Introduction
Single gene mutations that extend life span have provided important information about pathways that regulate aging and longevity [1]. A significant number of these mutations are in genes and pathways that are conserved between invertebrates and vertebrates, with the caveat that the pathways are often more complex in vertebrates. Many pathways implicated in longevity extension are related in some way to nutrient sensing or nutrient utilization, such as insulin signaling, energy production, and protein biosynthesis [1]. For example, the Tor pathway integrates signals from nutrient uptake, stress, growth factors, and cellular energy status [2]. Genetic manipulation of the Tor gene has been shown to affect longevity in yeast, worms, fruit flies, and mice [3–6].
Another category of genes implicated in longevity modulation encode regulatory proteins, such as transcription factors and histone deacetylases (HDACs). HDACs remove acetyl groups from the lysine residues within histones and a range of other proteins [7]. Nuclear HDACs regulate transcription by modifying chromatin structure and transcription factors. There are Zn-dependent HDACs (classes I, II, and IV) and NAD-dependent HDACs (class III, the sirtuins) [7]. Genetic studies have implicated two nuclear HDACs in longevity extension of Drosophila, dRPD3 (class I) [8] and dSIR2 (class III) [9]. While the complete removal of dRPD3 is lethal [10], flies heterozygous for rpd3 mutations have extended longevity [8]. Removal of dSIR2 is not lethal and has no longevity phenotype by itself, but overexpression of dSIR2 extends life span [9]. As with dRPD3, dosage appears to be crucial in that moderate global overexpression mediates life span increases but more pronounced overexpression is deleterious [11, 12]. Recently it was shown that knockdown of two class I HDACs in C. elegans increases life span, but worms null for these gene products have reduced life spans [13]. Several groups have shown that inhibitors of Zn-dependent HDACs such as suberoyanilide hydroxamic acid (SAHA) extend fly longevity [14–16]. Another inhibitor of Zn-dependent HDACs, beta-hydroxybutyrate, has been shown to extend C. elegans longevity [13]. In addition, removal of RPD3 in S. cerevisae leads to longevity extension [17, 18].
The capacity of single gene mutations to extend longevity was preceded by the discovery that nutrient uptake can affect both longevity and aging. Dietary restriction (DR) was first discovered in rodents, but has been shown to affect longevity in a variety of vertebrate and invertebrate species [19–21]. DR is defined as a level of nutrient intake that is reduced relative to ad libitum feeding without malnutrition. While this often means a restricted feeding schedule for rodents, for invertebrates other means need to be employed to restrict food uptake. DR in Drosophila can be achieved by formulating food of different nutrient levels using one of three general options: variation of carbohydrate, protein, or total nutrient (protein and carbohydrate) content [22–24]. Standard fly diets are usually optimized for reproduction (egg laying and larval growth), whereas peak DR-mediated longevity extension involves less nutrients than the standard diet. For the purposes of DR studies the standard diet can simply be regarded as a starting point for the manipulation of nutrient levels.
Our previous study on rpd3-mediated longevity changes in Drosophila melanogaster [8] used two nutrient conditions, standard (corn) and reduced (0.5N). However, a full range of nutrient levels is necessary to detect potential changes in life span relative to standard nutrients [23, 24, 26]. A report that Drosophila rpd3 mRNA level is affected by nutrients [27] lends further impetus to a more thorough examination of the interplay between dRPD3, nutrition, and longevity. In the studies presented here, we determined if there is rpd3-mediated longevity extension at different nutrient levels.
Results
Nutrients and RPD3 have additive effects upon longevity
This study utilizes the same alleles as our previous report [8], rpd3def (null), rpd3P-UTR (hypomorphic), and rpd3P-1.8 (control), and varies complete nutrients (both sucrose and yeast) from 0.3N to 1.5N relative to standard 1.0N conditions. The rpd3def and rpd3P-UTR alleles are lethal when homozygous [10]. In all survival experiments using the rpd3def allele, including the study of flies that are double mutants for rpd3 and Thor, crosses prior to the life span measurements yielded flies with matched genetic backgrounds (a hybrid mix of the rpd3 stock and the yw strain, the latter being wild type for all genes related to this study). rpd3P-UTR has greatly reduced transcription in all tissues, whereas rpd3P-1.8 only has reduced transcription in the eye [10, 28]. All survival experiments using the rpd3P-UTR and rpd3P-1.8 alleles also had matched genetic backgrounds, derived from crosses to CS flies. Heterozygous male flies (20 days old) carrying the rpd3def and rpd3P-UTR alleles have 40% and 25% reduced levels, respectively, of rpd3 mRNA (Supplementary Figure 1).
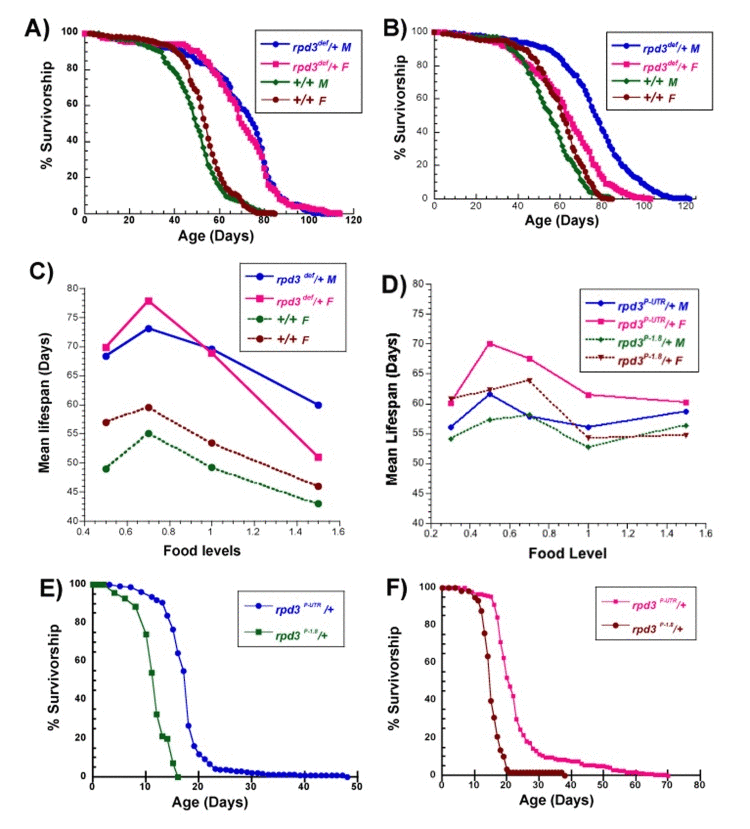
Figure 1. Longevity of rpd3 heterozygous mutant flies (A) Survivorship curves of male and female rpd3def/+ and +/+ control flies on 1.0N (A) and corn food (B). Male rpd3def/+ flies have increased mean and maximal longevity on both food levels compared to controls. Female rpd3def/+ flies have maximal longevity increased on both food levels but mean is increased on 1.0N. (C) Mean longevity of rpd3def/+ and +/+ control flies, male and female, on food levels ranging from 0.5N to 1.5N. (D) Mean longevity of rpd3P-UTR/+ and rpd3P-1.8/+ control flies, male and female, on food levels ranging from 0.3N to 1.5N. (E, F) Survivorship of male (E) and female (F) rpd3P-UTR/+ and rpd3P-1.8/+ controls, on 0.1N food. See Tables 1–4 for number of flies in each experiment and statistical analysis.
