTNNB1 mutation in human HCC are unclear [12, 42]. Paradoxical data exist concerning the prognostic value and clinicopathological significance of cytoplasmic and/or nuclear CTNNB1 accumulation. These discrepancies are most likely due to the gene type of CTNNB1. Cytoplasm and/or nucleusCTNNB1 accumulation can be associated with mutations in the CTNNB1 gene and other components of the signaling pathway [43]. The most common mechanism of CTNNB1 accumulation in HCC is mutations in CTNNB1 [44]. Mutations or wild-type CTNNB1 may influence the subcellular localization of β-catenin. Most studies on CTNNB1 mutations have focused on the consensus sequence for GSK-3β phosphorylation in exon 3 and the inactivation of APC and other factors [45]. Cui et al. found that mutation of exon3 of CTNNB1 is one of the most important factors activating the abnormal Wnt signaling pathway in HCC [46]. CTNNB1 mutations and nuclear overexpression may play a key role in HCC in Chinese people. And targeting the Wnt-beta-catenin pathway may represent a valid treatment option for Chinese HCC patients [15, 47, 48].
In the past 5 years, immune-checkpoint inhibitors have revolutionized the management of HCC [49]. However, several studies demonstrated that mutation of CTNNB1 have also been associated with scarcity of immune cells in the tumor microenvironment and poor clinical response to immune checkpoint inhibitor therapy [50]. Ogawa K et al. verified that Gain-of-function Inclusion mutation of CTNNB1 contributes to resistance of ICI monotherapy through the framework of non-T-cell-inflamed tumor microenvironment. Of note, the treatment effect of Atezolizumab plus bevacizumab in patients with HCC with MT CTNNB1 was comparable to those patients with WT CTNNB1. These results further implicate that bevacizumab added to Atezolizumab might improve immunosuppressive tumor microenvironment caused by CTNNB1 mutation [51]. In addition, Chen et al. proved that CTNNB1 alternation is a potential biomarker for immunotherapy prognosis in patients with HCC [52]. However, this meta-analysis concluded that HCC patients with CTNNB1 mutations appeared to have a favorable survival in comparison with wild-type CTNNB1 HCC. Additionally, CTNNB1 mutations were significantly associated with the differentiation grade, TNM stage, liver cirrhosis, and HBV infection. The reason may be closely related to the concept of CTNNB1 mutations in this meta-analysis refer to loss-of-function mutation.
However, our study has some limitations. Firstly, the clinical data used in this study were acquired from a relatively small cohort in each study, so selection bias or potential biases related to imbalanced clinical characteristics is inevitable. This will also lead to heterogeneity, which is a potential problem that may affect the results of all meta-analyses. In this study, significant heterogeneity was found when discussing the relationship between CTNNB1 mutations and 5-year overall survival and tumor size in the selected studies. Unfortunately, due to limited information, a meta-regression analysis could not be conducted. To eliminate variations across the included studies, we used a random effects model. Although this method may not completely eliminate the effects of heterogeneity, its adverse effects must be weakened. The second was publication bias, which can be seen in the publication bias evaluation. As is well known, this bias was unavoidable because positive results were more likely to be published than negative ones. Moreover, most studies included in this meta-analysis failed to elucidate the relationship between CTNNB1 expression and CTNNB1 mutation. Some studies provided the number of tissue samples with CTNNB1 expression, but failed to provide information on the gene type of CTNNB1, and vice versa. What’s more, the method of detecting CTNNB1 mutations was not standardized from author to author. Direct sequencing of exon 3, mass array, DNA sequence electropherograms, Sanger sequencing, SSCP analysis of exon 3, and PCR were performed. Therefore, a more standardized analysis should be performed and more prospective works and experimental clinical research should be conducted. Last but not the least, how CTNNB1 mutation effect clinical outcomes and survival of patients with HCC and how it serves as a valuable prognostic predictor is indeterminacy. Further research on the regulation of CTNNB1 expression by the CTNNB1 mutation-related signaling pathway is needed and will help to elucidate the new mechanism of drug resistance, providing a theoretical basis for the prediction of drug sensitivity in HCC and the development and application of new therapeutic targets for reversing drug resistance.
In conclusion, CTNNB1 mutation could serve as a potential predictor for the clinical and prognosis of HCC patients and act as a novel useful biomarker of molecular targeted therapies for HCC.
Supplementary Materials
Author Contributions
GLL designed this study. GLL and GQS completed the study selection and data extraction, JL completed the work of literature quality assessment, MC and GLL mainly focused on statistical analysis, and GLL and GQS wrote the paper.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
This study was supported by Quzhou City Science and Technology Guidance Project (2020122,2019136) from Min Chen.
