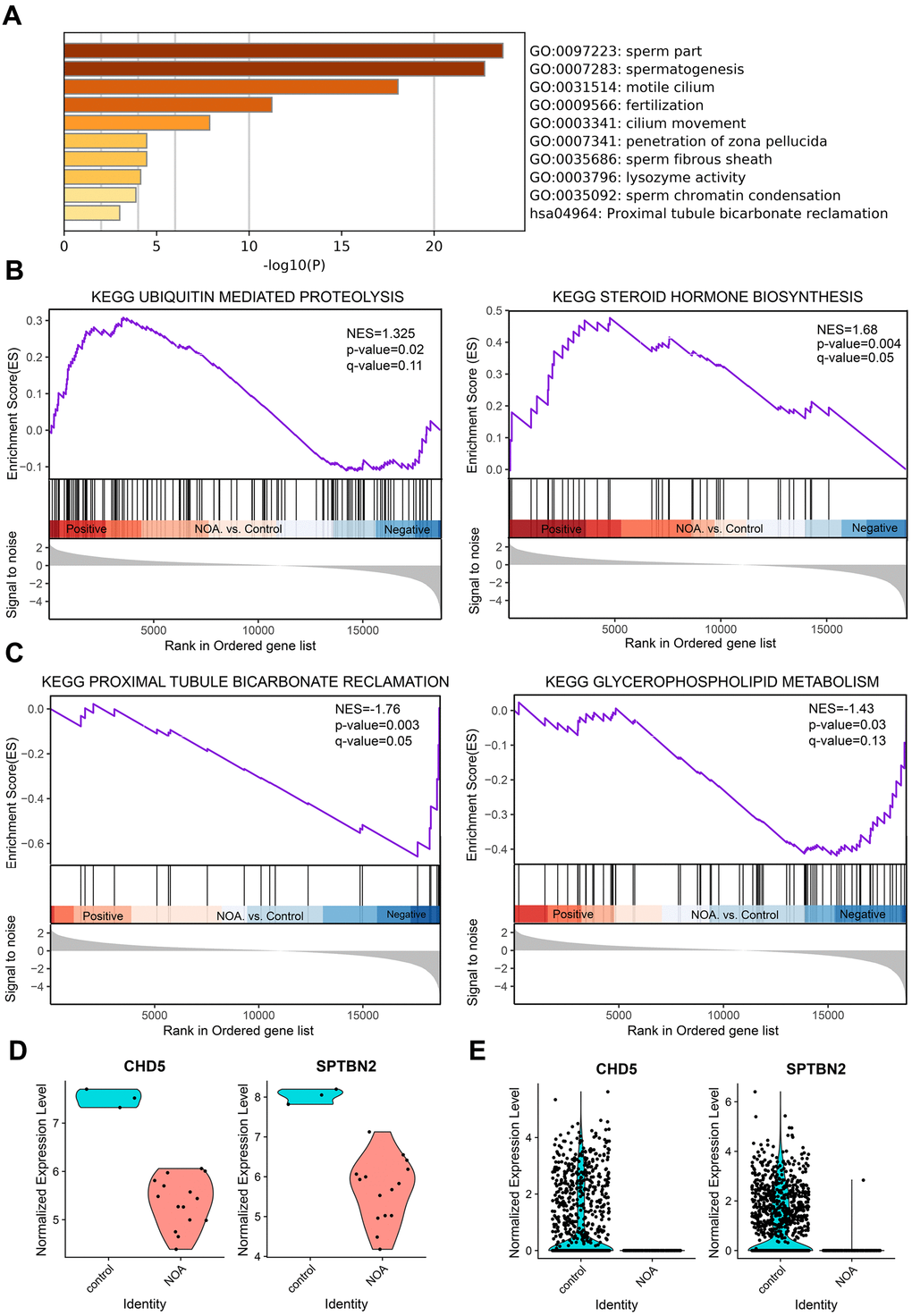
Figure 1.GO enrichment and Gene Set Enrichment Analysis of NOA, and validation of hub genes. (A) Top 10 clusters with their representative enriched term based on GO enrichment analysis of DEGs. (B) NOA samples were correlated positively with gene signatures related to Steroid Hormone Biosynthesis and Ubiquitin Mediated Proteolysis pathways. (C) NOA samples were correlated negatively with gene signatures related to Glycerophospholipid Metabolism and Proximal Tubule Bicarbonate Reclamation pathway. (D) The normalized expression validation of hub genes using the GSE45887 dataset. The Y-axis expressions were normalized by log2(TPM/10+1). (E) The normalized expression validation of hub genes using the GSE106487 dataset. The Y-axis expressions were normalized by log2(TPM/10+1).