Long-lived rpd3 mutant flies exhibited nutrient-mediated changes in longevity. While a diet containing corn meal and yeast is often considered standard (hereafter called corn), the standard food used in this study contained equivalent calories but was formulated using sucrose and yeast, called 1.0N [26, 29]. Males heterozygous for rpd3def have extensions of 37% and 42% in mean longevity on corn and 1.0N food, respectively, compared to genetic controls (Figure 1A, B). Female flies heterozygous for the same allele have 4% and 31% longevity extension on corn and 1.0N food (Figure 1A, B, Table 1, 2). Mean life spans at different nutrient levels are shown for heterozygotes carrying the rpd3def allele in Figure 1C and heterozygotes carrying the rpd3P-UTR and rpd3P-1.8 alleles in Figure 1D (Table 1, 2). Male and female flies carrying the rpd3def allele have extended longevity at all nutrient levels compared to genetic controls, while also showing increases and decreases in mean life span mediated by nutrients. Life span extensions for males vary between 36-41% compared to controls at the same food levels, while for females they vary between 12% at 1.5 food and 33% at 0.7N food. Male and female flies carrying the second allele, rpd3P-UTR, have smaller but consistent extensions in life span compared to flies carrying the control rpd3P-1.8 allele (at most food levels). Females have 10-13% extensions at 0.5N, 1.0N, and 1.5N food levels, whereas males have extensions of 6-7% at 0.3N, 0.5N and 1.0N food levels.
Table 1. Effect of rpd3def on longevity of flies on different food levels
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | +/+ | 1.0 | 261 | 49.7 | 71.3 (32) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 1.0 | 236 | 70.6 (42) | 250.01 | <0.001* | 93.5 (31) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.5 | 261 | 50.4 | 64.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 0.5 | 247 | 68.8 (37) | 283.64 | <0.001* | 95.8 (48%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.7 | 263 | 55.4 | 73.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 0.7 | 243 | 75.2 (36) | 254.15 | <0.001* | 103.6 (42) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 1.5 | 244 | 44.4 | 65 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 1.5 | 254 | 61.2 (39) | 205.28 | <0.001* | 84 (30) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | Corn | 263 | 55.7 | 76 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | Corn | 240 | 76.5 (37) | 243.83 | <0.001* | 105.3 (39) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.0 | 272 | 54.1 | 72.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 1.0 | 269 | 70.8 (31) | 241.92 | <0.001* | 95.5 (32) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.5 | 264 | 57.6 | 72.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 0.5 | 259 | 70.4 (22) | 230.55 | <0.001* | 84.1 (15) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.7 | 275 | 59.8 | 76.7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 0.7 | 251 | 79.6 (33) | 317.80 | <0.001* | 102.8 (34) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.5 | 275 | 46.1 | 63 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 1.5 | 278 | 51.5 (12) | 37.68 | <0.001* | 71 (12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | Corn | 257 | 60.2 | 78 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | Corn | 241 | 62.8 (4) | 24.68 | <0.001* | 88.8 (14) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) rpd3def/+ and +/+ control flies on different food levels. All values obtained for controls are compared to values of the rpd3def/+ group of the same sex and on the same food levels to determine the percent increase in mean and maximal lifespan. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table 2. Effect of DR on longevity of rpd3def/+ (Top) and +/+ control (Bottom) flies
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | rpd3def/+ | 1.0 | 236 | 70.6 | 93.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 0.5 | 247 | 68.8 (−3) | 3.6229 | 0.57 | 95.8 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 0.7 | 243 | 75.2 (7) | 16.91 | <0.001* | 103.6 (11) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 1.5 | 254 | 61.2 (−13) | 58.619 | <0.001* | 84 (−10) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | Corn | 240 | 76.5 (8) | 21.005 | <0.001* | 105.3 (13) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 1.0 | 269 | 70.8 | 95.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 0.5 | 259 | 70.4 (0) | 4.6814 | 0.0305* | 84.1 (−12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 0.7 | 251 | 79.6 (12) | 40.489 | <0.001* | 102.8 (8) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 1.5 | 278 | 51.5 (−27) | 266.83 | <0.001* | 71 (−26) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | Corn | 241 | 62.8 (−11) | 21.061 | <0.001* | 88.8 (7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 1.0 | 261 | 49.7 | 71.3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.5 | 261 | 50.4 (1) | 0.3308 | 0.5652 | 64.8 (−9) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.7 | 263 | 55.4 (11) | 21.498 | <0.001* | 73.1 (3) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 1.5 | 244 | 44.4 (−11) | 24.249 | <0.001* | 65 (−9) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | Corn | 263 | 55.7 (12) | 25.086 | <0.001* | 76 (7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.0 | 272 | 54.1 | 72.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.5 | 264 | 57.6 (6) | 17.045 | <0.001* | 72.9 (0) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.7 | 275 | 59.8 (10) | 44.583 | <0.001* | 76.7 (6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.5 | 275 | 46.1 (−15) | 62.478 | <0.001* | 63 (−13) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | Corn | 257 | 60.2 (11) | 52.928 | <0.001* | 78 (7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) rpd3def/+ (Top) and +/+ control (Bottom) flies on different food levels. All values are compared to values obtained for the flies of the same sex and of the same genotype kept on 1.0N food to determine the percent increase in mean and maximal lifespan. Food N: Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Some nutrient-mediated changes in life span differed for flies carrying the two rpd3 alleles. In male and female flies carrying the rpd3def allele and control flies for both alleles (rpd3def controls, and rpd3P-1.8) the peak extension due to DR (maximal extension of mean longevity compared to 1.0N food) occurs at 0.7N (Figure 1C,D and Table 1–4). However, in male and female flies carrying the rpd3P-UTR allele the peak extension due to DR occurs at 0.5N (Figure 1D, and Table 3, 4). The effect of high calories (1.5N) also differed. The mean life span is unchanged for rpd3P-UTR and rpd3P-1.8 flies on 1.5N and 1.0N food, but for rpd3def and rpd3def controls the mean life span is lower on 1.5N compared to 1.0N food. Since all of the survival studies reported here were performed at the same time with the same batches of food, precluding food preparation and fly culture as sources of variability, and since the two controls for each allele exhibited different responses to high calorie food (1.5N), it is likely that the response to high calorie food was affected by the genetic background. It has been shown by other groups that flies with different genetic backgrounds can differ in their responses to nutrients [30].