References
- 1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108. https://doi.org/10.3322/caac.21262 [PubMed]
- 2. Braillon A. Hepatocellular carcinoma. Lancet. 2012; 380:469. https://doi.org/10.1016/S0140-6736(12)61282-3 [PubMed]
- 3. Dhir M, Melin AA, Douaiher J, Lin C, Zhen WK, Hussain SM, Geschwind JF, Doyle MB, Abou-Alfa GK, Are C. A Review and Update of Treatment Options and Controversies in the Management of Hepatocellular Carcinoma. Ann Surg. 2016; 263:1112–25. https://doi.org/10.1097/SLA.0000000000001556 [PubMed]
- 4. Brown C. Targeted therapy: An elusive cancer target. Nature. 2016; 537:S106–8. https://doi.org/10.1038/537S106a [PubMed]
- 5. Cheng AL, Kang YK, Chen Z, Tsao CJ, Qin S, Kim JS, Luo R, Feng J, Ye S, Yang TS, Xu J, Sun Y, Liang H, et al. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: a phase III randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2009; 10:25–34. https://doi.org/10.1016/S1470-2045(08)70285-7 [PubMed]
- 6. Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF, de Oliveira AC, Santoro A, Raoul JL, Forner A, Schwartz M, Porta C, Zeuzem S, et al., and SHARP Investigators Study Group. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008; 359:378–90. https://doi.org/10.1056/NEJMoa0708857 [PubMed]
- 7. Carotenuto P, Fassan M, Pandolfo R, Lampis A, Vicentini C, Cascione L, Paulus-Hock V, Boulter L, Guest R, Quagliata L, Hahne JC, Ridgway R, Jamieson T, et al. Wnt signalling modulates transcribed-ultraconserved regions in hepatobiliary cancers. Gut. 2017; 66:1268–77. https://doi.org/10.1136/gutjnl-2016-312278 [PubMed]
- 8. Pez F, Lopez A, Kim M, Wands JR, Caron de Fromentel C, Merle P. Wnt signaling and hepatocarcinogenesis: molecular targets for the development of innovative anticancer drugs. J Hepatol. 2013; 59:1107–17. https://doi.org/10.1016/j.jhep.2013.07.001 [PubMed]
- 9. Monga SP. β-Catenin Signaling and Roles in Liver Homeostasis, Injury, and Tumorigenesis. Gastroenterology. 2015; 148:1294–310. https://doi.org/10.1053/j.gastro.2015.02.056 [PubMed]
- 10. Guichard C, Amaddeo G, Imbeaud S, Ladeiro Y, Pelletier L, Maad IB, Calderaro J, Bioulac-Sage P, Letexier M, Degos F, Clément B, Balabaud C, Chevet E, et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat Genet. 2012; 44:694–8. https://doi.org/10.1038/ng.2256 [PubMed]
- 11. Jia D, Dong R, Jing Y, Xu D, Wang Q, Chen L, Li Q, Huang Y, Zhang Y, Zhang Z, Liu L, Zheng S, Xia Q, et al. Exome sequencing of hepatoblastoma reveals novel mutations and cancer genes in the Wnt pathway and ubiquitin ligase complex. Hepatology. 2014; 60:1686–96. https://doi.org/10.1002/hep.27243 [PubMed]
- 12. Lee JM, Yang J, Newell P, Singh S, Parwani A, Friedman SL, Nejak-Bowen KN, Monga SP. β-Catenin signaling in hepatocellular cancer: Implications in inflammation, fibrosis, and proliferation. Cancer Lett. 2014; 343:90–7. https://doi.org/10.1016/j.canlet.2013.09.020 [PubMed]
- 13. Chen J, Liu J, Jin R, Shen J, Liang Y, Ma R, Lin H, Liang X, Yu H, Cai X. Cytoplasmic and/or nuclear expression of β-catenin correlate with poor prognosis and unfavorable clinicopathological factors in hepatocellular carcinoma: a meta-analysis. PLoS One. 2014; 9:e111885. https://doi.org/10.1371/journal.pone.0111885 [PubMed]
- 14. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010; 25:603–5. https://doi.org/10.1007/s10654-010-9491-z [PubMed]
