Introduction
Although the aging process is the leading cause of human mortality and morbidity, being associated with several diseases, scientists still debate its causes and mechanisms [1–3]. Among the biological pathways associated with aging, we can highlight stem cell exhaustion, which argues that during normal aging, the decrease in the number or activity of these cells contributes to physiological dysfunction in aged tissues [4]. This concept is supported by the observation that aging is associated with reduced tissue renewal and repair at advanced ages [5]. Moreover, longevity manipulations in mice often affect growth and cell division, which has been hypothesized to relate to stem cells [6].
Despite their importance, in vivo detection and quantification of stem cells are challenging, which makes it difficult to study their association with aging, especially in humans [7]. In this context, detecting stemness-associated expression signatures is a promising strategy for studying stem cell biology. Stemness refers to a distinctive attribute marked by a series of molecular processes that delineate the essential properties of stem cells, enabling the generation of daughter cells and self-renewal. While widely employed in oncology, the exploration of this concept in gerontology has been comparatively limited [8–10].
In this study, we applied a machine learning method to detect stemness signatures from transcriptome data of healthy human tissues (see Methods). The methodology, developed by Malta et al. [9], was trained on stem cell classes and their differentiated progenitors and went through rigorous validation steps by Malta et al., including tests in several datasets from tumor and non-tumor samples. Although initially used to study oncogenic dedifferentiation, this approach has also been employed to study normal and pathological (non-tumorous) samples [11–15]. Therefore, we first downloaded expression data of 17,382 samples, divided into 30 tissues aged between 20 and 79 years, from GTEx in transcripts per million (TPM) [16]. After that, we followed the methodology by Malta et al. and assigned a stemness score to all GTEx samples [9]. Briefly, the stemness score varies from 0 (lowest stemness of the samples) to 1 (highest stemness of the samples). All the data generated are in Supplementary Table 1 and include the stemness score and clinical data from GTEx.
Results
First, in Figure 1A, we show the distribution of stemness in the 27 human tissues with at least 50 samples. We observe that the highest stemness scores are in testis, which are known to have a higher number of stem cells [17]. Notably, blood ranks as the second tissue with the highest stemness. Even though most hematopoietic stem cells (HSC) are typically located in the bone marrow, a fraction can be identified in the bloodstream, presenting direct potential for clinical applications [18, 19].
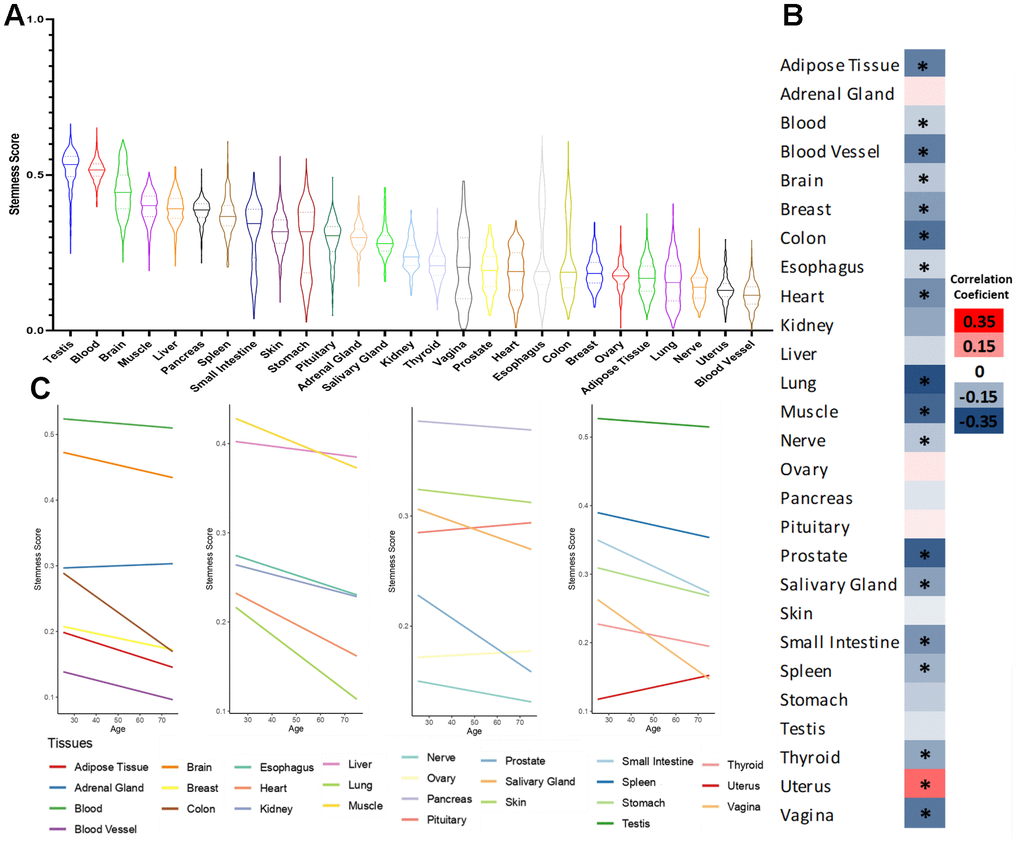
Figure 1. Stemness levels during human aging. (A) Distribution of stemness in human tissues. (B) Heatmap of Pearson’s correlation coefficient between stemness scores and age in human tissues. (C) Linear trend between stemness scores and age in human tissues. *FDR <0.05.