Table 3. Effect of rpd3P-UTR/+ mutation on longevity of flies on different food levels
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | rpd3P-1.8/+ | 1.0 | 228 | 54.2 | 70.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 1.0 | 219 | 57.4 (6) | 10.212 | 0.0014 | 72.61 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.1 | 70 | 11.6 | 15.4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.1 | 237 | 17.6 (51) | 206.86 | <0.001* | 26.1 (70) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.3 | 226 | 54.5 | 66.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.3 | 206 | 58.1 (7) | 36.901 | <0.001* | 71.1 (7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.5 | 223 | 58.7 | 74.4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.5 | 225 | 62.4 (7) | 23.307 | <0.001* | 85.7 (15) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.7 | 227 | 60.3 | 75.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.7 | 213 | 58.7 (−3) | 5.7376 | 0.0166* | 74.6 3 (−1) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 1.5 | 228 | 58.0 | 73.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 1.5 | 231 | 60.0 (3) | 4.4888 | 0.0341* | 76.8 (5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | Corn | 227 | 65.5 | 87.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | Corn | 218 | 68.1 (1) | 0.0645 | 0.7995 | 82.6 (−6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 1.0 | 216 | 56.2 | 69.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 1.0 | 223 | 63.0 (12) | 83.590 | <0.001* | 79.4 (24) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.1 | 58 | 15.6 | 23.2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.1 | 219 | 23.6 (52) | 120.64 | <0.001* | 48.7 (110) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.3 | 225 | 61.0 | 78 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.3 | 199 | 61.9 (2) | 9.0925 | 0.0026 | 80.7 (4) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.5 | 224 | 64.0 | 77.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.5 | 218 | 72.4 (13) | 112.55 | <0.001* | 89.2 (15) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.7 | 224 | 67.3 | 80.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.7 | 232 | 68.8 (2) | 13.142 | 0.0003* | 85.5 ()6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 1.5 | 230 | 56.6 | 71.4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 1.5 | 226 | 63.0 (10) | 60.616 | <0.001* | 80.6 (13) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | Corn | 224 | 55.0 | 76.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | Corn | 230 | 57.0 (4) | 24.759 | <0.001* | 88.8 (17) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) rpd3P-UTR/+ and rpd3P-1.8/+ control flies on different food levels. All values for control group are compared to values of rpd3P-UTR/+ flies of the same sex and on the same food levels to determine the percent increase in mean and maximal lifespan. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days except for longevity experiments on 0.1N. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table 4. Effects of DR on mean and maximal longevity of rpd3P-UTR/+ (Top) and rpd3P-1.8/+ (Bottom) flies
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | rpd3P-UTR/+ | 1.0 | 219 | 57.4 | 72.61 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.1 | 237 | 17.6 (−70) | 506.18 | <0.001* | 26.1(−64) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.3 | 206 | 58.1 (1) | 0.0011 | 0.9732 | 71.1 (−2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.5 | 225 | 62.4 (8) | 31.336 | <0.001* | 85.7 (18) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 0.7 | 213 | 58.7 (2) | 6.6398 | 0.0100* | 74.6 (3) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | 1.5 | 231 | 60.0 (4) | 8.9391 | 0.0028 | 76.8 (6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-UTR/+ | Corn | 218 | 68.1 (19) | 129.48 | <0.001* | 82.6 (14) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 1.0 | 223 | 63.0 | 79.4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.1 | 219 | 23.6 (−62) | 423.72 | <0.001* | 48.7 (−39) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.3 | 199 | 61.9 (−2) | 0.7415 | 0.3892 | 80.7 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.5 | 218 | 72.4 (15) | 90.451 | <0.001* | 89.2 (12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 0.7 | 232 | 68.8 (9) | 52.246 | <0.001* | 85.5 (8) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | 1.5 | 226 | 63.0 (0) | 1.4257 | 0.2325 | 80.6 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-UTR/+ | Corn | 230 | 57.0 (−10) | 2.8411 | 0.0919 | 88.8 (12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 1.0 | 228 | 54.2 | 70.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.1 | 70 | 11.6 (−79) | 437.4 | <0.001* | 15.4 (−78) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.3 | 226 | 54.5 (0) | 4.0704 | 0.0436* | 66.6 (−6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.5 | 223 | 58.7 (8) | 19.005 | <0.001* | 74.4 (5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 0.7 | 227 | 60.3 (11) | 53.962 | <0.001* | 75.6 (7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | 1.5 | 228 | 58.0 (7) | 19.156 | <0.001* | 73.5 (4) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3P-1.8/+ | Corn | 227 | 65.5 (21) | 126.69 | <0.001* | 87.8 (24) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 1.0 | 216 | 56.2 | 69.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.1 | 58 | 15.6 (−72) | 398.44 | <0.001* | 23.2 (−67) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.3 | 225 | 61.0 (8) | 26.974 | <0.001* | 78 (12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.5 | 224 | 64.0 (14) | 80.351 | <0.001* | 77.5 (11) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 0.7 | 224 | 67.3 (20) | 163.90 | <0.001* | 80.8 (16) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | 1.5 | 230 | 56.6 (1) | 0.3637 | 0.5465 | 71.4 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3P-1.8/+ | Corn | 224 | 55.0 (−2) | 1.2871 | 0.2566 | 76.1 (9) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) rpd3P-UTR/+ (Top) and rpd3P-1.8/+ (Bottom) control flies on different food levels. All values are compared to values obtained for flies of the same sex and the same genotype kept on 1.0N food to determine the percent increase in mean and maximal lifespan. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
It is of interest that there is a DR peak for flies carrying both the null and hypomorphic rpd3 alleles, despite the potentially confounding effects of the two hybrid genetic backgrounds used in these experiments. For males carrying either allele, the DR peak is 7-9% higher than the mean life span at 1.0N, whereas for females carrying either allele the DR peak is 12-15% higher. Male controls for both alleles have a DR peak of 11%, whereas the female controls for rpd3def have a DR peak of 11% and the female controls for rpd3P-UTR have a DR peak of 20%. Finally, there is a trend when comparing mutants to controls. For both males and females carrying the rpd3P-UTR allele, the DR peak is lower in mutants compared to controls. A similar trend is seen for males but not females carrying the rpd3def allele.
