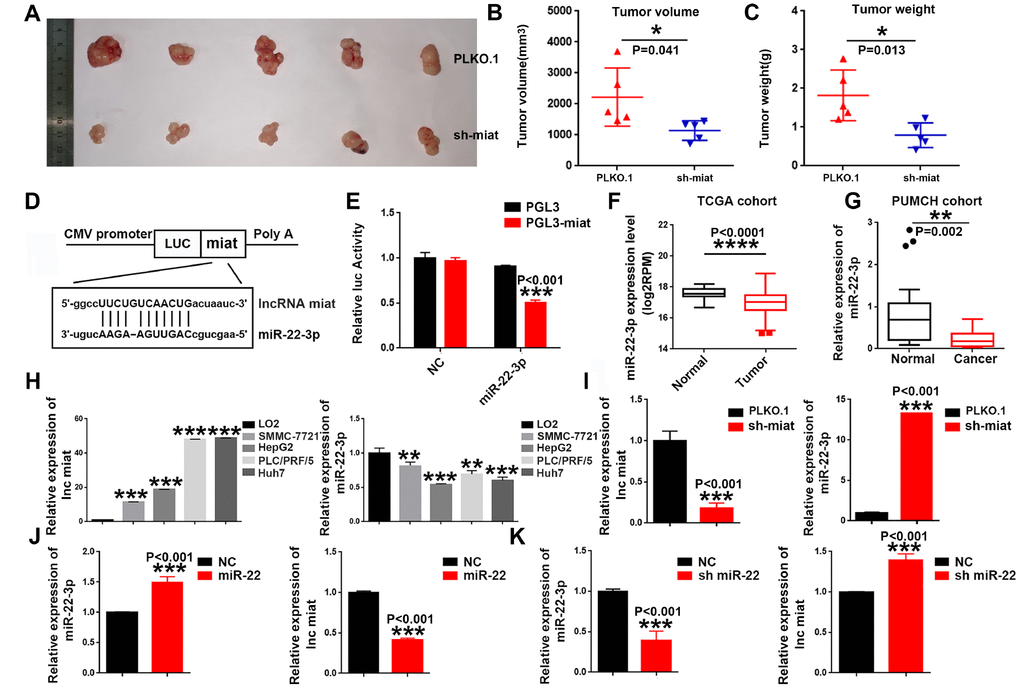
Figure 3.Knockdown of miat suppresses hepatocarcinogenesis in vivo, miR-22-3p bonded to and suppressed miat expression. (A–C) Subcutaneous injection of SMMC-7721 cells with or without miat knockdown into nude mice. Representative images of the resected subcutaneous tumors from each group are shown. Tumor volumes and tumor weights were measured (n = 6). *P < 0.05, **P< 0.01, *** P< 0.001. (D) Bioinformatics prediction using miRcode indicated that the miat sequence contained the putative binding site of miR-22-3p. (E) The cDNA of miat was cloned downstream of the luciferase gene (PGL3-miat) and transfected into HepG2 cells with miR-22-3p or control oligonucleotides. Luciferase activity was detected 48 h after transfection. The bars represent the mean and SD of three independent experiments, *P < 0.05, **P< 0.01, *** P< 0.001. (F) MiR-22-3p expression analyses in HCC and nontumor tissues in TCGA datasets. *P < 0.05, **P< 0.01, *** P< 0.001, **** P< 0.0001. (G) MiR-22-3p levels in 20 HCC and paired nontumor tissues in PUMCH cohort. *P < 0.05, **P< 0.01, *** P< 0.001, **** P< 0.0001. (H) QRT-PCR analysis of miat and miR-22-3p expression in human normal liver cell line (LO2) and HCC cell lines (SMMC-7721, HepG2, PLC/PRF/5 and Huh7). Data are expressed as the mean ± SD; n=3, *P < 0.05, **P< 0.01, *** P< 0.001, **** P< 0.0001 compared with the control group. (I) MiR-22-3p expression was increased in HepG2 cells transfected with sh-miat. The bars represent the mean and SD of three independent experiments, *P < 0.05, **P< 0.01, *** P< 0.001. (J) Miat expression was decreased in HepG2 cells transfected with the miR-22-3p(miR-22). The bars represent the mean and SD of three independent experiments, *P < 0.05, **P< 0.01, *** P< 0.001. (K) Miat expression was increased in HepG2 cells transfected with the miR-22-3p inhibitor(sh-miR-22); The bars represent the mean and SD of three independent experiments, *P < 0.05, **P< 0.01, *** P< 0.001.