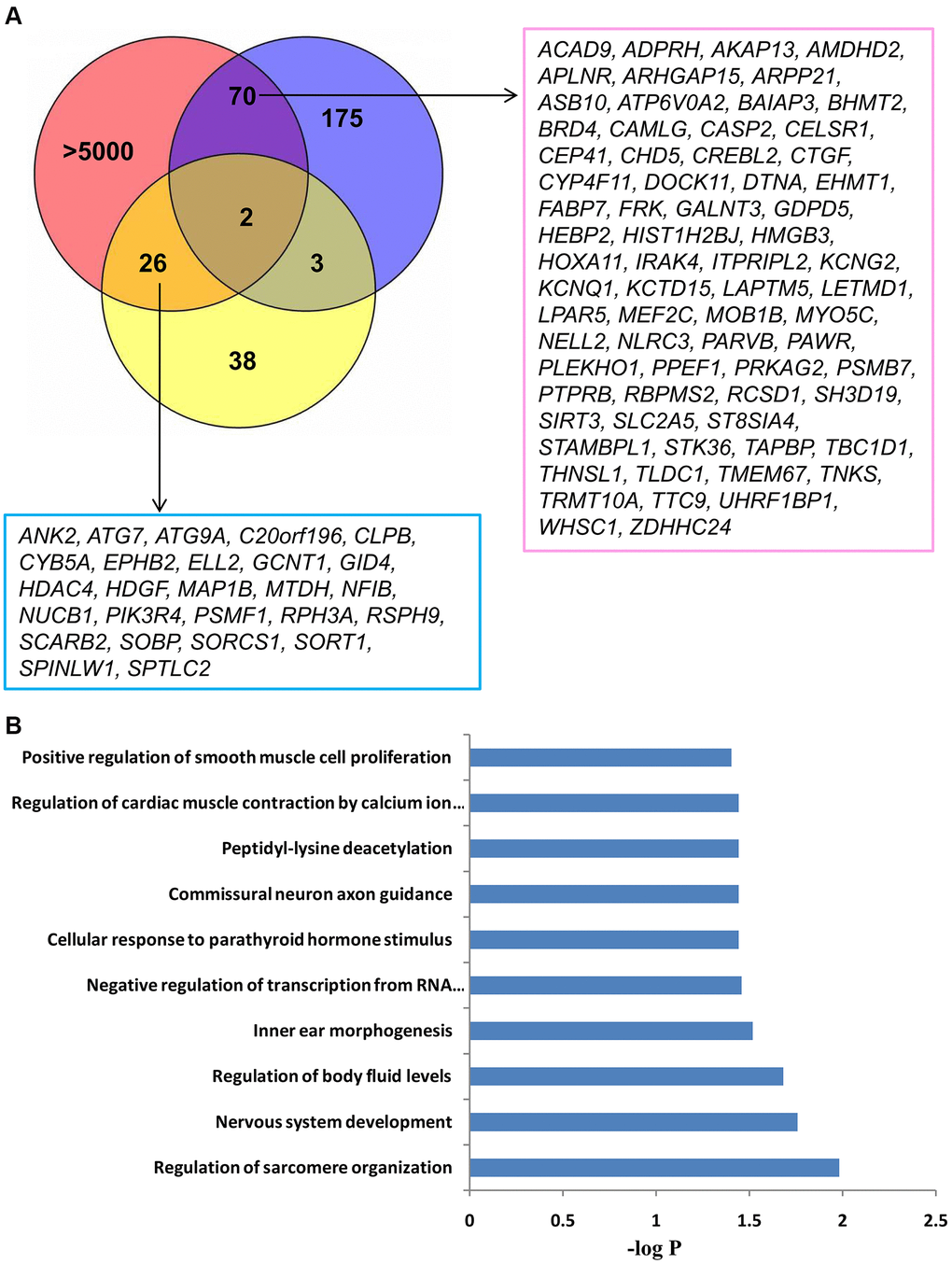
Figure 3.The identification and GO analysis of up-regulated targets of miRNAs associated with LA. (A) An integrated analysis of microarray LA miRNA and mRNA expression profiling identified 98 up-regulated genes, which could be inhibited by hsa-miR-1972, hsa-miR-26b-5p, or hsa-miR-3141. Two targets showed increased expression in both WML tissue and whole blood of LA subjects. Blue and yellow circles show 250 up-regulated genes in WML tissue and 79 up-regulated genes in whole blood, respectively. The pink circle includes more than 5000 potential targets of miR-1972, miR-3141, and miR-26b-5p, which were predicted from multiple predictive tools. (B) Top 10 biological processes significantly enriched with 98 overlapping genes. Functional enrichment analysis was performed using a DAVID bioinformatics database.