rpd3 mutants live longer under conditions close to starvation
Decreasing food below the level that yields DR induces increasing starvation. Flies raised on 0.1N food since eclosion have much shorter life spans compared to flies raised on all higher nutrient levels, indicating a condition close to starvation. rpd3P-UTR flies live longer on 0.1N food compared to controls, indicating increased resistance to severe food deprivation (Figure 1E,F).
Effects of rpd3 on the expression of 4E-BP and Tor
Our results indicate that decreased rpd3 mRNA levels has a minor effect on DR-mediated longevity extension and a major effect on resistance to food deprivation. In order to gain more insight into the interaction between RPD3 and nutrient responses, we determined whether as a transcriptional regulator RPD3 might be affecting the mRNA levels for two post-transcriptional modulators of nutrient signaling, 4E-BP and Tor (Figure 2).
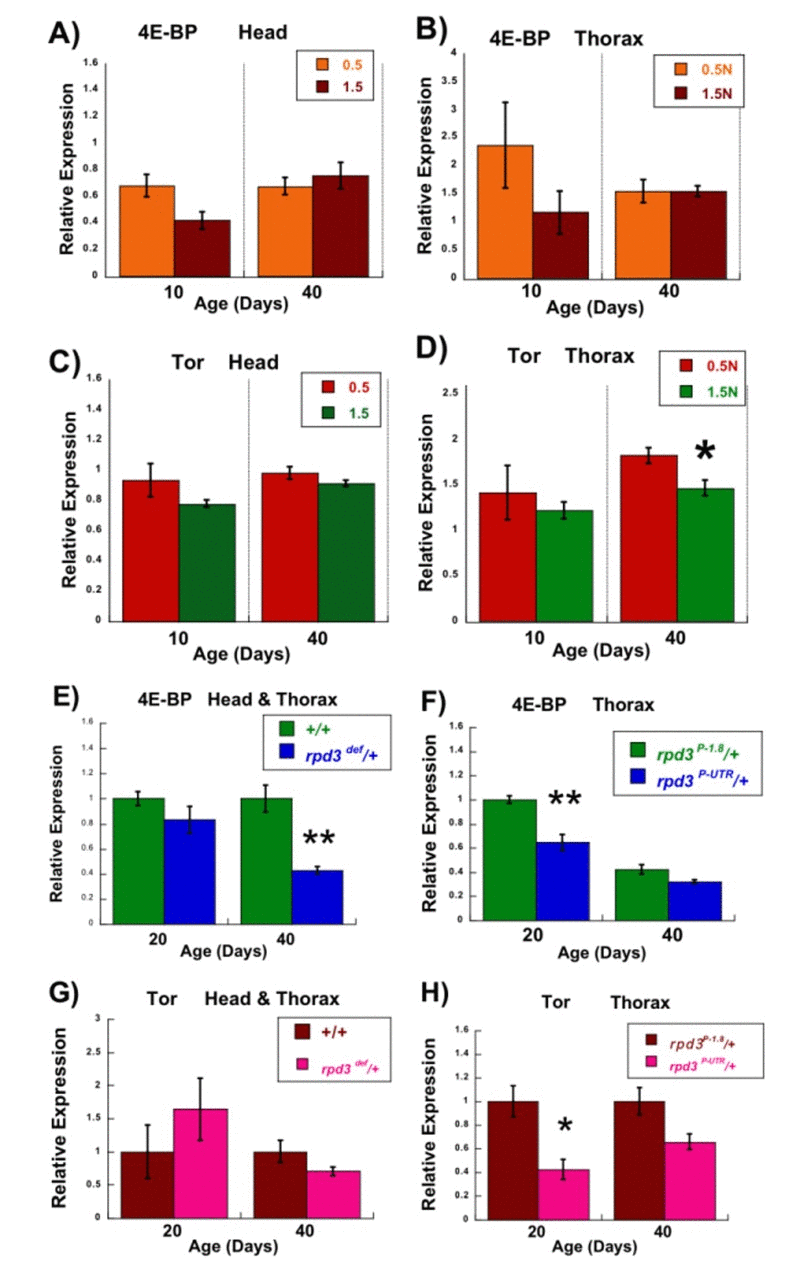
Figure 2. Transcription of the 4E-BP and Tor genes under conditions extending longevity. (A-D) Effects of DR on levels of 4E-BP and Tor mRNA Average expression of mRNA for the 4E-BP (A, B) and Tor (C, D) genes in three biological replicates isolated from the heads (A, C) and thoraces (B, D) of wild type CS female flies kept on 0.5N and 1.5N food at 10 or 40 days of age by Q-PCR. *p<0.05, 25 heads or thoraces per replicate. (E- H) Effects of rpd3 mutations on the levels of 4E-BP and Tor mRNA (E) Average expression of mRNA for the 4E-BP gene isolated from the heads and thoraces of rpd3def/+ and +/+ control male flies at 20 and 40 days of age by Q-PCR. (F) Average expression of mRNA for the 4E-BP gene isolated from thoraces of rpd3P-UTR/+ and control rpd3P-1.8/+ male flies at 20 or 40 days of age by Q-PCR. (G) Average expression of mRNA for the Tor gene isolated from the heads and thoraces of rpd3def/+ and +/+ control male flies at 20 and 40 days of age by Q-PCR. (H) Average expression of mRNA for the Tor gene isolated from the thoraces of rpd3P-UTR/+ and control rpd3P-1.8/+ male flies at 20 and 40 days of age by Q-PCR. In each case three biological replicates were used. Graphs are plotted as the means +/− standard errors. *p<0.05 **p<0.01, 25 heads + thoraces or thoraces per replicate.
