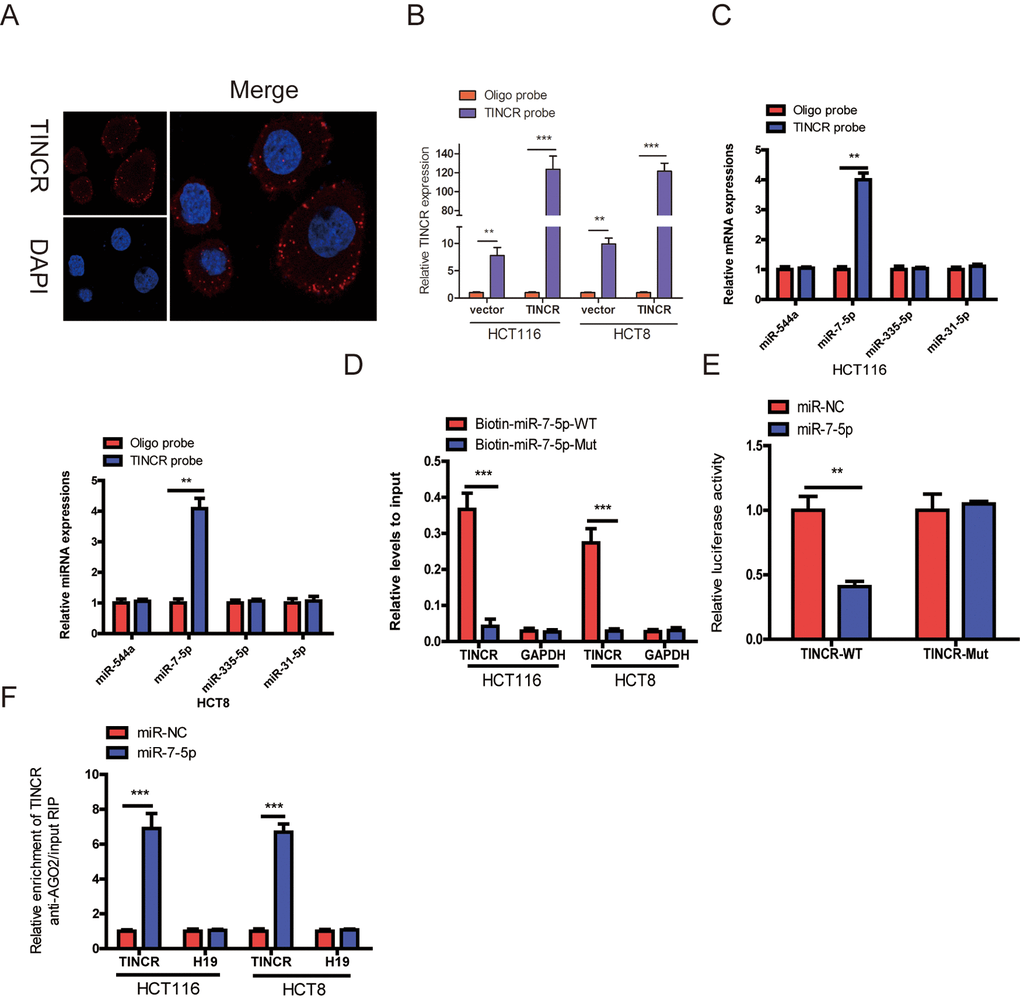
Figure 5.TINCR functioned as a ceRNA by sponging miR-7-5p. (A) RNA-FISH were employed to verify that TINCR was located mainly in the cytoplasm. (B) Lysates from HCT116 and HCT8 cells with TINCR overexpression were subject to biotinylated-TINCR pull-down assay and the expression of TINCR were measured by qRT-PCR. (C) The expression of four candidate miRNAs predicted by starbase database were quantified by qRT-PCR after the biotinylated-TINCR pull-down assay in HCT116 and HCT8 cells. (D) The biotinylated wild-type/mutant miR-7-5p was, respectively, transfected into HCT116 and HCT8 cells with TINCR overexpression. The expression of TINCR were measured by qRT-PCR after streptavidin capture. (E) Luciferase activity in HCT116 and HCT8 cells cotransfected with luciferase reporter containing TINCR sequences with wild-type or mutated miR-7-5p binding sites and miR-7-5p or its control. (F) Anti-AGO2 RIP was used in HCT116 and HCT8 cells overexpressing miR-7-5p, followed by qRT-PCR to assess the expression of TINCR or H19 (control) associated with AGO2. The data are presented as the mean±S.D. of three independent experiments. **P<0.01. ***P<0.001.