Then, we analyzed the relationship between stemness and aging. In Figure 1B, we have a heatmap of the correlation coefficient (Pearson correlation test followed by Benjamini-Hochberg correction, Supplementary Table 2) between stemness score and age, and in Figure 1C, we show the linear trend of the same variables. We observe a pan-tissue loss of stemness in most tissues studied, with the only significant exception being the uterus. Interestingly, our group previously showed that the uterus also tends to behave differently concerning cellular senescence and gene expression signatures during aging [20, 21].
As death and sex can influence gene expression (and consequently the stemness score) [22], we conducted a linear model where we adjusted the stemness variation (log2FC) with the variables present in GTEx (see Methods for details). With this model, we observe a variation in stemness, which aligns with the direction of the age correlation results, indicating a decrease in stemness with age in most tissues (Supplementary Table 3). The only tissues that exhibit a different behavior are the liver and nerve, which, overall, corroborates our earlier observation.
Moreover, we constructed a correlation matrix (Figure 2A) to examine the association between stemness in different tissues from the same individual. Most correlations are positive, suggesting that stemness is a global attribute of the organism. A noteworthy exception is the brain, whose stemness exhibits a flat or negative correlation with other tissues.
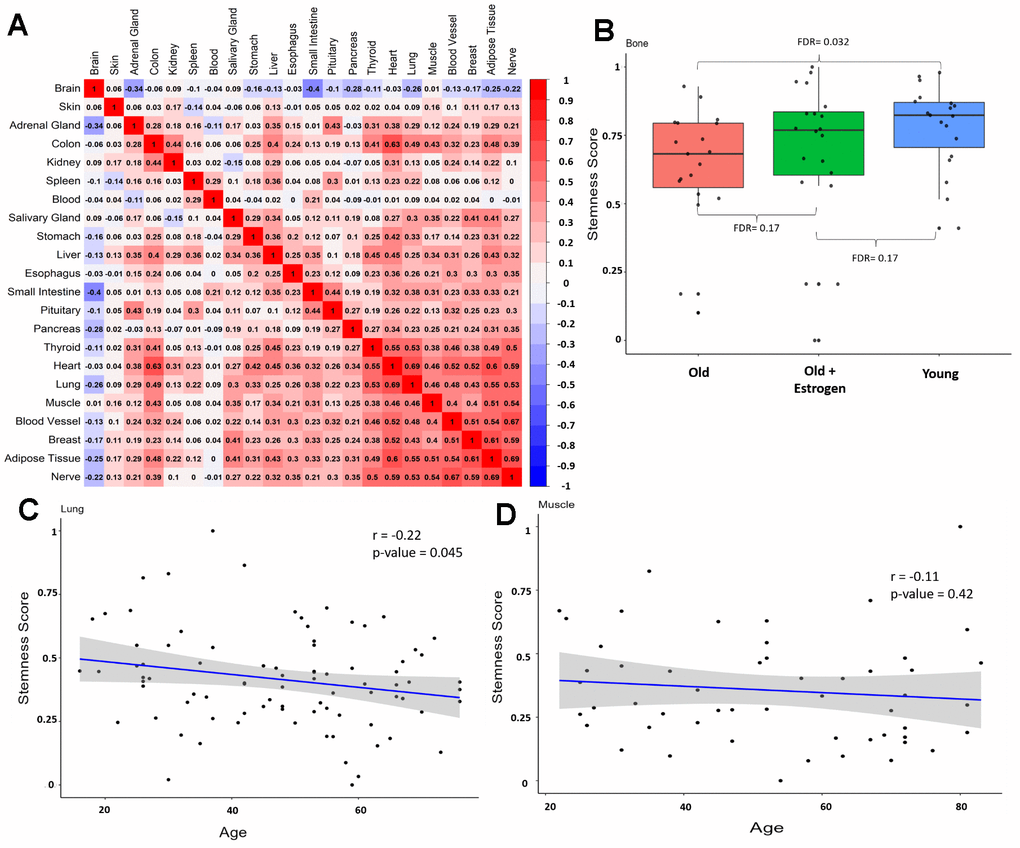
Figure 2. Stemness in same individual and validation datasets. (A) Correlation matrix between stemness across tissues from the same individual (excluding sex-related tissues) from GTEx. Numbers in each square represent the Pearson correlation coefficient. (B) Stemness levels of different groups in the alternative bone dataset, data from Weivoda et al. [30]. (C) Correlation between stemness and aging in an alternative lung dataset (r = -0.22, p-value = 0.045), data from Lee et al. [31]. (D) Correlation between stemness and aging in an alternative muscle dataset (r = -0.11, p-value = 0.42), data from Tumasian et al. [32].
To validate the GTEx results, we used additional datasets for lung, bone, and muscle (Figure 2B–2D). In bone, we observe a difference between the old and young groups (FDR = 0.032), and notably, women treated with estrogen replacement seem to have more stemness than the “natural” old group, suggesting that physiological variations (in this case, hormonal) can affect stemness (Figure 2B). Also, in lungs there is a significant decline in stemness with aging (Figure 2C) and a declining trend in muscle (Figure 2D). Taken together, these results indicate a trend toward a decline in stemness in human tissues during aging. Additionally, sample size appeared to be crucial in our analyses, and future studies should consider individual parameters, beyond age, that influence stemness.
To explore potential mechanisms, we associated stemness with cell proliferation and senescence, two processes associated with aging. We investigated the correlation between stemness and cell proliferation by acquiring gene expression data (in TPM) for the MKI67 proliferation marker from GTEx. Subsequently, we conducted a correlation analysis between the MKI67 expression and the stemness score of the samples. Figure 3A shows the correlation between stemness and proliferation in all GTEx samples (Pearson’s correlation test). Figure 3B shows a heatmap of the correlation coefficient in all tissues from GTEx (Pearson’s correlation test followed by Benjamini-Hochberg correction).