4E-BP regulates mRNA translation and the Tor kinase has a central role in nutrient-mediated signal transduction [31]. We first determined whether 4E-BP and Tor mRNA levels are affected by nutrients in wild type Drosophila. Heads and thoraces were assayed separately, in case changes were localized to one area of the body. Young and old flies were also assayed, in case changes were affected by aging. Young (10 day) wild type female flies have modestly increased 4E-BP mRNA levels at 0.5N compared to 1.5N (similar trends in head and thorax, though the changes are not signifi-cant), whereas older (40 day) flies show no changes (Figure 2A,B). Another group has shown that 4E-BP protein levels are increased under DR conditions [32].
Tor mRNA shows much smaller changes, comparing 0.5N and 1.5N of young and old flies (Figure 2C,D), though the change in 40 day thoraces is significant. It was then determined whether rpd3 mutations affect the transcription of 4E-BP and Tor. When heterozygotes for the rpd3def allele are compared to genetic controls, young mutant male flies (20 days) show no changes in 4E-BP mRNA levels but older mutant flies (40 days) have 60% less mRNA (Figure 2E). Male flies heterozygous for the rpd3P-UTR allele have a 40% reduction in 4E-BP mRNA levels when younger and a smaller decrease relative to controls when older (only thoraces were assayed, see Methods) (Figure 2F). Flies heterozygous for the rpd3def allele have no significant changes in Tor mRNA level at both ages compared to genetic controls (Figure 2G). Flies heterozygous for the rpd3P-UTR allele have a 60% reduction in Tor mRNA when young and tend towards a decrease relative to controls when older (though this is not significant) (Figure 2H). In summary, the transcription of 4E-BP is affected by both rpd3 alleles, whereas only one allele affects Tor transcription. In all cases where there were significant changes, reduced dRPD3 levels led to decreased transcription. DR and rpd3 mutants differed in their effects upon 4E-BP and Tor transcription, providing further evidence that dRPD3 and nutrient status differ in their mechanism of life span extension.
RPD3 and 4E-BP have distinct effects on longevity
Several groups have reported that flies lacking any 4E-BP have lowered life spans at all nutrient levels [24, 32–34]. Since rpd3-mutant flies have moderately reduced levels of 4E-BP mRNA, we asked whether a moderate reduction in 4E-BP mRNA might affect life span. This was measured by using a hypomorphic allele of the 4E-BP gene, Thork07736. Flies carrying this allele were backcrossed to yw flies for 10 generations. Hetero-zygotes carrying the backcrossed Thork07736 (4E-BP) allele have a 30% reduction in 4E-BP mRNA (Figure 3A). Heterozygous Thor males have mean life spans that are reduced by 19%, 7%, and 5% relative to yw controls at 1.0N, 0.7N, and 0.5N food levels, respectively (no change at 1.5N) (Figure 3B, Table 5, 6). Heterozygous Thor females have mean life spans that are reduced by 12%, 33%, and 9% relative to yw controls at 1.5N, 1.0N, and 0.5N (no change at 0.7N) (Figure 3B, Table 5, 6). There were several additional differences between Thor mutants and their controls. Over a food range of 0.5N to 1.5N, yw controls show a peak life span at 1.0N, whereas Thor mutants have a peak at 0.5N (Figure 3B, Table, 5, 6). In addition, 1.5N food decreased the mean life spans of yw controls relative to 1.0N food but not the mean life spans of the Thor mutants. These differences between Thor mutants and the controls provide some evidence that 4E-BP can affect the response to nutrients.
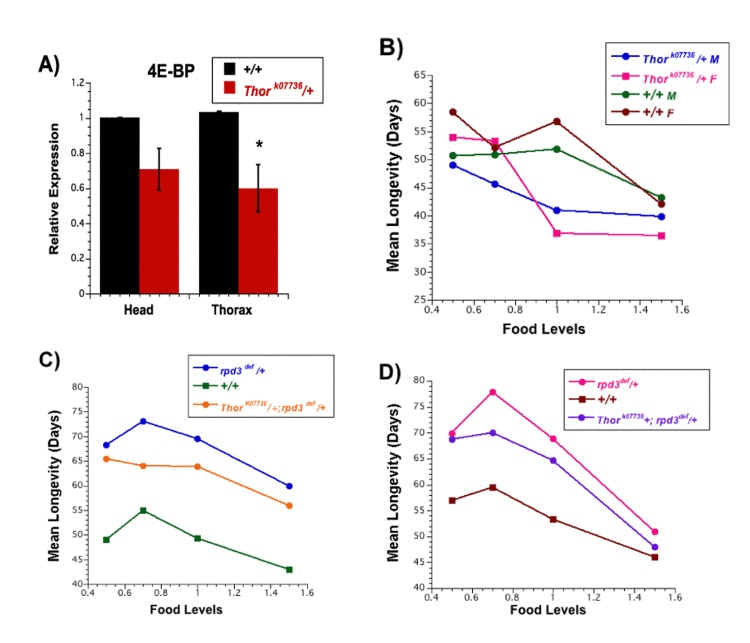
Figure 3. Longevity of flies carrying mutations in both rpd3 and Thor (4E-BP) (A) Average expression of mRNA for the 4E-BP gene in two (yw) or three (Thor mutant) biological replicates isolated from the heads and thoraces of female flies at 10 days of age by Q-PCR. Black, +/+ controls, magenta, Thork07736/+. Graphs are plotted as the means +/− standard errors. (B) Mean life span of Thork0773/+ and +/+ control (yw background) flies, male and female, on 0.5N, 0.7N, 1.0N, and 1.5N food levels.(C, D) Mean life span of male (C) and female (D) rpd3def/+, Thork0773 /+; rpd3def/+ double mutants and +/+ control flies on 0.5N, 0.7N, 1.0N, and 1.5 food levels.
