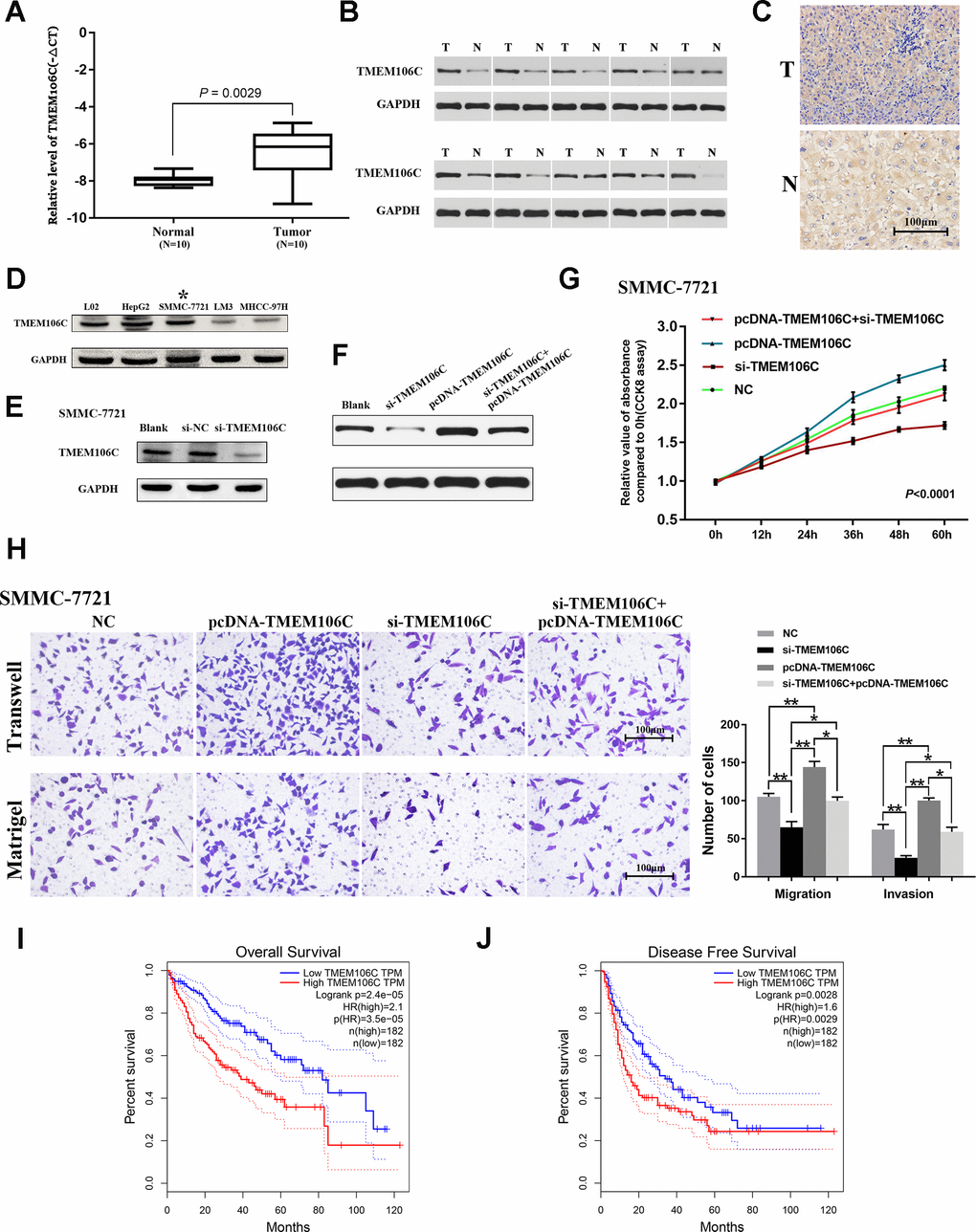
Figure 3.Expression validation, functional exploration and prognostic value of TMEM106C in HCC. (A) Relative expression level of TMEM106C in 10 pairs of HCC samples (tumor tissues and adjacent normal liver tissues), as assessed by real-time PCR. (B) The protein expression level of TMEM106C in 10 pairs of HCC samples, as assessed by western blot. (C) TMEM106C IHC staining in HCC and adjacent normal liver tissue. 400×. (D) The protein expression level of TMEM106C in the normal liver cell line L02 and in different HCC cell lines. The cell line SMMC-7721 was selected for further study. (E) siRNAs targeting TMEM106C were transfected into SMMC-7721 cells for 48 h, and then all cell lysates were harvested for western blotting. (F) TMEM106C levels under si-TMEM106C, pcDNA-TMEM106C plasmid or si-TMEM106C plus pcDNA-TMEM106C plasmid assessed by western blot. (G) si-TMEM106C (50 nM) and pcDNA-TMEM106C plasmid plus si-TMEM106C were transfected into SMMC-7721 cells and were compared to untransfected control cells. Every 12 h, cell numbers were measured by CCK8 assay. NC represents pcDNA3.1, si-NC, and blank control, which were proven to not be different from each other. P < 0.01. 32 (H) Transwell migration and invasion assays of SMMC-7721 cells after transient transfection with si-TMEM106C or pcDNA-TMEM106C plasmid plus si-TMEM106C or not. The migration and invasion cell numbers are shown in histograms (mean ± SD). NC represents pcDNA3.1, si-NC, and blank control, which were proven to not be different from each other. 400×. *P < 0.05, **P < 0.01. (I) The overall survival rates of 364 HCC patients were compared between the TMEM106C high and low expression groups using Kaplan-Meier analysis (GEPIA). (J) The disease-free survival rates of 364 HCC patients were compared between the TMEM106C high and low expression groups using Kaplan-Meier analysis (GEPIA).