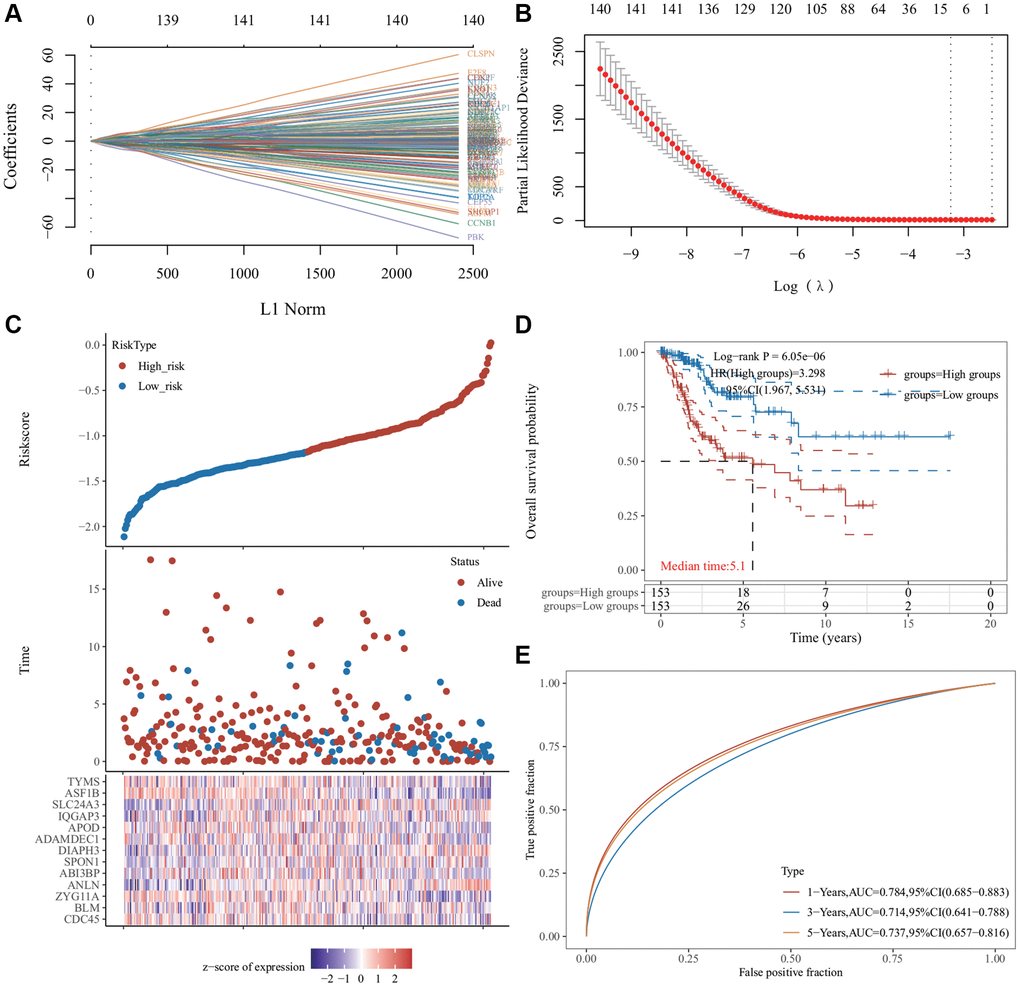
Figure 4.Least absolute shrinkage and selection operator (LASSO) regression analysis of common differentially expressed genes. (A) The LASSO coefficient curve. The abscissa represents the value of the independent variable lambda; the ordinate represents the coefficient of the independent variable. (B) It plots the relationship between partial likelihood deviation and log (λ) using the LASSO-Cox regression model. (C) The risk score, survival time, and survival status of the dataset selected, wherein the top represents the risk score scatter map from low to high, and different colors represent different risk groups; the middle represents the scatter plot distribution of survival time and survival state corresponding to different sample risk score; the bottom diagram represents the expression heat map of genes contained in the signature. (D) The Kaplan−Meier (KM) survival curve distribution of the risk model in the data set, in which different groups were tested using log-rank. High-risk (HR) represents the risk coefficient of the high-risk group relative to the low-risk group. HR >1 represents the risk model; HR <1 represents the protection model; 95% CI represents the HR confidence interval. The mean of time represents the time in which the survival rates of the HR and the low-risk groups were 50% (i.e., the median survival time) in unit years. (E) The receiver operator characteristic (ROC) curve and area under the ROC curve (AUC) of the risk model at different times, in which the higher the AUC value, the stronger the prediction ability of the model. (Riskscore = (−0.0685) × CDC45 + (−0.451) × BLM + (−0.2443) × ZYG11A + (0.465) × ANLN + (−0.1119) × ABI3BP + (0.1099) × SPON1 + (0.3614) × DIAPH3 + (−0.1009) × ADAMDEC1 + (−0.0683) × APOD + (0.3228) × IQGAP3 + (0.1394) × SLC24A3 + (−0.5415) × ASF1B + (−0.0287) × TYMS. The cutoff value of riskcore = −1.137103259).