Table 5. Effect of 4E-BP mutation on longevity of flies on different food levels
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | +/+ | 1.0 | 273 | 52.7 | 73.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 1.0 | 209 | 42.8 (−19) | 55.70 | <0.001* | 66.1 (−10) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.5 | 286 | 51.7 | 72.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 0.5 | 231 | 49.2 (−5) | 5.62 | 0.0178* | 66 (−8) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.7 | 277 | 51.1 | 73.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 0.7 | 221 | 47.6 (−7) | 9.68 | 0.0019* | 69.8 (−5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 1.5 | 277 | 43.3 | 64.93 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 1.5 | 201 | 41.8 (−3) | 3.38 | 0.0658 | 58.3 (−10) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | Corn | 280 | 54.0 | 71.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | Corn | 220 | 42.8 (1) | 3.58 | 0.0583 | 79.0 (10) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.0 | 273 | 57.6 | 74.2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 1.0 | 226 | 38.3 (−33) | 317.08 | <0.001* | 56.0 (−25) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.5 | 247 | 59.9 | 78.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 0.5 | 231 | 54.6 (−9) | 22.99 | <0.001* | 72 (−8) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.7 | 282 | 52.5 | 71 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 0.7 | 234 | 54.3 (3) | 12.69 | 0.0004* | 74.5 (5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.5 | 284 | 42.7 | 59.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 1.5 | 220 | 37.5 (−12) | 36.98 | <0.001* | 52.9 (−11) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | Corn | 273 | 58.6 | 75.9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | Corn | 222 | 59.4 (1) | 4.26 | 0.039 | 78.0 (3) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) +/+ control and 4E-BP (Thork07736/+) mutant flies on different food levels. All +/+ values are compared to values of Thork07736/+ group on the same food level and same sex to determine the percent increase in mean and maximal lifespan in 4E-BP mutant flies. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table 6. Effect of DR on longevity of 4E-BP hypomorphic (Top) and control (Bottom) flies
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | Thork07736/+ | 1.0 | 209 | 42.8 | 66.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 0.5 | 231 | 49.2 () | 16.914 | <0.001* | 66 (0) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 0.7 | 221 | 47.6 () | 12.807 | 0.0003* | 69.8 (6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | 1.5 | 201 | 41.8 () | 0.7227 | 0.3953 | 58.3 (−12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+ | corn | 220 | 42.8 () | 80.044 | <0.001* | 79.0 (20) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 1.0 | 226 | 38.3 () | 56.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 0.5 | 231 | 54.6 () | 208.70 | <0.001* | 72 (29) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 0.7 | 234 | 54.3 () | 191.31 | <0.001* | 74.5 (33) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | 1.5 | 220 | 37.5 () | 0.2207 | 0.6385 | 52.9 (6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+ | corn | 222 | 59.4 () | 282.08 | <0.001* | 78.0 (40) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 1.0 | 273 | 52.7 | 73.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.5 | 286 | 51.7 | 1.745 | 0.1865 | 72.6 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 0.7 | 277 | 51.1 | 0.5569 | 0.4555 | 73.5 (0) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | 1.5 | 277 | 43.3 | 58.062 | <0.001* | 64.93 (−12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | +/+ | corn | 280 | 54.0 | 2.1704 | 0.1407 | 71.8 (−3) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.0 | 273 | 57.6 | 74.2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.5 | 247 | 59.9 | 10.077 | 0.0015* | 78.9 (6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 0.7 | 282 | 52.5 | 28.028 | <0.001* | 71 9 (−4) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | 1.5 | 284 | 42.7 | 226.87 | <0.001* | 59.6 (−20) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | +/+ | corn | 273 | 58.6 | 2.81 | 0.0937 | 75.9 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) 4E-BP (Thork07736/+) (Top) mutant and control (Bottom) +/+ flies on different food levels. All values are compared to values of flies the same sex and genotype kept on 1.0N food to determine the percent increase in mean and maximal lifespan. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
We next examined the longevity of flies heterozygous for both Thork07736 (4E-BP) and rpd3def to directly assess the combined effects of these gene products (Figure 3C,D, Table 7, 8). Preliminary experiments indicated that rpd3 mutations mediated a decrease in 4E-BP mRNA when the flies also carried a Thor mutation (double heterozygotes) (Fig. Supplemental 2), similar to the decrease mediated by rpd3 mutations in the absence of Thor (Fig. 2). Double heterozygotes, single hetero-zygotes, and control flies shared the same genetic background. rpd3-mediated life span extension was attenuated but not eliminated by the presence of the Thor allele (Figure 3 C,D, Table, 7, 8). The reduction in mean life span of the double mutants relative to flies singly mutant for rpd3def ranged from 4% - 14% for males and 3% - 12% for females. While female flies that were doubly heterozygous maintained a smaller DR peak of 0.7N, male flies that were doubly heterozygous had no DR-mediated increase in life span. Doubly heterozygous flies of both sexes had decreased life spans at 1.5N relative to 1.0N. The simplest interpretation would be an overall superimposition of the life span decrease mediated by reduced 4E-BP on the life span extension mediated by reduced dRPD3.
Table 7. 4E-BP (Thork07736) mutation affects longevity of rpd3def/+ flies on different food levels
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | rpd3def/+ | 1.0 | 236 | 70.6 | 93.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 1.0 | 256 | 64.3 (−9) | 142.91 | <0.001* | 88.8 (−5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 0.5 | 247 | 68.8 | 3.6229 | 0.57 | 95.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 0.5 | 259 | 65.9 (−4) | 237.93 | <0.001* | 87.4 (−9) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 0.7 | 243 | 75.2 | 16.912 | <0.001* | 103.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 0.7 | 244 | 64.7 (−14) | 101.36 | <0.001* | 87.0 (−16) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | 1.5 | 254 | 61.2 | 58.619 | <0.001* | 84 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 1.5 | 228 | 57.1 (−7) | 118.91 | <0.001* | 75.1 (11) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | rpd3def/+ | Corn | 240 | 76.5 | 21.005 | <0.001* | 105.3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | Corn | 242 | 68.5 (10) | 124.82 | <0.001* | 90.9 (14) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 1.0 | 269 | 70.8 | 95.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 1.0 | 276 | 64.8 (−8) | 130.99 | <0.001* | 89.0 (−7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 0.5 | 259 | 70.4 | 4.6814 | 0.0305* | 84.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 0.5 | 274 | 68.6 (−3) | 164.07 | <0.001* | 87.9 (5) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 0.7 | 251 | 79.6 | 40.489 | <0.001* | 102.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 0.7 | 275 | 69.9 (−12) | 105.99 | <0.001* | 94.5 (−8) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | 1.5 | 278 | 51.5 | 266.83 | <0.001* | 71 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 1.5 | 263 | 49.5 (−4) | 16.049 | <0.001* | 70.1 (0) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | rpd3def/+ | Corn | 241 | 62.8 | 21.061 | <0.001* | 88.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | Corn | 258 | 62.2 (−1) | 5.401 | 0.0201* | 78 (−12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) rpd3def/+ and Thork07736/+;rpd3def/+ mutant flies on different food levels. All rpd3def/+ values are compared to Thork07736/+;rpd3def/+ value mutant of the same sex and on the same food level to determine the percent increase in mean and maximal lifespan. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table 8. 4E-BP mutation affects longevity of rpd3def/+ flies on different food levels
