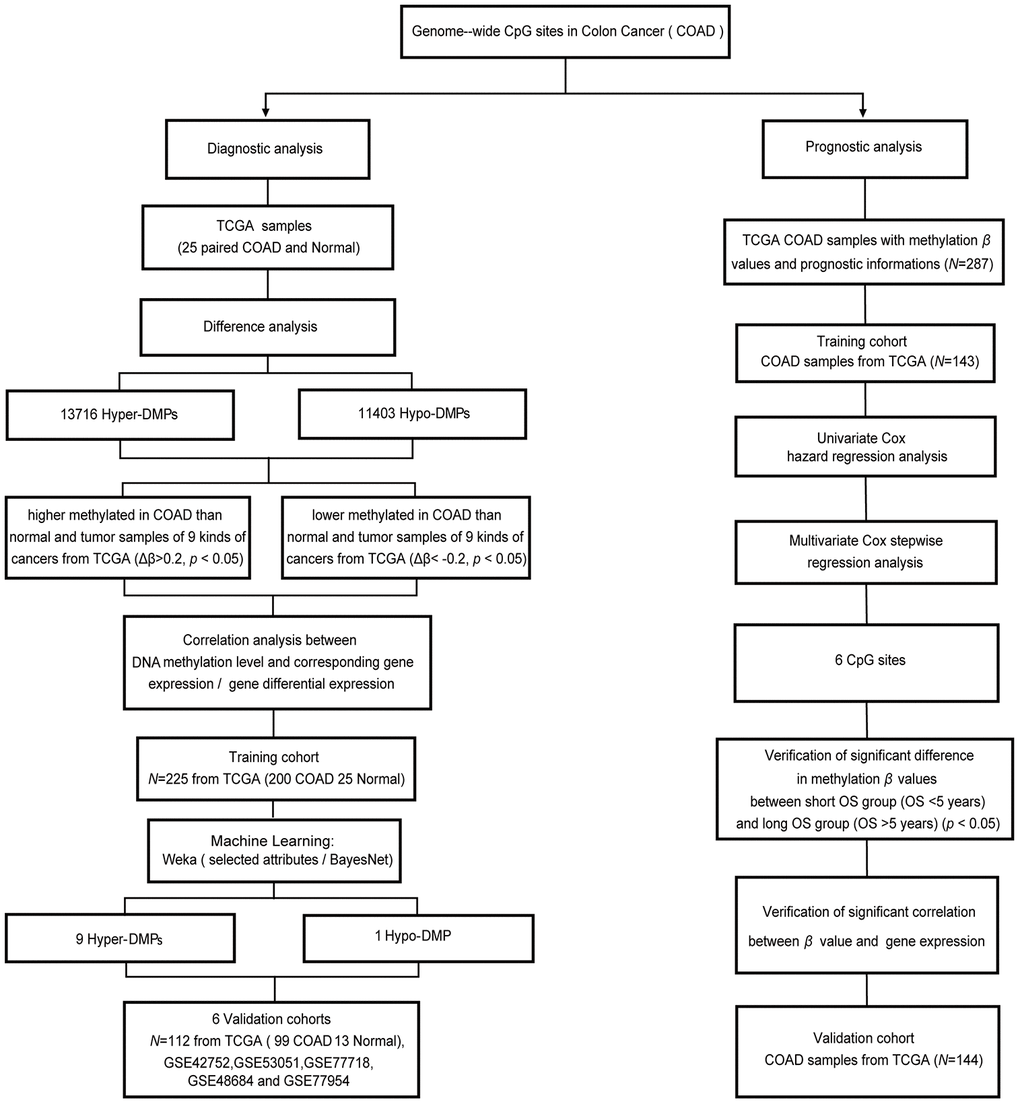
Figure 1.Workflow diagram for biomarker screening and model construction. The DNA methylation levels of genome-wide CpG sites were used to screen biomarkers and construct diagnostic and prognostic models of COAD. Left side: diagnostic biomarker selection and COAD-specific diagnostic model construction. Conditional screening and machine learning using the selected attributes and BayesNet functions of WEKA were performed to obtain the final nine Hyper-DMPs and one Hypo-DMP as potential biomarkers in the training cohort from TCGA (including 200 COAD and 25 normal samples). BayesNet was used to evaluate the COAD-specific diagnostic model based on these DMPs in the validation cohort from TCGA (including 99 COAD and 13 normal samples) and five independent GEO cohorts (GSE42752, GSE53051, GSE77718, GSE48684 and GSE77954). Right side: prognostic biomarker selection and COAD prognostic model construction. Univariate Cox hazard regression analysis and multivariate Cox stepwise regression analysis were applied to 143 TCGA COAD samples as the training cohort to obtain six CpG sites as potential biomarkers. The prognostic model based on these six CpG sites was evaluated using 144 TCGA COAD samples as the validation cohort.