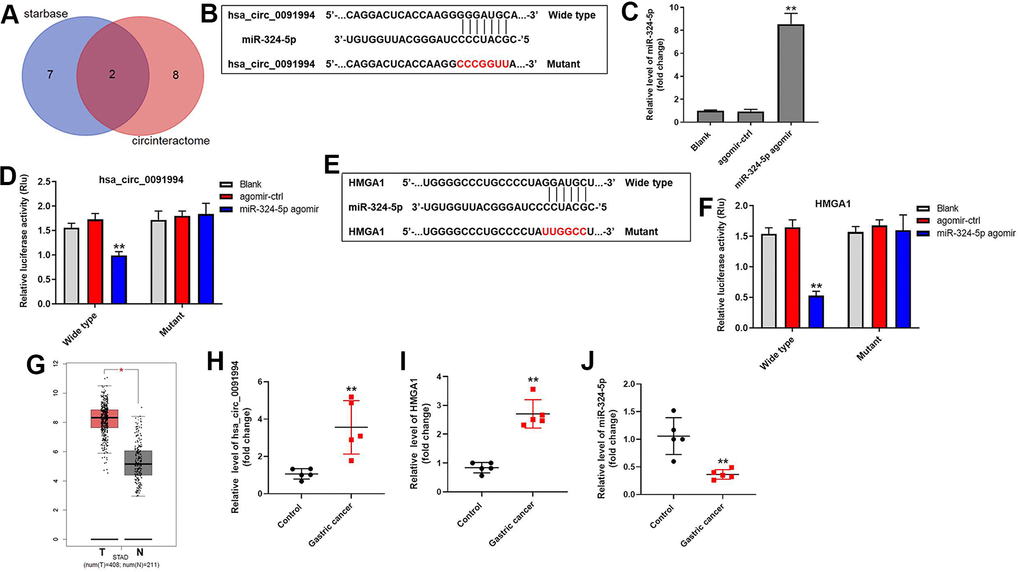
Figure 4.Downstream target of hsa_circ_0091994. (A) The downstream targets of hsa_circ_0091994 were predicted using Starbase and CircInteractome. (B) MiR-324-5p was predicted as the downstream target of hsa_circ_0091994. Sequences of wild type and mutant 3’ UTR of miR-324-5p. (C) MiR-324-5p agomir or the negative control (agomir-ctrl) was transfected into AGS cells for 48 hr. The efficacy of miR-324-5p agomir was detected by RT-qPCR. (D) Dual luciferase assay for hsa_circ_0091994 was conducted at 48 hr after transfection. (E) HMGA1 was predicted as the downstream target of miR-324-5p. Sequences of wild type and mutant 3’ UTR of HMGA1 were presented. (F) Dual luciferase assay for HMGA1 was conducted at 48 hr after transfection. (G) Bioinformatics analysis using GEPIA tool was performed to analyze the levels of HMGA1 in GC tissue and normal gastric tissue. (H–J). The expressions of hsa_circ_0091994, miR-324-5p and HMGA1 in 5 pairs of clinical GC and adjacent normal tissues were detected with RT-qPCR. **P<0.01, compared with blank; n = 3.