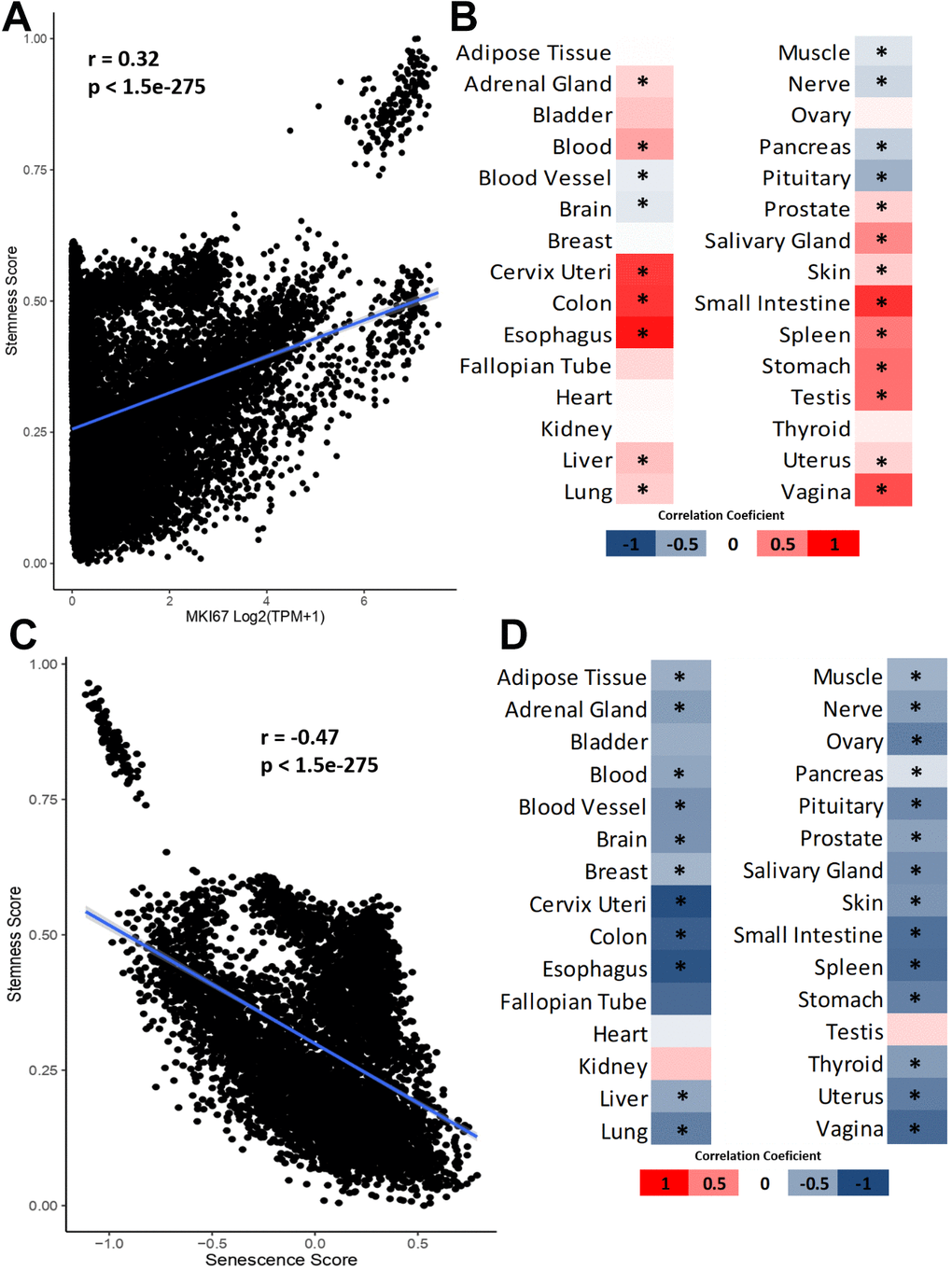
Figure 3. Relationship between stemness, cellular proliferation and senescence. (A) Correlation between stemness score and MKI67 expression. (B) Heatmap of Pearson’s correlation coefficient between stemness scores and MKI67 expression. (C) Correlation between stemness score and senescence scores. (D) Heatmap of Pearson’s correlation coefficient between stemness score and senescence score. *FDR <0.05.
We observe a general positive correlation trend between stemness and cell proliferation, but this correlation is not observed in all tissues as exceptions include blood vessel, brain, muscle, nerve, pancreas, and pituitary results (Figure 3A, 3B). To validate this result, we employed the gene list provided in the study by Ramaker et al. [23]. In summary, the authors established and validated a proliferation index for healthy tissues and cells using the gene list from Venet et al. [24]. We correlated it with the stemness score using the average expression of all genes in the proliferation list, thereby obtaining a proliferation value per sample. The results reveal an even greater number of positive correlations, but still demonstrate negative and significant correlations, now specifically for the pancreas and pituitary (Supplementary Figure 1A). All the results of these correlations can be found in Supplementary Table 4. This suggests that, although cell proliferation is important for stemness, their relationship is complex.
Then, we used Wang et al. senescence score data to study the association between stemness and cellular senescence [25]. In total, we have 7123 samples with stemness and senescence values simultaneously, and the correlations were performed as before. We observe a negative correlation between stemness and senescence when considering all available GTEx samples (Figure 3C) and when we separate by tissue (Figure 3D) without any significant exception. Similar to the validation for cell proliferation, we employed the SenMayo genes to obtain an alternative senescence signature [26]. Subsequently, we correlated it with the stemness score (Supplementary Figure 1B). The results are similar, with all significant correlations between stemness and senescence being negative. All the results of these correlations can be found in Supplementary Table 5. These results indicate that although senescent cells and stem cells are not technically opposite states, they behave in opposite ways in vivo at the transcriptomic level.