- 15. Ding X, Yang Y, Han B, Du C, Xu N, Huang H, Cai T, Zhang A, Han ZG, Zhou W, Chen L. Transcriptomic characterization of hepatocellular carcinoma with CTNNB1 mutation. PLoS One. 2014; 9:e95307. https://doi.org/10.1371/journal.pone.0095307 [PubMed]
- 16. Lin ZZ, Jeng YM, Hu FC, Pan HW, Tsao HW, Lai PL, Lee PH, Cheng AL, Hsu HC. Significance of Aurora B overexpression in hepatocellular carcinoma. Aurora B Overexpression in HCC. BMC Cancer. 2010; 10:461. https://doi.org/10.1186/1471-2407-10-461 [PubMed]
- 17. Lu LC, Shao YY, Lee YH, Hsieh MS, Hsiao CH, Lin HH, Kao HF, Ma YY, Yen FC, Cheng AL, Hsu CH. β-catenin (CTNNB1) mutations are not associated with prognosis in advanced hepatocellular carcinoma. Oncology. 2014; 87:159–66. https://doi.org/10.1159/000362821 [PubMed]
- 18. Mao TL, Chu JS, Jeng YM, Lai PL, Hsu HC. Expression of mutant nuclear beta-catenin correlates with non-invasive hepatocellular carcinoma, absence of portal vein spread, and good prognosis. J Pathol. 2001; 193:95–101. https://doi.org/10.1002/1096-9896(2000)9999:9999<::AID-PATH720>3.0.CO;2-3 [PubMed]
- 19. Wong CM, Fan ST, Ng IOL. beta-Catenin mutation and overexpression in hepatocellular carcinoma: clinicopathologic and prognostic significance. Cancer. 2001; 92:136–45. https://doi.org/10.1002/1097-0142(20010701)92:1<136::aid-cncr1301>3.0.co;2-r [PubMed]
- 20. Yuan RH, Chang KT, Chen YL, Hsu HC, Lee PH, Lai PL, Jeng YM. S100P expression is a novel prognostic factor in hepatocellular carcinoma and predicts survival in patients with high tumor stage or early recurrent tumors. PLoS One. 2013; 8:e65501. https://doi.org/10.1371/journal.pone.0065501 [PubMed]
- 21. Cavard C, Terris B, Grimber G, Christa L, Audard V, Radenen-Bussiere B, Simon MT, Renard CA, Buendia MA, Perret C. Overexpression of regenerating islet-derived 1 alpha and 3 alpha genes in human primary liver tumors with beta-catenin mutations. Oncogene. 2006; 25:599–608. https://doi.org/10.1038/sj.onc.1208860 [PubMed]
- 22. Cieply B, Zeng G, Proverbs-Singh T, Geller DA, Monga SP. Unique phenotype of hepatocellular cancers with exon-3 mutations in beta-catenin gene. Hepatology. 2009; 49:821–31. https://doi.org/10.1002/hep.22695 [PubMed]
- 23. Hsu HC, Jeng YM, Mao TL, Chu JS, Lai PL, Peng SY. Beta-catenin mutations are associated with a subset of low-stage hepatocellular carcinoma negative for hepatitis B virus and with favorable prognosis. Am J Pathol. 2000; 157:763–70. https://doi.org/10.1016/s0002-9440(10)64590-7 [PubMed]
- 24. Huang H, Fujii H, Sankila A, Mahler-Araujo BM, Matsuda M, Cathomas G, Ohgaki H. Beta-catenin mutations are frequent in human hepatocellular carcinomas associated with hepatitis C virus infection. Am J Pathol. 1999; 155:1795–801. https://doi.org/10.1016/s0002-9440(10)65496-x [PubMed]
- 25. Kim YD, Park CH, Kim HS, Choi SK, Rew JS, Kim DY, Koh YS, Jeung KW, Lee KH, Lee JS, Juhng SW, Lee JH. Genetic alterations of Wnt signaling pathway-associated genes in hepatocellular carcinoma. J Gastroenterol Hepatol. 2008; 23:110–8. https://doi.org/10.1111/j.1440-1746.2007.05250.x [PubMed]
- 26. Laurent-Puig P, Legoix P, Bluteau O, Belghiti J, Franco D, Binot F, Monges G, Thomas G, Bioulac-Sage P, Zucman-Rossi J. Genetic alterations associated with hepatocellular carcinomas define distinct pathways of hepatocarcinogenesis. Gastroenterology. 2001; 120:1763–73. https://doi.org/10.1053/gast.2001.24798 [PubMed]
