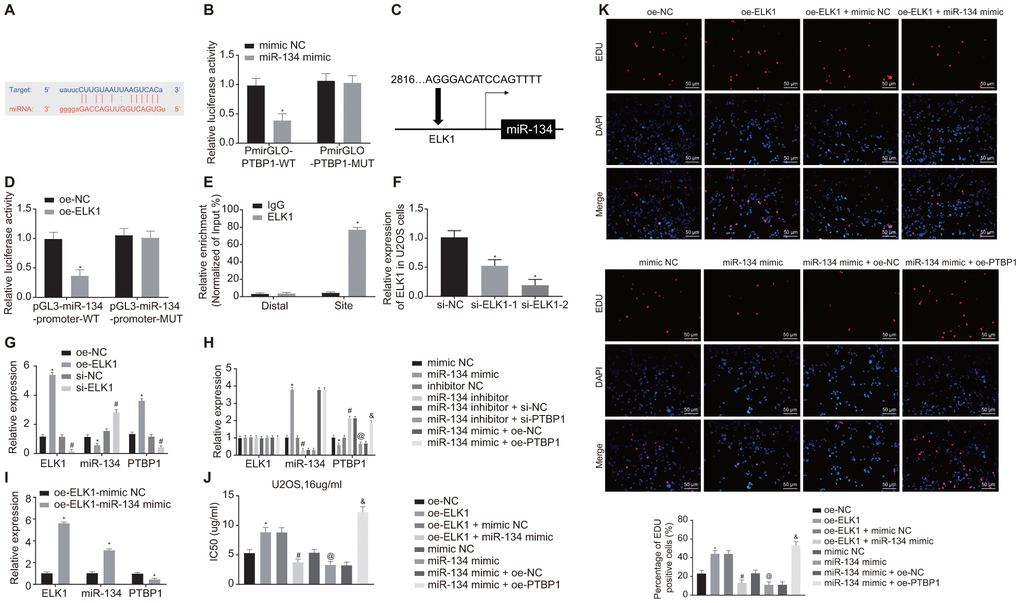
Figure 6.ELK1 promotes chemoresistance of osteosarcoma cells to DXR by inhibiting miR-134 and elevating PTBP1 expression. (A) Prediction of binding site between miR-134 and PTBP1 using online analysis website microRNA (MicroRNA.org). (B) Detection of luciferase activity using dual-luciferase reporter gene assay. (C) Prediction of binding site between ELK1 and miR-134 using online analysis website PROMO8.3 (http://alggen.lsi.upc.es/). (D) Detection of luciferase activity using dual-luciferase reporter gene assay. (E) Intersection role between ELK1 and miR-134 promoter detected by ChIP assay. U2OS cells were treated with expression vectors containing ELK1 gene or si-targeting ELK1 gene. (F) The knockdown efficiency of si-ELK1 in U2OS cells verified using RT-qPCR. (G) Expression of ELK1, miR-134, and PTBP1 (normalized to GAPDH and U6) in U2OS cells determined using RT-qPCR. PTBP1-overexpressed or -depleted U2OS cells were treated with exogenous miR-134 mimic or miR-134 inhibitor. (H) Expression of ELK1, miR-134, and PTBP1 (normalized to GAPDH and U6) in U2OS cells determined using RT-qPCR. (I) Expression of ELK1, miR-134, and PTBP1 (normalized to GAPDH and U6) in U2OS cells determined using RT-qPCR. U2OS cells were treated with expression vector containing ELK1 gene, miR-134 mimic, or PTBP1 alone or in combination after DXR treatment. (J) Cell viability and IC50 in U2OS cells detected using CCK8. (K) Cell proliferation in U2OS cells detected using EdU assay. Values obtained from three independent experiments in triplicate are analyzed by unpaired t test between two groups, by ANOVA followed by Tukey's test among three or more groups, and by repeated measurement ANOVA followed by Bonferroni test at indicated time points. The values at different time points were analyzed by repeated measurement ANOVA, followed by Bonferroni test. In panel C, D, *p < 0.05 U2OS cells treated with oe-NC. In panel F, *p < 0.05 U2OS cells treated with si-NC. In panel G, *p < 0.05 U2OS cells treated with oe-NC; # p < 0.05 U2OS cells treated with si-NC. In panel H, *p < 0.05 U2OS cells treated with mimic NC; # p < 0.05 U2OS cells treated with inhibitor NC; @ p < 0.05 U2OS cells treated with miR-134 inhibitor + si-NC; & p < 0.05 U2OS cells treated with miR-134 mimic + oe-PTBP1. In panel I, *p < 0.05 U2OS cells treated with oe-ELK1.