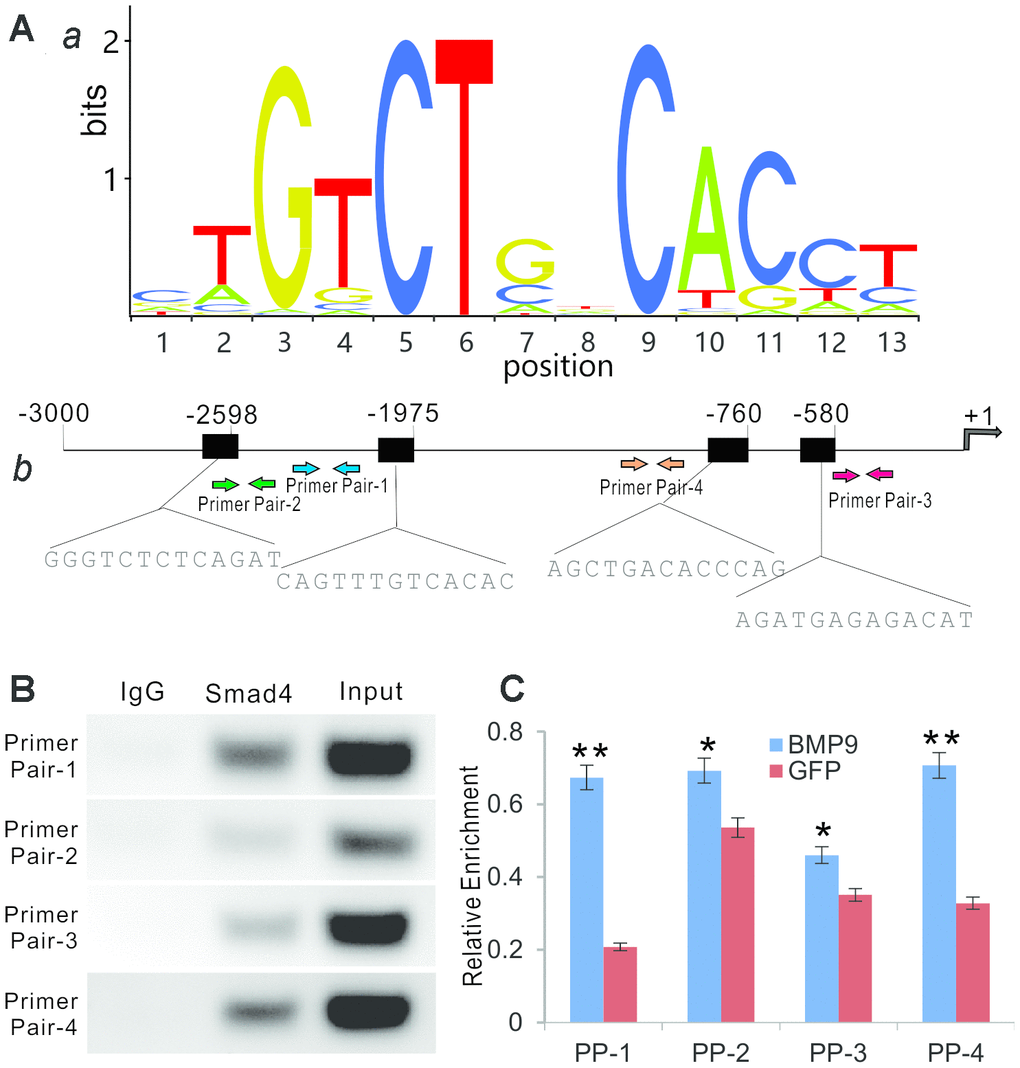
Figure 5.BMP9 regulates Rmst expression through Smad signaling pathway. (A) Bioinformatic prediction of putative Smad4 binding motif sequences using JASPAR. The representative position weight matrix for motif enriched in Smad4 binding sites by Chip-seq database (a). The sequences of putative binding sites and locations of PCR primer pairs are shown in (b). (B) ChIP analysis was performed with specific antibody for Smad4 in iMADs. Isotype matched IgG was used as a negative control. A whole cell extract (Input) was used as a positive control. (C) BMP9-induced binding of Smad4 to Rmst promoter. The iMADs were infected with Ad-BMP9 or Ad-GFP for 48h, and then subjected to anti-Smad4 ChIP pull-down as described in (B). RT-qPCR analysis was carried out to determine relative Smad4 promoter enrichment with different primer pairs. “*”, p<0.05, “**”, p<0.01, Ad-BMP9 group vs. Ad-GFP group.