Finally, GTEx samples come from bulk RNAseq containing multiple cell types. It is plausible that varying tissue types harbor distinct ratios of stem cells to somatic cells, thereby influencing the stemness score. However, the functional understanding of how stemness variation in stem cells concerning age remains elusive. Thus, it is uncertain whether there exists any age-related difference in the stemness attribute specifically within stem cells from older individuals. To address this uncertainty, we investigated whether the directly measured stemness in stem cells exhibits variation with age. To do so, we utilized data from Adelman et al. [27] and compared stemness levels directly in HSC isolated from two age groups: young (ages 18-30) and old (ages 65-75).
We observed a quite unexpected result; despite not having a statistically significant difference, we see that both the comparison between the groups (Figure 4A, p=0.07) and the direct correlation (Figure 4B, r=0.42 and p=0.068), have an indicative trend of HSC from older donors to have more stemness. This result is interesting when we consider that literature shows evidence of increasing HSC numbers with age, which may suggest that age-related stem cell problems are functional rather than quantitative [27]. Nevertheless, more robust studies must be performed before we can draw more assertive conclusions.
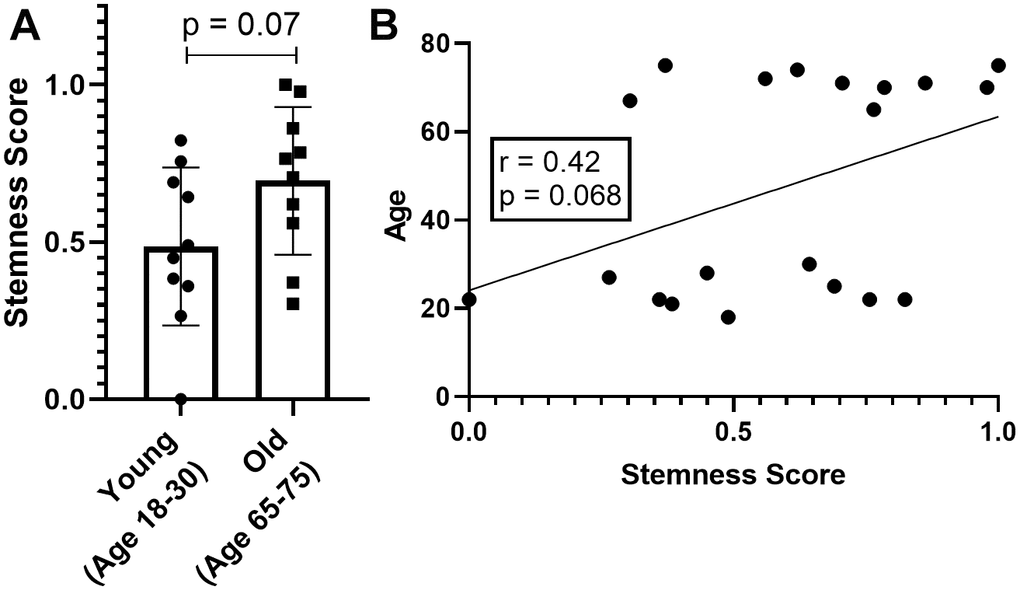
Figure 4. Stemness levels in hematopoietic stem cells. (A) Direct comparison between young and old groups. (B) Correlation between stemness score and age of the HSC donors.
Discussion
In this work, we studied the relationship between stemness and aging in humans. We found a loss of stemness in most aged tissues, which corroborates that stem cell exhaustion is important for aging. Although stem cell depletion in aging has already been observed in some tissues, such as satellite cells in the muscle [28], and hematopoietic stem cells in blood and bone marrow [29], as far as we know, we are the first to provide evidence of this in a pan-tissue manner.
We also show a positive trend between stemness and cell proliferation, but it is not global, with exceptions in some tissues. When we consider cellular senescence, we observe that the two phenomena are opposite in almost all tissues at the transcriptomic level.
It is important to note that many questions remain open. Our analyses are mainly correlation tests; therefore, we cannot determine cause and effect. This is important as we cannot assure whether the loss of stemness contributes to aging or is a cause or response of aging processes. Furthermore, we do not know whether the decrease in stemness a direct reduction of the stem cell pool is or refers to intrinsic characteristics of different cells in the tissue. In this sense, our HSC analyses corroborate a functional problem in aged stem cells, at least in blood stem cells. Ultimately, it is crucial to determine what drives these changes and which patterns and genes are associated with this process. Here is essential to highlight some papers that suggest that epigenetic modifications regulate stemness and could be a promising area for future studies [33, 34]. Additionally, our analyses suggest that stemness is a global attribute of the organism which correlates across tissues (except the brain), and future analyses may investigate how genetic polymorphisms contribute to individual stemness. Specifically, genome-wide association studies could be insightful in elucidating mechanisms associated with stemness. Additionally, the correlation of stemness among tissues could reflect systemic factors such as hormones (as suggested by the effect of estrogen in Figure 2B) or inflammation that impact cell proliferation [35, 36]. This presents three potential scenarios: the pan-tissue decrease in stemness may indicate either cell-intrinsic mechanisms, a systemic phenomenon, or a combination of both.
