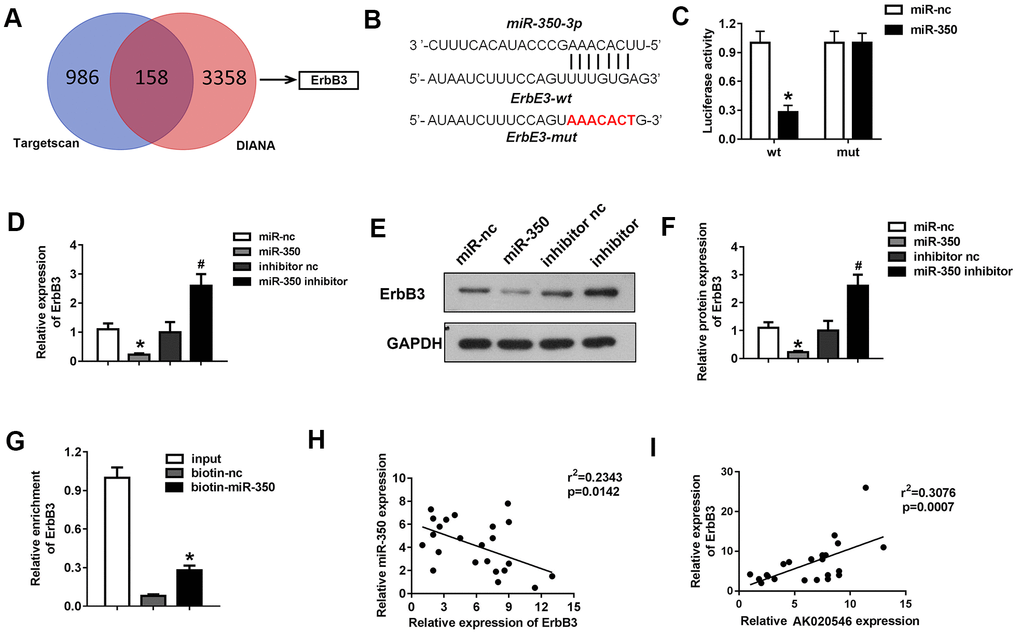
Figure 7.MiR-350-3p directly targeted ErbB3 in H9c2 cardiomyocytes. (A) Bioinformatics analysis using TargetScan and DIANA predicted ErbB3 as the target of miR-350-3p. (B) The potential targeting region between miR-350-3p and ErbB3 predicted by bioinformatics analysis. (C) Luciferase assay was performed to verify whether miR-350-3p targeted ErbB3 in H9c2 cardiomyocytes (n = 5). (D) qPCR was used to detect the expression of ErbB3. (E, F) Western blot was carried out to evaluate the expression level of ErbB3 (n = 5). (G) Luciferase assay was performed to verify whether miR-350-3p targeted ErbB3 in H9c2 cardiomyocytes (n = 5). (H, I) Pearson’s analysis was performed to investigate the correlation between AK020546 and ErbB3 as well as miR-350-3p and ErbB3 (n = 5). *p < 0.05 vs miR-nc or biotin-nc, #p < 0.05 vs inhibitor nc group.