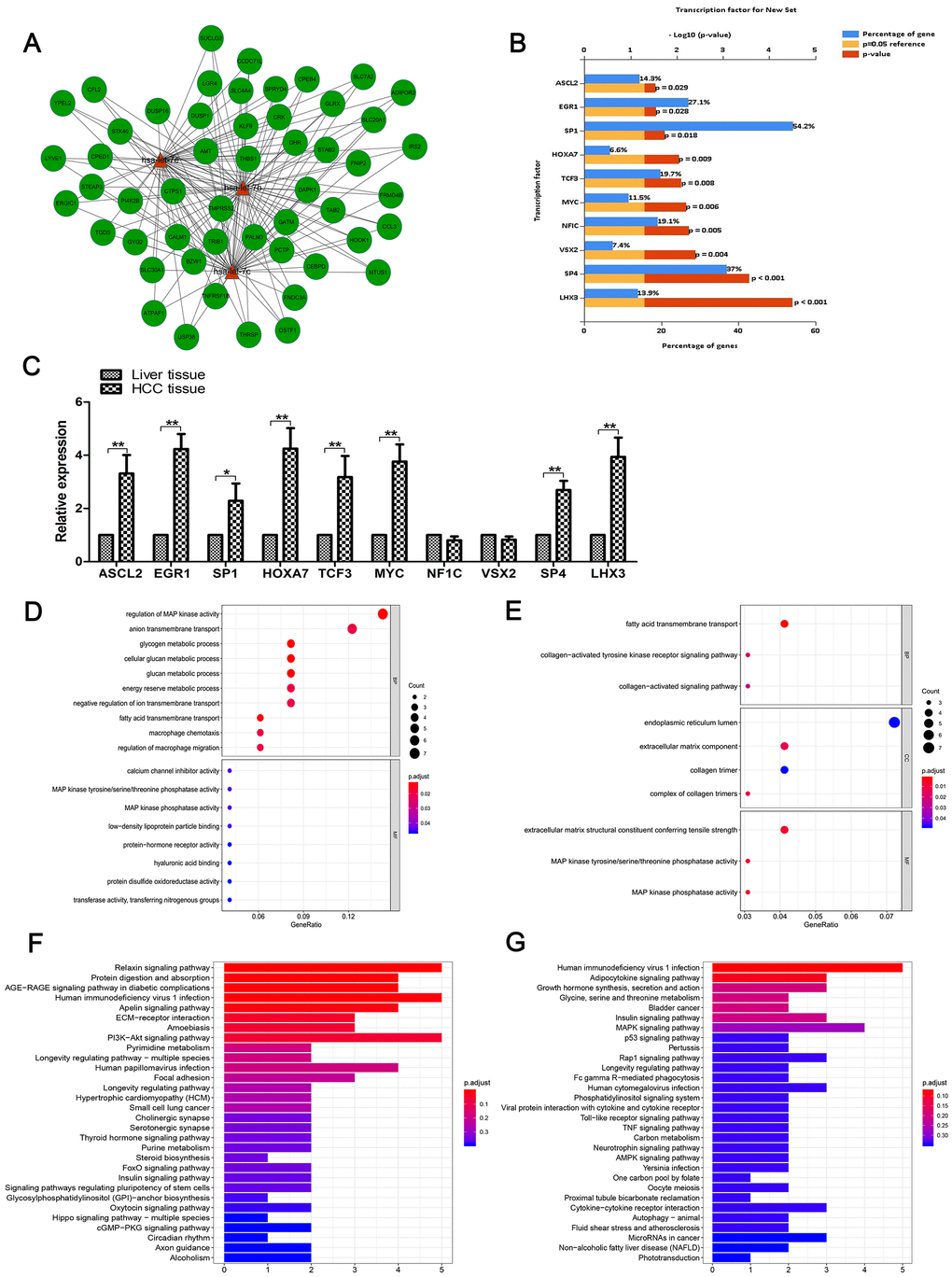
Figure 2.Functional enrichment analysis. (A) The network of cross-linked genes of let-7b, let-7c and let-7e was constructed of these three miRNAs and 52 genes. (B) Top 10 relevant enriched TFs of let-7b, let-7c and let-7e were analyzed, including ASCL2, EGR1, SP1, HOXA7, TCF3, MYC, NF1C, VSX2, SP4 and LHX3. (C) Expression of the aforementioned 10 TFs in HCC tissues and matched normal tissues as shown by RT-qPCR. (D) GO analysis of genes negatively correlated with the three miRNAs. (E) GO analysis of genes positively correlated with these miRNAs. (F) KEGG pathway enrichment analysis was conducted to identify the potential pathways of genes positively correlated with the miRNAs. (G) KEGG pathway enrichment analysis of genes negatively correlated with the miRNAs.