One limitation of our study is that it is based on a data-driven inference of stemness from transcriptomic signatures. Additionally, our method does not consider differences between types of stem cells in different tissues. Ideally, experimental validation will be necessary to corroborate our findings. Nevertheless, it is currently challenging to delineate an approach to measure stemness in aged human tissues experimentally, and thus we believe that our work can serve as a starting point for future research.
In conclusion, we provide the first evidence of a pan-tissue decrease of stemness during human aging and report an association between stemness and cell proliferation and senescence. This study also assigned a stemness score to more than 17,000 human samples, and these data can be useful for the scientific community for further studies.
Materials and Methods
Transcriptome data from healthy tissues
RNA-Seq-based gene expression data from human tissues were downloaded from the GTEx portal (https://gtexportal.org) in transcripts per million (version 8) [16]. The raw RNA-Seq data were previously aligned to the human reference genome GRCH38/hg38 by the GTEx consortium.
We reduced the number of genes from the GTEx data to the same pairs found in the training matrix, i.e., we retained only the genes utilized by Malta et al. [9]. to calculate stemness. The training matrix is generated through a machine learning methodology (see below) and is the same as that in the original stemness paper by Malta et al. [9]. The resulting matrix contained 12,471 mRNA expression values and was used to calculate the stemness score. Here it is important to note that GTEx does not provide the actual age of each sample but rather age ranges (20-29, 30-39, 40-49, 50-59, 60-69, and 70-79). We then approximate the age of each sample to 25, 35, 45, 55, 65 and 75 years, respectively, as previously [20].
Stemness score
To assign the stemness score of the GTEx samples we used a machine learning methodology built by Malta et al. [9].
In brief, Malta et al. built a predictive model using a one-class logistic regression on the pluripotent stem cell samples (ESC and iPSC) from the PCBC dataset [37–39]. The data were mean-centered, and the logistic regression was applied to the stem cell labeled samples to obtain the training signature. We then applied this signature to the GTEx transcriptome data, using Spearman correlations between the model’s weight vector and the sample’s expression profile. As a result, we have a stemness score for all GTEx samples ranging from low (zero) to high (one) stemness. Further details and validation of the methodology can be found in the original study [9].
Additionally, the highest stemness scores observed in Figure 1 (below 0.7) are due to the highest values >0.7 being associated with EBV-transformed lymphocyte samples, which are not represented in the figure. The GTEx data is segmented into sub-regions known as “tissue detail,” particularly within blood and skin, including cells derived from patients, fibroblasts, and lymphocytes transformed with EBV. Our study focuses on the overall stemness during aging, emphasizing major tissues without subdivisions. Cell lines from skin and blood were excluded in downstream analyses to prevent significant data distortion, although we calculated their stemness scores for potential use in future studies. Notably, transformed cells, expected to exhibit stem-like behavior compared to tissues, consistently display the highest stemness scores.
All this process was done using R and the code available on GitHub associated with the original paper [9]. We followed the author’s guidelines step by step.
Linear model
To confirm that the stemness effect was not an artifact resulting from the interference of cause of death, sex, or tissue subregion, we applied a linear model similar to what we had done previously [40].
For each tissue, fold change with age was calculated using the model below. If any variable is not present in the tissue (e.g., sex for vagina or region for thyroid), it is disregarded in the analysis. All information on the subjects was taken directly from the GTEx portal (https://gtexportal.org/home/datasets).
The variables are defined as follows:
Y_ij: The stemness score j in sample i.
Age_i: The age of sample i – continuous variable.
Sex_i: The sex of sample i – categorical variable.
Death_i: The death classification of sample i based on the 4-point Hardy scale – categorical variable [41].
Region_i: The tissue region cells were extracted from for sample i – categorical variable.
ε_ij: The error term for stemness score i in sample j.
Linear model was generated using the R package limma, using the lmFit() function [42, 43].
Correlation matrix
To assess the stemness behavior between tissues across the same individual, we constructed a correlation matrix using Pearson coefficients. For the few cases in which a specific subject had two or more samples from the same tissue, we used the average stemness score to calculate the correlation matrix.
For the Figure 2, we employed the R package ‘corrplot’ with hierarchical clustering option [44]. In this analysis, we excluded sex tissues (testis, prostate, vagina, uterus, and ovary) as they cannot be correlated with each other and would interfere with the hierarchical classification. In Supplementary Figure 2, we included the correlation matrix with all GTEx samples with more than 50 samples, without clustering.
Validation datasets
For validation, we selected three datasets on the aging of healthy tissues. For lung data, we utilized the dataset from Lee et al. [31], which comprises 86 samples with an age range of 16 to 76 years. For muscle data, we utilized the dataset from Tumasian et al. [32] which comprises 53 samples with an age range of 22 to 81 years. The bone data is from Weivoda et al. [30] and consists of 58 female samples divided into three groups: 19 young women (30.3 ± 5.4 years), 19 old women (73.1 ± 6.6 years), and 20 old women treated with 3 weeks of Estrogen therapy (70.5 ± 5.2 years).
All datasets are from bulk RNA-seq, like GTEx. Additionally, the stemness score was calculated independently for each tissue, following the same procedure as in the GTEx analysis. For correlations, we used the Pearson coefficient, and for the analysis of bone data, we employed the Kruskal-Wallis test.
Correlation with senescence and cell proliferation
To correlate the stemness score with cell proliferation, we initially retrieved the expression data for the proliferative marker MKI67 from the GTEx database in TPM (transcripts per million). To validate this result, we employed the gene list provided in the study by Ramaker et al. [23]. In summary, the authors established and validated a proliferation index for healthy tissues and cells using the gene list from Venet et al. [24]. The gene list for constructing the proliferation index was identified from the top 1% of genes most positively correlated with the proliferation marker PCNA (proliferating cell nuclear antigen) across 36 healthy tissues. We applied the same gene list and calculated the average expression of all genes within the proliferation list, thus obtaining a proliferation value per sample for correlation with the stemness score.
