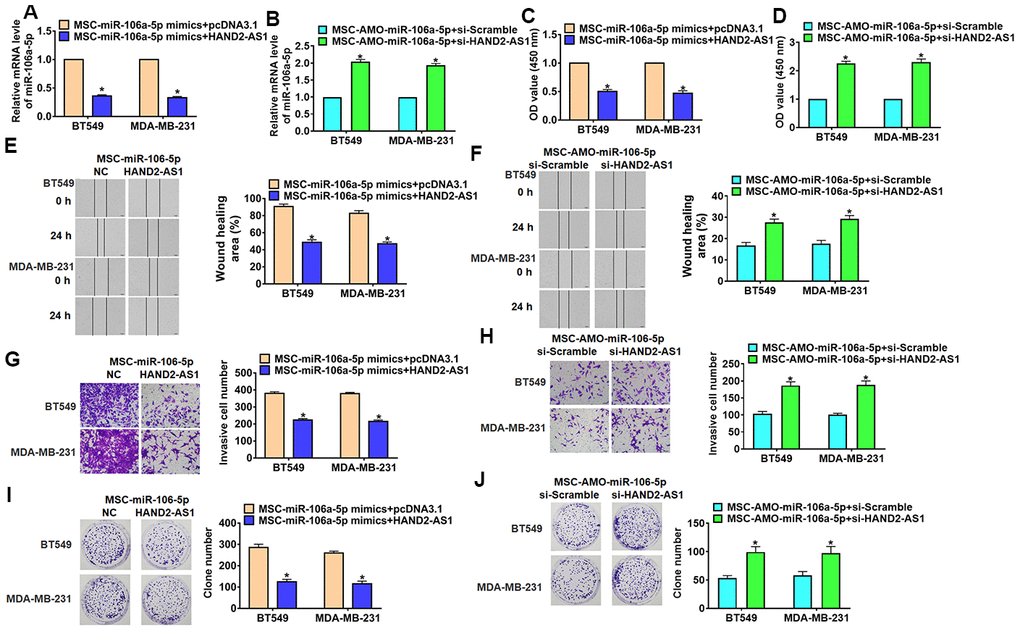
Figure 6.Overexpression lncRNA HAND2-AS1 inhibited progression of TNBC cells by regulating exo-miR-106-5p. BT549 and MDA-MB-231 cells were transfected with HAND2-AS1 or si-HAND2-AS1 and incubated with exosomes from MSCs transfected miR-106a-5p or AMO-miR-106a-5p. (A, B) qRT-PCR analyzed miR-106a-5p expression in BT549 and MDA-MB-231 cells. n = 6, *p<0.05. (C, D) MTT for BT549 and MDA-MB-231 cells. n = 10, *p<0.05. (E, F) Migrative ability was detected by wound healing assay. n = 4, *p<0.05 vs MSC-miR-NC or MSC-AMO-miR-NC. (G, H) Invasive ability was detected by Transwell assay. n = 4, *p<0.05. (I, J) Proliferative ability was detected by clone formation assay. n = 4, *p<0.05. The above measurement data were expressed as mean ± standard deviation. Data among multiple groups were analyzed by one-way ANOVA, followed by a Tukey post hoc test. The experiment was repeated in triplicate.