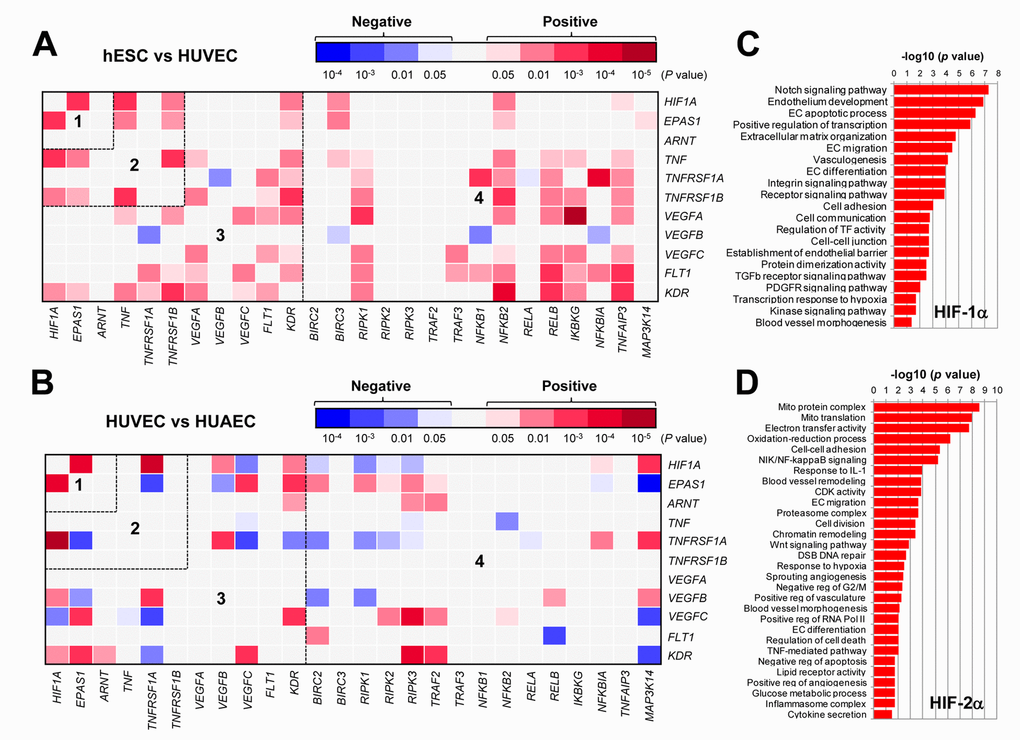
Figure 1. The GEP analyses map the potential correlations among the HIFs, TNFα, VEGFs, and NF-κB signaling pathways in ECs. (A, B) By analyzing the datasets involving gene expression profiling (GEP) in ECs using the R2: Genomics Analysis and Visualization Platform, the heatmaps were generated based on P values for analyses of correlations (red and blue indicating positive and negative correlations, respectively) between each pair of genes as indicated on longitudinal and transverse axes, in (A; dataset, Exp HUVEC vs ESC - James - 12 - MAS5.0 - u133p2) human stem cells (hESC) vs human umbilical vein endothelial cells (HUVEC) and (B; dataset, Normal Endothelial Cells HUAEC/HUVEC - Luttun - 38 - MAS5.0 - u133p2) HUVEC vs human umbilical artery endothelial cells (HUAEC). Numbers in the heatmaps indicate the areas (outlined by dash line) clustered for each pathway. (C, D) Gene ontology (GO) analyses were performed to categorize (C) HIF-1α- and (D) HIF-2α-related genes according to their functions, in all types of ECs, including (dataset, Normal Endothelial Cells HUAEC/HUVEC - Luttun - 38 - MAS5.0 - u133p2).