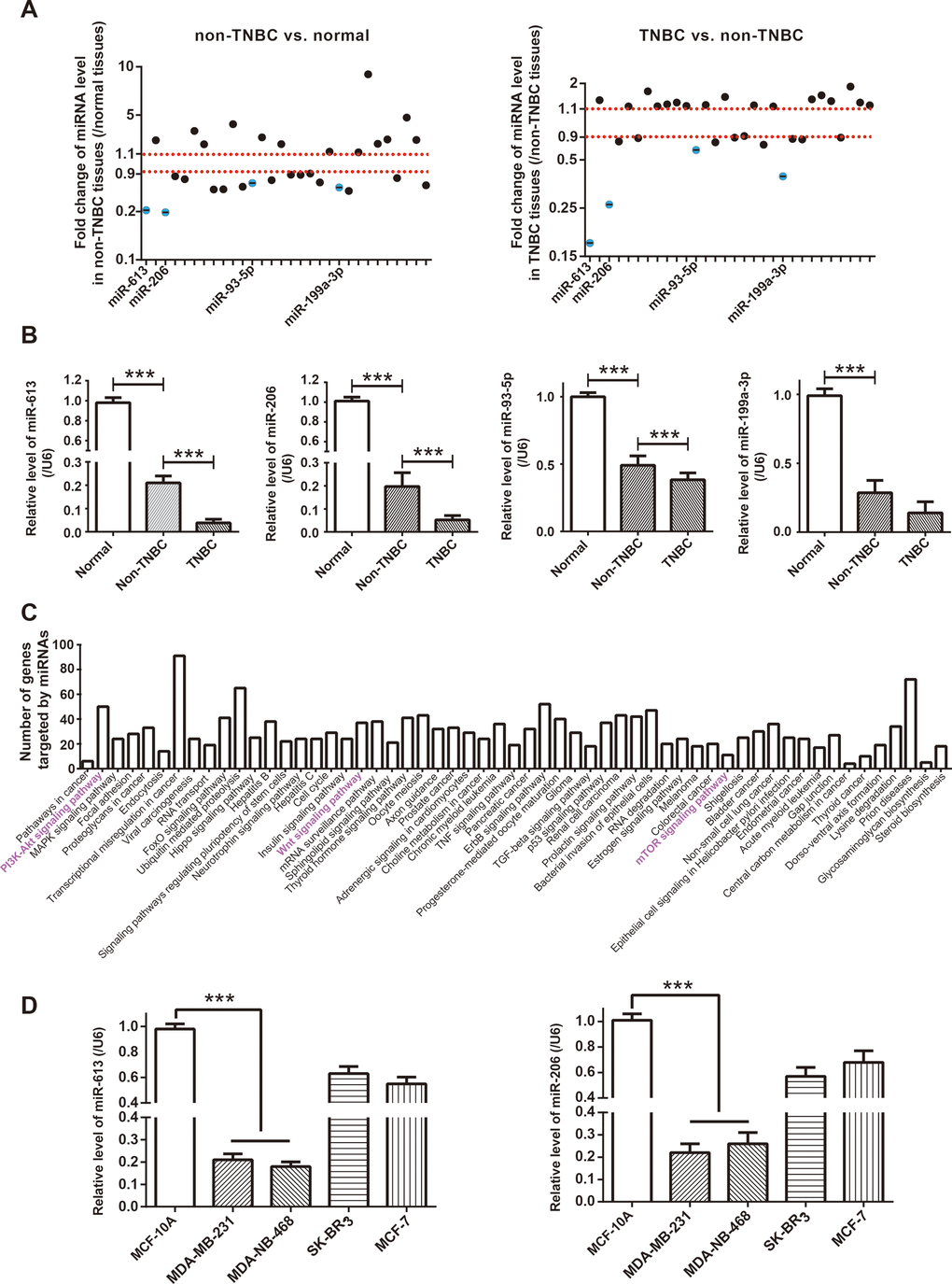
Figure 2.Identification of miRNA network of circRNA AKT3 in triple-negative breast cancer (TNBC). (A) Fold change of miRNAs, potentially sponged by significant AKT3-derived circRNAs, were determined in BC tissues of other subtypes (i.e. non-TNBC) as relative to normal tissues, as well as in TNBC tissues as relative to BC tissues of other subtypes. (B) Expressions of miR-613, miR-206, miR-93-5p and miR-199a-3p were determined in normal tissues, TNBC tissues and BC tissues of other subtypes (i.e. non-TNBC). ***: P<0.001. (C) KEGG pathways enriched by genes targeted by miR-613, miR-206, miR-93-5p and miR-199a-3p were drawn from miRPath online tool (http://snf-515788.vm.okeanos.grnet.gr/). (D) MiR-613 and miR-206 expressions were compared among MCF-10A, MDA-MB-231, MDA-MB-468, SK-BR-3 and MCF-7 cell lines. ***: P<0.001.