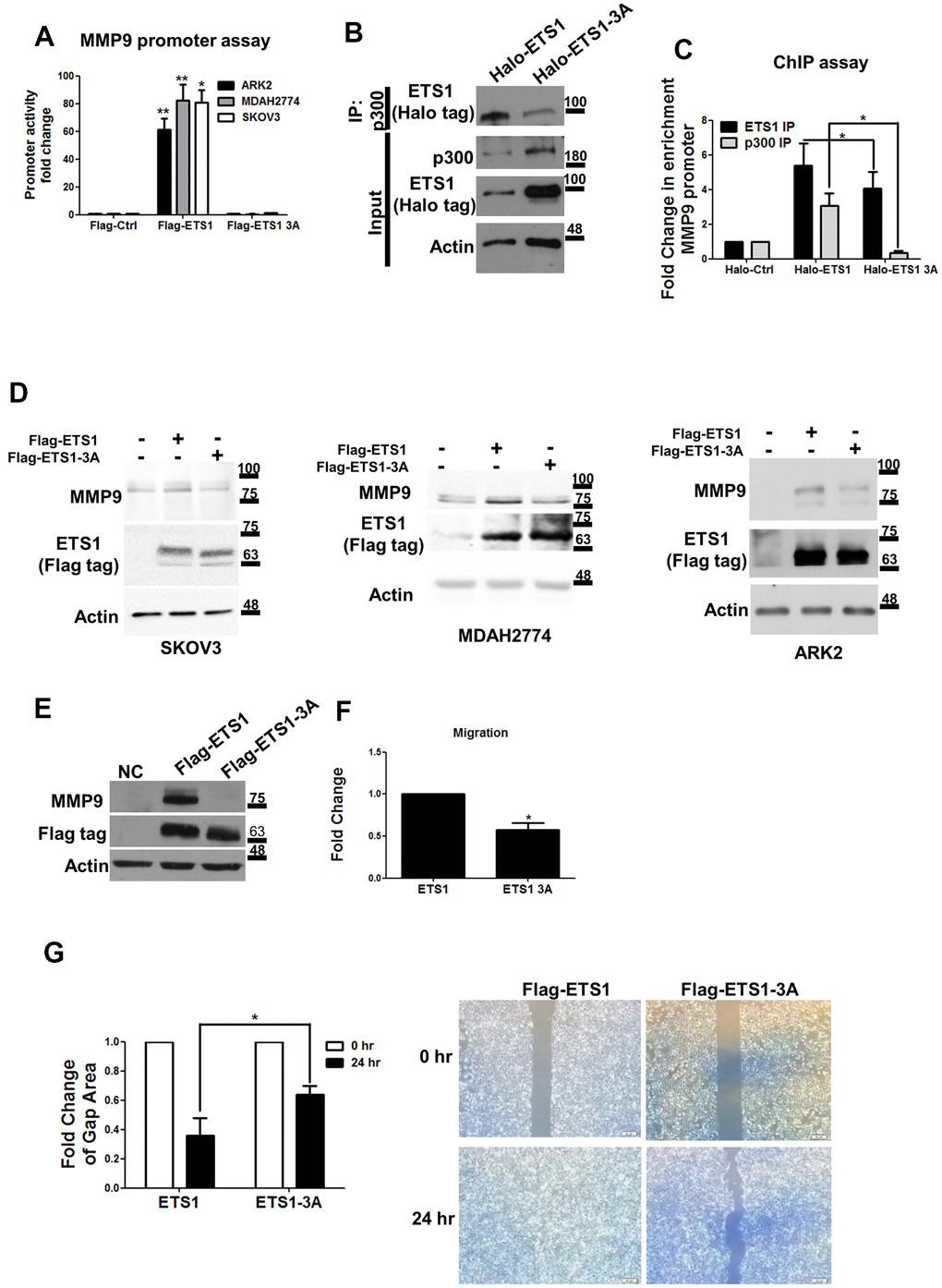
Figure 5.Mutant ETS1 is unable to activate MMP-9 expression. (A) MMP-9 promoter luciferase assays were co-transfected with the MMP-9 promoter construct using Flag-control, Flag-ETS1 or Flag-ETS1-3A in ARK2 (black box), MDAH2774 (gray box), and SKOV3 (white box) cells 48 h later. The MMP-9 promoter activity in transfected Flag-control cells was set as 1. Results are means ± standard errors of the mean from three independent experiments. (B) An anti-p300 antibody was able to precipitate Halo-ETS1 or Halo-ETS1-3A in overexpressing 293 cells. Halo-ETS1 or Halo-ETS1-3A in association with p300 was identified with an anti-Halo tag antibody. (C) ChIP assays were analyzed with Halo-ETS1, Halo-ETS1-3A, and endogenous p300 on the MMP-9 promoter in overexpressing 293 cells. Chromatin DNA was crosslinked with Halo-ETS1 or Halo-ETS1-3A and subsequently purified with a Halo-tag resin. Associated p300 was precipitated with an anti-p300 antibody. Enrichment of the MMP-9 promoter was identified by qPCR. Results are means ± standard errors of the mean from three independent experiments. (D) MMP-9 protein levels were increased by Halo-ETS1 or Halo-ETS1-3A in SKOV3, MDAH2774 and ARK2 cells. (E) MMP-9 expression was detected in 293 cells stably transfected Flag-ETS1. (F) The results of the transwell migration assay revealed that the migration ability of cells expressing wild-type ETS1 was higher than that of cells expressing ETS1-3A. Quantification of cell migration ability was performed by direct counting. (G) In wound closure assay, the cell-free gap area was quantified with the image J software (left panel). Cells characterized by higher migration ability produced smaller cell-free gap areas. Cell-free gap areas at 0 and 24 h are shown as open and black bars, respectively. Results in (F, G) are means ± standard errors of the mean from three independent experiments. Asterisks denote statistical significance (*p < 0.05, **p < 0.01).