- 27. Li M, Zhao H, Zhang X, Wood LD, Anders RA, Choti MA, Pawlik TM, Daniel HD, Kannangai R, Offerhaus GJ, Velculescu VE, Wang L, Zhou S, et al. Inactivating mutations of the chromatin remodeling gene ARID2 in hepatocellular carcinoma. Nat Genet. 2011; 43:828–9. https://doi.org/10.1038/ng.903 [PubMed]
- 28. Park JY, Park WS, Nam SW, Kim SY, Lee SH, Yoo NJ, Lee JY, Park CK. Mutations of beta-catenin and AXIN I genes are a late event in human hepatocellular carcinogenesis. Liver Int. 2005; 25:70–6. https://doi.org/10.1111/j.1478-3231.2004.0995.x [PubMed]
- 29. Taniguchi K, Roberts LR, Aderca IN, Dong X, Qian C, Murphy LM, Nagorney DM, Burgart LJ, Roche PC, Smith DI, Ross JA, Liu W. Mutational spectrum of beta-catenin, AXIN1, and AXIN2 in hepatocellular carcinomas and hepatoblastomas. Oncogene. 2002; 21:4863–71. https://doi.org/10.1038/sj.onc.1205591 [PubMed]
- 30. Tornesello ML, Buonaguro L, Tatangelo F, Botti G, Izzo F, Buonaguro FM. Mutations in TP53, CTNNB1 and PIK3CA genes in hepatocellular carcinoma associated with hepatitis B and hepatitis C virus infections. Genomics. 2013; 102:74–83. https://doi.org/10.1016/j.ygeno.2013.04.001 [PubMed]
- 31. Zucman-Rossi J, Benhamouche S, Godard C, Boyault S, Grimber G, Balabaud C, Cunha AS, Bioulac-Sage P, Perret C. Differential effects of inactivated Axin1 and activated beta-catenin mutations in human hepatocellular carcinomas. Oncogene. 2007; 26:774–80. https://doi.org/10.1038/sj.onc.1209824 [PubMed]
- 32. Xu C, Xu Z, Zhang Y, Evert M, Calvisi DF, Chen X. β-Catenin signaling in hepatocellular carcinoma. J Clin Invest. 2022; 132:e154515. https://doi.org/10.1172/JCI154515 [PubMed]
- 33. Qu C, He D, Lu X, Dong L, Zhu Y, Zhao Q, Jiang X, Chang P, Jiang X, Wang L, Zhang Y, Bi L, He J, et al. Salt-inducible Kinase (SIK1) regulates HCC progression and WNT/β-catenin activation. J Hepatol. 2016; 64:1076–89. https://doi.org/10.1016/j.jhep.2016.01.005 [PubMed]
- 34. Schulze K, Imbeaud S, Letouzé E, Alexandrov LB, Calderaro J, Rebouissou S, Couchy G, Meiller C, Shinde J, Soysouvanh F, Calatayud AL, Pinyol R, Pelletier L, et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat Genet. 2015; 47:505–11. https://doi.org/10.1038/ng.3252 [PubMed]
- 35. Mo Z, Wang Y, Cao Z, Li P, Zhang S. An Integrative Analysis Reveals the Underlying Association Between CTNNB1 Mutation and Immunotherapy in Hepatocellular Carcinoma. Front Oncol. 2020; 10:853. https://doi.org/10.3389/fonc.2020.00853 [PubMed]
- 36. Wang Z, Sheng YY, Gao XM, Wang CQ, Wang XY, Lu XU, Wei JW, Zhang KL, Dong QZ, Qin LX. β-catenin mutation is correlated with a favorable prognosis in patients with hepatocellular carcinoma. Mol Clin Oncol. 2015; 3:936–40. https://doi.org/10.3892/mco.2015.569 [PubMed]
- 37. Nault JC, Rebouissou S, Zucman Rossi J. NRF2/KEAP1 and Wnt/β-catenin in the multistep process of liver carcinogenesis in humans and rats. Hepatology. 2015; 62:677–9. https://doi.org/10.1002/hep.27828 [PubMed]
- 38. Wang W, Pan Q, Fuhler GM, Smits R, Peppelenbosch MP. Action and function of Wnt/β-catenin signaling in the progression from chronic hepatitis C to hepatocellular carcinoma. J Gastroenterol. 2017; 52:419–31. https://doi.org/10.1007/s00535-016-1299-5 [PubMed]
- 39. Chowdhury MK, Montgomery MK, Morris MJ, Cognard E, Shepherd PR, Smith GC. Glucagon phosphorylates serine 552 of β-catenin leading to increased expression of cyclin D1 and c-Myc in the isolated rat liver. Arch Physiol Biochem. 2015; 121:88–96. https://doi.org/10.3109/13813455.2015.1048693 [PubMed]