| Gender | Genotype | Food (N) | Number | Mean Lifespan (% Change to controls) | X2 | p | Maximal Lifespan (% Change) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M | Thork07736/+;rpd3def/+ | 1.0 | 256 | 64.3 | 88.8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 0.5 | 259 | 65.9 (2) | 2.1279 | 0.1446 | 87.4 (−2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 0.7 | 244 | 64.7 (0) | 1.34 | 0.247 | 87.0 (−2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | 1.5 | 228 | 57.1 (−11) | 38.915 | <0.001* | 75.1 (−15) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| M | Thork07736/+;rpd3def/+ | Corn | 242 | 68.5 (6) | 15.525 | <0.001* | 90.9 (2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 1.0 | 276 | 64.8 | 89.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 0.5 | 274 | 68.6 (6) | 8.6198 | 0.0033* | 87.9 (−1) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 0.7 | 275 | 69.9 (8) | 11.562 | 0.0007* | 94.5 (6) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | 1.5 | 263 | 49.5 (−24) | 165.63 | <0.001* | 70.1 (−21) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| F | Thork07736/+;rpd3def/+ | Corn | 258 | 62.2 (−4) | 6.7179 | 0.0095* | 78 (−12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| The mean and maximal lifespan of male (M) and female (F) Thork07736/+;rpd3def/+ mutant flies on different food levels. All values are compared to flies of the same sex kept on 1.0N food to determine the percent increase in mean and maximal lifespan. Food (N): standard food is 1.0N. Number: number of flies used in the experiment. Mean and maximal lifespan are in days. Data are censored for 0-10 days. Long-rank analyses were performed using the JMP 11 program. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In summary, DR-mediated longevity extension is maintained in the presence of rpd3 mutations, though it may have a moderately reduced magnitude. Our epistasis experiments using flies doubly mutant for rpd3 and 4E-BP suggest potential crosstalk between dRPD3 and nutrient signaling.
Discussion
Many single gene mutations that extend life span in Drosophila implicate particular pathways in longevity regulation, such as insulin and Tor signaling [1]. Dietary restriction is thought to extend life span through nutrient signaling and multiple effector pathways (intermediary metabolism, flux through the mitochondrial respiratory chains, control of protein synthesis rates, to name a few). The complexities of nutrient signaling at the cellular and tissue level are paralleled by the complexities of measuring nutrient effects in survival studies, since many variables can affect nutrient effects upon life span. Our earlier study showed that flies heterozygous for mutations in rpd3 extend longevity [8]. Connecting this transcriptional regulator to nutrient signaling would facilitate the identification of effectors for nutrient-mediated regula-tion, since some of the transcriptional targets of dRPD3 would be candidate effectors. The earlier study obtained preliminary evidence of a potential connection to DR. However, since only two nutrient levels were compared, corn and 0.5N, the present study aimed to more systematically test for a relationship between nutrient signaling and dRPD3-mediated longevity control. Our data indicate longevity extension by DR and dRPD3 are distinct, with some potential crosstalk or overlap.
Genetic background and gender have strong effects on longevity in Drosophila. The finding that reduced dRPD3 extends longevity has been shown using several genetic backgrounds and both sexes. In our earlier study, rpd3def flies were crossed to wild type CS flies prior to longevity measurements, whereas rpd3P-UTR and rpd3P-1.8 were crossed to a wm4 fly line (rpd3P-UTR had also been crossed into a wm4 genetic background prior to the longevity studies). In the present study rpd3def flies were crossed to yw flies, whereas rpd3P-UTR and rpd3P-1.8 flies were crossed to CS prior to longevity measurements (yw and wm4 are control strains for any relevant genes). The amount of life span extension varies from males to females and also varies for each allele depending upon the genetic background. It is therefore noteworthy that longevity extension was consistently obtained under all conditions, and that this effect has been replicated by a different research group using a different genetic background [25]. The Thor (4E-BP) allele was backcrossed into a yw background. In the present study all experiments with the rpd3def allele involve crosses with flies containing a yw background, so that every longevity experiment using this allele (whether singly or in combination with the Thor allele) had the same background. This entails a 50%/50% mix of the genetic background from the rpd3 mutant stocks (all rpd3 alleles are from a P-element mutagenesis that used a rosy strain) and the yw stock.
Every longevity experiment using the rpd3P-UTR and rpd3P-1.8 alleles has a 50%/50% mix of the rpd3 stock and CS strain.
Longevity studies in budding yeast have shown that rpd3 deletion extends replicative life span and rpd3 deletion mutants raised on DR have no additional extension of life span [18]. A subsequent study implicated RPD3 in a pathway that can regulate life span independently of DR but that converges on a common effector, the yeast homolog of S6K, indicating a potential connection [35]. Ours is the first study in metazoans to examine the longevity effects of RPD3 on a range of nutrient levels. We aimed to engage all potential nutrient-sensing pathways by varying both sugar and protein (yeast).
Three types of nutrient effects were examined in this study: DR (0.5N and 0.7N food), severe nutrient deprivation (0.1N and 0.3N food), and nutrient excess (1.5N food). Earlier studies used only one nutrient condition for DR (0.5N) [8, 9]. We observed DR-mediated longevity extension with both rpd3 alleles and both sexes. Decreased longevity on high calories (1.5N) relative to control food (1.0N) was only observed for the rpd3def allele (both sexes), though it was tested using both. The most severe nutrient deprivation (0.1N) was only tested using the rpd3P-UTR allele, with the mutant flies showing increased resistance to near starvation. This correlates with some but not all studies on the starvation resistance of DR flies. Such studies raise flies on standard or DR food, and then transfer the flies to conditions providing moisture but no nutrients. Virgin female flies subjected to DR were reported to have increased starvation resistance at younger ages but decreased resistance at later ages [36]. Young mated flies (both male and female) under DR were reported to have increased starvation resistance, but mated young male flies were also reported to have decreased resistance [37–39]. Genetic background, gender, and types of nutrients prior to starvation may account for some of the differences seen in starvation studies.
