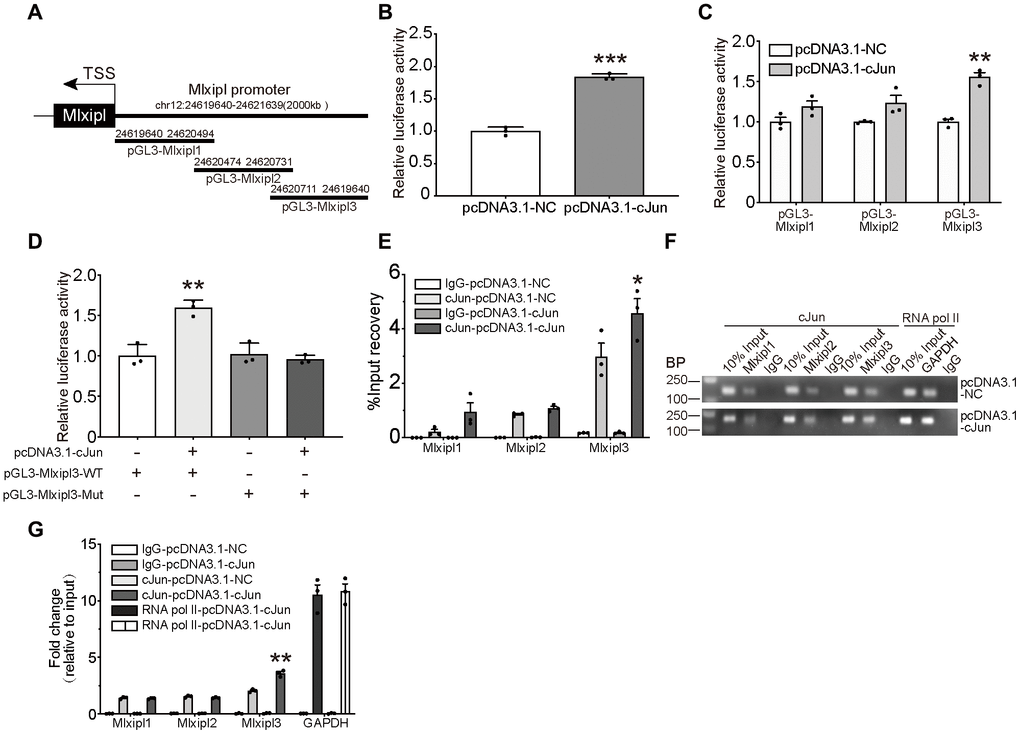
Figure 4.cJun directly regulated Mlxipl expression at the transcriptional level. (A) Schematic diagrams showed the Mlxipl promoter and its fragments containing predicting binding sites. The diagrams show the Mlxipl promoter and its fragments, which were used to predict potential binding sites. (B) Luciferase reporter assays of full-length Mlxipl reporter. PcDNA3.1-cJun or pcDNA3.1-NC was co-transfected with full-length Mlxipl reporter in microglia. PcDNA3.1-cJun promoted luciferase activity of full-length Mlxipl reporter luciferase activity. The data were relative to the pcDNA3.1-NC group. N = 3. ***P < 0.001 vs. pcDNA3.1-NC. (C) Luciferase reporter assays of fragments of Mlxipl reporter. PcDNA3.1-cJun or pcDNA3.1-NC was co-transfected with fragmented Mlxipl promoters (pGL3-Mlxipl1, pGL3-Mlxipl2 and pGL3-Mlxipl3), respectively. PcDNA3.1-cJun promoted luciferase activity of pGL3-Mlxipl3 reporter luciferase activity. N = 3. **P < 0.01 vs. pcDNA3.1-NC. (D) Luciferase reporter assays of mutation of pGL3-Mlxipl3 reporter. PcDNA3.1-cJun or pcDNA3.1-NC was co-transfected with pGL3-Mlxipl3 reporter with or without mutation. N = 3. **P < 0.01 vs. pcDNA3.1-cJun+pGL3-Mlxipl3. (E and F) ChIP assays with anti-cJun antibody were performed followed by qPCR or agarose gel electrophoresis. The DNA product of ChIP was detected by fragmented Mlxipl promoter primers. PcDAN3.1-cJun transfection enriched more Mlxipl3 fragments. IgG was used as a negative control; RNA pol II was used as a positive control system; ChIP DNA mixture was used as Input, and the sample loading was 10% of other groups. Quantification of agarose gel electrophoresis (G). N= 3. *P < 0.05, **P < 0.01 vs. pcDNA3.1-NC. TSS, transcription starting site.