For the correlation with cellular senescence, we utilized the study by Wang et al. [25], which calculated a senescence score for GTEx samples. Again, for validation, we employed the SenMayo gene list [26], one of the most used by the scientific community for senescence studies. We employed the same strategy as before, using the average expression of SenMayo genes to correlate with the stemness score.
Analysis of hematopoietic stem cells
We first downloaded normalized counts of bulk RNA-seq from the work of Adelman et al. [27]. In this study, the authors isolated human hematopoietic stem cells from two groups with ten samples each: young (age 18-30) and old (age 65-75). We applied the same approach as above to generate stemness scores in these samples. We then compared stemness with age by direct comparison between the two groups (Student’s t-test) and by Pearson correlation. The graphs for this analysis were constructed using GraphPad Prism 8.
Statistical analysis and graphs
We applied Pearson correlations to all analyses performed in this work using R functions. In tissue-specific analyses, the p-value was corrected using Benjamini-Hochberg’s FDR methodology. All correlations coefficients, p-values and FDR are presented in Supplementary Tables 2, 4, 5.
Linear trends and the respective graphs were built using the ggplot2 (version 3.3.6) with standard parameters [45]. Heatmaps and violin plot were built using Microsoft Excel and GraphPad Prism 8, respectively.
Supplementary Materials
Author Contributions
GAS: Initial concept of the study, conducted the analyses, discussion and interpretation of results, and writing and editing the manuscript. GDVM: Conducted and reviewed the analyses, contributed to the discussion and interpretation of results. JPM: Initial concept and supervision of the study, discussion and interpretation of results, review and manuscript writing.
Acknowledgments
We are grateful to current and past members of the Genomics of Ageing and Rejuvenation Lab for valuable discussions.
Conflicts of Interest
JPM is CSO of YouthBio Therapeutics, an advisor/consultant for the Longevity Vision Fund, 199 Biotechnologies, and NOVOS, and the founder of Magellan Science Ltd, a company providing consulting services in longevity science.
Funding
Gabriel Arantes dos Santos is financed by the scholarship “Bolsa de Excelência em Medicina Domingos Nelson Martins” of the Faculty of Medicine of the University of São Paulo (FMUSP). Work in our lab is supported by grants from the Wellcome Trust (208375/Z/17/Z), Longevity Impetus Grants, LongeCity and the Biotechnology and Biological Sciences Research Council (BB/R014949/1 and BB/V010123/1).
References
- 1. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. Hallmarks of aging: An expanding universe. Cell. 2023; 186:243–78. https://doi.org/10.1016/j.cell.2022.11.001 [PubMed]
- 2. Gems D, de Magalhães JP. The hoverfly and the wasp: A critique of the hallmarks of aging as a paradigm. Ageing Res Rev. 2021; 70:101407. https://doi.org/10.1016/j.arr.2021.101407 [PubMed]
- 3. de Magalhães JP. Ageing as a software design flaw. Genome Biol. 2023; 24:51. https://doi.org/10.1186/s13059-023-02888-y [PubMed]
- 4. Rudolph KL. Stem cell aging. Mech Ageing Dev. 2021; 193:111394. https://doi.org/10.1016/j.mad.2020.111394 [PubMed]
- 5. Clevers H, Watt FM. Defining Adult Stem Cells by Function, not by Phenotype. Annu Rev Biochem. 2018; 87:1015–27. https://doi.org/10.1146/annurev-biochem-062917-012341 [PubMed]
- 6. de Magalhães JP, Faragher RG. Cell divisions and mammalian aging: integrative biology insights from genes that regulate longevity. Bioessays. 2008; 30:567–78. https://doi.org/10.1002/bies.20760 [PubMed]
- 7. Srivastava AK, Bulte JW. Seeing stem cells at work in vivo. Stem Cell Rev Rep. 2014; 10:127–44. https://doi.org/10.1007/s12015-013-9468-x [PubMed]
- 8. Mushtaq M, Kovalevska L, Darekar S, Abramsson A, Zetterberg H, Kashuba V, Klein G, Arsenian-Henriksson M, Kashuba E. Cell stemness is maintained upon concurrent expression of RB and the mitochondrial ribosomal protein S18-2. Proc Natl Acad Sci USA. 2020; 117:15673–83. https://doi.org/10.1073/pnas.1922535117 [PubMed]
- 9. Malta TM, Sokolov A, Gentles AJ, Burzykowski T, Poisson L, Weinstein JN, Kamińska B, Huelsken J, Omberg L, Gevaert O, Colaprico A, Czerwińska P, Mazurek S, et al, and Cancer Genome Atlas Research Network. Machine Learning Identifies Stemness Features Associated with Oncogenic Dedifferentiation. Cell. 2018; 173:338–54.e15. https://doi.org/10.1016/j.cell.2018.03.034 [PubMed]