- 40. Xiao X, Mo H, Tu K. CTNNB1 mutation suppresses infiltration of immune cells in hepatocellular carcinoma through miRNA-mediated regulation of chemokine expression. Int Immunopharmacol. 2020; 89:107043. https://doi.org/10.1016/j.intimp.2020.107043 [PubMed]
- 41. Akasu M, Shimada S, Kabashima A, Akiyama Y, Shimokawa M, Akahoshi K, Kudo A, Yamaoka S, Tanabe M, Tanaka S. Intrinsic activation of β-catenin signaling by CRISPR/Cas9-mediated exon skipping contributes to immune evasion in hepatocellular carcinoma. Sci Rep. 2021; 11:16732. https://doi.org/10.1038/s41598-021-96167-0 [PubMed]
- 42. Aoki T, Nishida N, Kudo M. Clinical Significance of the Duality of Wnt/β-Catenin Signaling in Human Hepatocellular Carcinoma. Cancers (Basel). 2022; 14:444. https://doi.org/10.3390/cancers14020444 [PubMed]
- 43. Alomar SY, Mansour L, Abuderman A, Alkhuriji A, Arafah M, Alwasel S, Harrath AH, Almutairi M, Trayhyrn P, Dar JA. β-Catenin accumulation and S33F mutation of CTNNB1 gene in colorectal cancer in Saudi Arabia. Pol J Pathol. 2016; 67:156–62. https://doi.org/10.5114/pjp.2016.61452 [PubMed]
- 44. Wands JR, Kim M. WNT/β-catenin signaling and hepatocellular carcinoma. Hepatology. 2014; 60:452–54. https://doi.org/10.1002/hep.27081 [PubMed]
- 45. Ding Q, Xia W, Liu JC, Yang JY, Lee DF, Xia J, Bartholomeusz G, Li Y, Pan Y, Li Z, Bargou RC, Qin J, Lai CC, et al. Erk associates with and primes GSK-3beta for its inactivation resulting in upregulation of beta-catenin. Mol Cell. 2005; 19:159–70. https://doi.org/10.1016/j.molcel.2005.06.009 [PubMed]
- 46. Cui J, Zhou XD, Liu YK, Tang ZY, Zile MH. Abnormal beta-catenin gene expression with invasiveness of primary hepatocellular carcinoma in China. World J Gastroenterol. 2001; 7:542–46. https://doi.org/10.3748/wjg.v7.i4.542 [PubMed]
- 47. Wang J, Yu H, Dong W, Zhang C, Hu M, Ma W, Jiang X, Li H, Yang P, Xiang D. N6-Methyladenosine-Mediated Up-Regulation of FZD10 Regulates Liver Cancer Stem Cells’ Properties and Lenvatinib Resistance Through WNT/β-Catenin and Hippo Signaling Pathways. Gastroenterology. 2023; 164:990–1005. https://doi.org/10.1053/j.gastro.2023.01.041 [PubMed]
- 48. Lin D, Chen Y, Koksal AR, Dash S, Aydin Y. Targeting ER stress/PKA/GSK-3β/β-catenin pathway as a potential novel strategy for hepatitis C virus-infected patients. Cell Commun Signal. 2023; 21:102. https://doi.org/10.1186/s12964-023-01081-9 [PubMed]
- 49. Chakraborty E, Sarkar D. Emerging Therapies for Hepatocellular Carcinoma (HCC). Cancers (Basel). 2022; 14:2798. https://doi.org/10.3390/cancers14112798 [PubMed]
- 50. Kwee SA, Tiirikainen M. Beta-catenin activation and immunotherapy resistance in hepatocellular carcinoma: mechanisms and biomarkers. Hepatoma Res. 2021; 7:8. https://doi.org/10.20517/2394-5079.2020.124 [PubMed]
- 51. Ogawa K, Kanzaki H, Chiba T, Ao J, Qiang N, Ma Y, Zhang J, Yumita S, Ishino T, Unozawa H, Kan M, Iwanaga T, Nakagawa M, et al. Effect of Atezolizumab plus Bevacizumab in Patients with Hepatocellular Carcinoma Harboring CTNNB1 Mutation in Early Clinical Experience. J Cancer. 2022; 13:2656–61. https://doi.org/10.7150/jca.71494 [PubMed]
- 52. Chen L, Zhou Q, Liu J, Zhang W. CTNNB1 Alternation Is a Potential Biomarker for Immunotherapy Prognosis in Patients With Hepatocellular Carcinoma. Front Immunol. 2021; 12:759565. https://doi.org/10.3389/fimmu.2021.759565 [PubMed]