- 10. Saygin C, Matei D, Majeti R, Reizes O, Lathia JD. Targeting Cancer Stemness in the Clinic: From Hype to Hope. Cell Stem Cell. 2019; 24:25–40. https://doi.org/10.1016/j.stem.2018.11.017 [PubMed]
- 11. Peng XY, Dong B, Liu X. Cancer metastasis is related to normal tissue stemness. PLoS One. 2022; 17:e0277811. https://doi.org/10.1371/journal.pone.0277811 [PubMed]
- 12. Lu GM, Rong YX, Liang ZJ, Hunag DL, Ma YF, Luo ZZ, Wu FX, Liu XH, Liu Y, Mo S, Qi ZQ, Li HM. Multiomics global landscape of stemness-related gene clusters in adipose-derived mesenchymal stem cells. Stem Cell Res Ther. 2020; 11:310. https://doi.org/10.1186/s13287-020-01823-3 [PubMed]
- 13. Devall M, Plummer SJ, Bryant J, Jennelle LT, Eaton S, Dampier CH, Huyghe JR, Peters U, Powell SM, Casey G. Ethanol exposure drives colon location specific cell composition changes in a normal colon crypt 3D organoid model. Sci Rep. 2021; 11:432. https://doi.org/10.1038/s41598-020-80240-1 [PubMed]
- 14. Ye Z, Zhang Y, Huang N, Chen S, Wu X, Li L. Immune repertoire and evolutionary trajectory analysis in the development of diabetic nephropathy. Front Immunol. 2022; 13:1006137. https://doi.org/10.3389/fimmu.2022.1006137 [PubMed]
- 15. Li Y, Ju S, Li X, Li W, Zhou S, Wang G, Cai Y, Dong Z. Characterization of the microenvironment of diabetic foot ulcers and potential drug identification based on scRNA-seq. Front Endocrinol (Lausanne). 2023; 13:997880. https://doi.org/10.3389/fendo.2022.997880 [PubMed]
- 16. GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020; 369:1318–30. https://doi.org/10.1126/science.aaz1776 [PubMed]
- 17. Kanatsu-Shinohara M, Shinohara T. Spermatogonial stem cell self-renewal and development. Annu Rev Cell Dev Biol. 2013; 29:163–87. https://doi.org/10.1146/annurev-cellbio-101512-122353 [PubMed]
- 18. Zhang Y, Huang B. Peripheral blood stem cells: phenotypic diversity and potential clinical applications. Stem Cell Rev Rep. 2012; 8:917–25. https://doi.org/10.1007/s12015-012-9361-z [PubMed]
- 19. Eaves CJ. Hematopoietic stem cells: concepts, definitions, and the new reality. Blood. 2015; 125:2605–13. https://doi.org/10.1182/blood-2014-12-570200 [PubMed]
- 20. Chatsirisupachai K, Palmer D, Ferreira S, de Magalhães JP. A human tissue-specific transcriptomic analysis reveals a complex relationship between aging, cancer, and cellular senescence. Aging Cell. 2019; 18:e13041. https://doi.org/10.1111/acel.13041 [PubMed]
- 21. Avelar RA, Ortega JG, Tacutu R, Tyler EJ, Bennett D, Binetti P, Budovsky A, Chatsirisupachai K, Johnson E, Murray A, Shields S, Tejada-Martinez D, Thornton D, et al. A multidimensional systems biology analysis of cellular senescence in aging and disease. Genome Biol. 2020; 21:91. https://doi.org/10.1186/s13059-020-01990-9 [PubMed]
- 22. Marttila S, Chatsirisupachai K, Palmer D, de Magalhães JP. Ageing-associated changes in the expression of lncRNAs in human tissues reflect a transcriptional modulation in ageing pathways. Mech Ageing Dev. 2020; 185:111177. https://doi.org/10.1016/j.mad.2019.111177 [PubMed]
- 23. Ramaker RC, Lasseigne BN, Hardigan AA, Palacio L, Gunther DS, Myers RM, Cooper SJ. RNA sequencing-based cell proliferation analysis across 19 cancers identifies a subset of proliferation-informative cancers with a common survival signature. Oncotarget. 2017; 8:38668–81. https://doi.org/10.18632/oncotarget.16961 [PubMed]
- 24. Venet D, Dumont JE, Detours V. Most random gene expression signatures are significantly associated with breast cancer outcome. PLoS Comput Biol. 2011; 7:e1002240. https://doi.org/10.1371/journal.pcbi.1002240 [PubMed]
- 25. Wang X, Ma L, Pei X, Wang H, Tang X, Pei JF, Ding YN, Qu S, Wei ZY, Wang HY, Wang X, Wei GH, Liu DP, Chen HZ. Comprehensive assessment of cellular senescence in the tumor microenvironment. Brief Bioinform. 2022; 23:bbac118. https://doi.org/10.1093/bib/bbac118 [PubMed]
- 26. Saul D, Kosinsky RL, Atkinson EJ, Doolittle ML, Zhang X, LeBrasseur NK, Pignolo RJ, Robbins PD, Niedernhofer LJ, Ikeno Y, Jurk D, Passos JF, Hickson LJ, et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat Commun. 2022; 13:4827. https://doi.org/10.1038/s41467-022-32552-1 [PubMed]
- 27. Adelman ER, Huang HT, Roisman A, Olsson A, Colaprico A, Qin T, Lindsley RC, Bejar R, Salomonis N, Grimes HL, Figueroa ME. Aging Human Hematopoietic Stem Cells Manifest Profound Epigenetic Reprogramming of Enhancers That May Predispose to Leukemia. Cancer Discov. 2019; 9:1080–101. https://doi.org/10.1158/2159-8290.CD-18-1474 [PubMed]
- 28. Sousa-Victor P, García-Prat L, Muñoz-Cánoves P. Control of satellite cell function in muscle regeneration and its disruption in ageing. Nat Rev Mol Cell Biol. 2022; 23:204–26. https://doi.org/10.1038/s41580-021-00421-2 [PubMed]
- 29. de Haan G, Lazare SS. Aging of hematopoietic stem cells. Blood. 2018; 131:479–87. https://doi.org/10.1182/blood-2017-06-746412 [PubMed]
- 30. Weivoda MM, Chew CK, Monroe DG, Farr JN, Atkinson EJ, Geske JR, Eckhardt B, Thicke B, Ruan M, Tweed AJ, McCready LK, Rizza RA, Matveyenko A, et al. Identification of osteoclast-osteoblast coupling factors in humans reveals links between bone and energy metabolism. Nat Commun. 2020; 11:87. https://doi.org/10.1038/s41467-019-14003-6 [PubMed]
- 31. Lee S, Islam MN, Boostanpour K, Aran D, Jin G, Christenson S, Matthay MA, Eckalbar WL, DePianto DJ, Arron JR, Magee L, Bhattacharya S, Matsumoto R, et al. Molecular programs of fibrotic change in aging human lung. Nat Commun. 2021; 12:6309. https://doi.org/10.1038/s41467-021-26603-2 [PubMed]
- 32. Tumasian RA 3rd, Harish A, Kundu G, Yang JH, Ubaida-Mohien C, Gonzalez-Freire M, Kaileh M, Zukley LM, Chia CW, Lyashkov A, Wood WH 3rd, Piao Y, Coletta C, et al. Skeletal muscle transcriptome in healthy aging. Nat Commun. 2021; 12:2014. https://doi.org/10.1038/s41467-021-22168-2 [PubMed]
- 33. Musella M, Guarracino A, Manduca N, Galassi C, Ruggiero E, Potenza A, Maccafeo E, Manic G, Mattiello L, Soliman Abdel Rehim S, Signore M, Pietrosanto M, Helmer-Citterich M, et al. Type I IFNs promote cancer cell stemness by triggering the epigenetic regulator KDM1B. Nat Immunol. 2022; 23:1379–92. https://doi.org/10.1038/s41590-022-01290-3 [PubMed]
- 34. Kim BN, Ahn DH, Kang N, Yeo CD, Kim YK, Lee KY, Kim TJ, Lee SH, Park MS, Yim HW, Park JY, Park CK, Kim SJ. TGF-β induced EMT and stemness characteristics are associated with epigenetic regulation in lung cancer. Sci Rep. 2020; 10:10597. https://doi.org/10.1038/s41598-020-67325-7 [PubMed]
- 35. Liu L, Zhang L, Li C, Qiu Z, Kuang T, Wu Z, Deng W. Effects of hormones on intestinal stem cells. Stem Cell Res Ther. 2023; 14:105. https://doi.org/10.1186/s13287-023-03336-1 [PubMed]
- 36. Vadivel S, Vincent P, Sekaran S, Visaga Ambi S, Muralidar S, Selvaraj V, Palaniappan B, Thirumalai D. Inflammation in myocardial injury- Stem cells as potential immunomodulators for myocardial regeneration and restoration. Life Sci. 2020; 250:117582. https://doi.org/10.1016/j.lfs.2020.117582 [PubMed]
- 37. Sokolov A, Paull EO, Stuart JM. ONE-CLASS DETECTION OF CELL STATES IN TUMOR SUBTYPES. Pac Symp Biocomput. 2016; 21:405–16. [PubMed]
- 38. Daily K, Ho Sui SJ, Schriml LM, Dexheimer PJ, Salomonis N, Schroll R, Bush S, Keddache M, Mayhew C, Lotia S, Perumal TM, Dang K, Pantano L, et al. Molecular, phenotypic, and sample-associated data to describe pluripotent stem cell lines and derivatives. Sci Data. 2017; 4:170030. https://doi.org/10.1038/sdata.2017.30 [PubMed]
- 39. Salomonis N, Dexheimer PJ, Omberg L, Schroll R, Bush S, Huo J, Schriml L, Ho Sui S, Keddache M, Mayhew C, Shanmukhappa SK, Wells J, Daily K, et al. Integrated Genomic Analysis of Diverse Induced Pluripotent Stem Cells from the Progenitor Cell Biology Consortium. Stem Cell Reports. 2016; 7:110–25. https://doi.org/10.1016/j.stemcr.2016.05.006 [PubMed]
- 40. Dos Santos GA, Chatsirisupachai K, Avelar RA, de Magalhães JP. Transcriptomic analysis reveals a tissue-specific loss of identity during ageing and cancer. BMC Genomics. 2023; 24:644. https://doi.org/10.1186/s12864-023-09756-w [PubMed]
- 41. Ferreira PG, Muñoz-Aguirre M, Reverter F, Sá Godinho CP, Sousa A, Amadoz A, Sodaei R, Hidalgo MR, Pervouchine D, Carbonell-Caballero J, Nurtdinov R, Breschi A, Amador R, et al. The effects of death and post-mortem cold ischemia on human tissue transcriptomes. Nat Commun. 2018; 9:490. https://doi.org/10.1038/s41467-017-02772-x [PubMed]
- 42. Phipson B, Lee S, Majewski IJ, Alexander WS, Smyth GK. ROBUST HYPERPARAMETER ESTIMATION PROTECTS AGAINST HYPERVARIABLE GENES AND IMPROVES POWER TO DETECT DIFFERENTIAL EXPRESSION. Ann Appl Stat. 2016; 10:946–63. https://doi.org/10.1214/16-AOAS920 [PubMed]
- 43. Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015; 43:e47. https://doi.org/10.1093/nar/gkv007 [PubMed]
- 44. Wei TSV. R package ‘corrplot’: Visualization of a Correlation Matrix. (Version 0.92), 2021. https://github.com/taiyun/corrplot.
- 45. Wickham H. gplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York. 2016.