Since long-lived rpd3 flies exhibited DR-mediated life span extension, it is likely that dRPD3 and DR operate by distinct mechanisms. However, our longevity data does not rule out an effect of DR upon dRPD3-mediated longevity regulation. Potential interactions could be either upstream or downstream of dRPD3. An example of upstream regulation would be evidence of nutrient effects on rpd3 transcription. In one study newly eclosed flies were first fed a yeast-free diet (4 days) and then fed yeast (calorie and protein increase); addition of yeast led to an increase in rpd3 transcription [27]. The same study examined the transcription profile of Drosophila S2 cells (embryonic origin) that express a constitutively active form of the transcription factor dFOXO. Since decreased insulin signaling and DR activate dFOXO, this serves as a surrogate for those conditions. Cells with activated dFOXO had a marked decrease in rpd3 transcription. It is also possible that DR could have an effect downstream from dRPD3. For example, dRPD3 may modulate transcription of a gene that was also modulated by nutrients. We tested two genes that produce proteins representing convergence points for nutrient signaling, Tor and 4E-BP. Tor is a kinase that phosphorylates S6K and 4E-BP, thereby regulating metabolism and protein synthesis [31]. 4E-BP regulates translation in response to DR and other conditions [32]. Transcription of the Drosophila4E-BP gene is regulated by the dFOXO transcription factor in response to stress or starvation (40, 41]. 4E-BP protein levels are increased during DR in Drosophila and this increase has been linked to a variety of metabolic changes stimulated by DR [32]. We found that transcription of the 4E-BP gene is also regulated by dRPD3, providing some evidence for downstream crosstalk between DR and dRPD3.
Both alleles of rpd3 affected 4E-BP transcription, whereas only one allele affected Tor mRNA levels. We therefore asked whether longevity modulation by dRPD3 and DR involved crosstalk mediated by 4E-BP. A hypomorphic 4E-BP allele reduced 4E-BP transcription to a level roughly similar to the reduction obtained with rpd3 mutations. This 4E-BP allele was shown to reduce longevity, making it likely that we could detect the potential crosstalk. Since the rpd3 alleles increased longevity but maintained DR-mediated extension, and since the 4E-BP allele decreased longevity but maintained DR-mediated extension, we also asked whether double mutants would show maintenance of DR-mediated extension. dRPD3-mediated longevity extension was decreased by the presence of a 4E-BP mutation at all nutrient levels, suggesting distinct effects upon longevity. However, a DR effect was absent in males that are double mutants and was moderately decreased in females that are double mutants. This suggests either that 4E-BP helps mediate the DR effect, or more likely, a protein whose level is regulated by 4E-BP helps mediate the interaction between DR and dRPD3. A recent study examined whether Drosophila longevity extension mediated by chico mutations was dependent upon 4E-BP [42]; the chico gene product is part of the insulin signaling pathway. The longevity phenotype of chico heterozygotes was found to be partially dependent upon 4E-BP, but not the longevity phenotype of chico homozygotes. They found that 4E-BP-null flies have no altered longevity, but flies that were both 4E-BP null and chico heterozygotes had mean life spans intermediate between controls and chico heterozygotes (wild type for 4E-BP). Our study and the chico study illustrate that longevity regulation is complex, with multiple pathways affecting longevity and crosstalk between the pathways. In addition, longevity regulation seems to be highly sensitive to the dosage of proteins linked to this regulation. In summary, our results show that DR and dRPD3 can have discrete and therefore additive effects upon longevity. However, DR and dRPD3 show some interaction. Additional evidence is presented that implicates 4E-BP in the crosstalk between longevity regulation mediated by nutrients and dRPD3.
Material and methods
Fly strains and maintenance
rpd3-deficient (rpd3def24/TM6,Sb), rpd3–hypomorphic (rpd3P-UTR/TM3,Sb,Ser), and control for rpd3–hypomorphic (rpd3P-1.8/TM6,Tb) strains were used in our experiments. The hypomorphic rpd3P-UTR allele has a P-element inserted in the 5′UTR region of the rpd3 gene, close to the transcription start site, which affects expression throughout the fly's body [10]. The control rpd3P-1.8 allele has a P-element inserted 1.8 kb upstream from the transcriptional start site, which only decreases expression in the eye [10]. The rpd3P-UTR and rpd3P-1.8 alleles are derived from the same mutagenesis. The rpd3def allele was derived by excision of the P-element from rpd3P-UTR [10]. Canton S (CS), yw and Thork07736 flies were kindly provided by the Bloomington Stock center. Thork07736 flies were backcrossed to the yw strain for 10 generations to minimize the effects of genetic background.
Maintenance of aging flies
Flies were collected within 24 hours of eclosion and maintained using standard culture media in plastic vials. They were kept at 25°C in a humidified incubator on a 12 hour light: dark cycle. About 25 males and 25 females are kept together in each vial. For life span studies flies were passed every 2 days up to age 30 days and every day after that and the number of dead flies were counted. The number of flies in each life span study is listed in Tables 1 – 8. Standard corn food was prepared as previously described [43, 44]. The nutrient media used are dilutions (0.1N, 0.3N, 0.5N, 0.7N) or concentrations (1.5N) of an arbitrary normal condition, 1.0N, containing 100 g/L brewer's yeast (MP Biomedicals, Inc.), 100 g/L sucrose (MP Biomedicals, Inc.), 20 g/L agar, and 10 mg/L tegosept. Our nutrient protocol coordinately changes both sugar and protein by varying sucrose and yeast [26, 29]. The concentrations of agar and tegosept are not varied for the various media.
Genetic crosses
Male yw flies were crossed to female rpd3def24/TM6,Sb. F1 progeny not carrying the TM6 balancer were used for life span measurements (flies carrying the mutation). F1 progeny carrying the balancer were crossed to each other and F2 progeny not carrying the balancer and not yw were used as the life span controls. Male flies homozygous for Thork07736 (backcrossed into a yw genetic background) were crossed to female rpd3def24/TM6,Sb flies. F1 progeny not carrying the balancer were used for life span measurements (doubly mutant flies). Life span measurements with flies only heterozygous for Thork07736 were performed with progeny from crosses of Thork07736 homozygotes and yw. For studies using the other rpd3 alleles, rpd3P-UTR/TM3,Sb or rpd3P-1.8/TM6,Sb were crossed to wild type CS and F1 progeny not containing the balancer were used for life span measurements.
Quantitative PCR (Q-PCR)
Flies were separated by sex and frozen at the appropriate age. Thoraces of rpd3P-UTR /+ and rpd3P-1.8/+ flies were dissected since the rpd3P-1.8 allele has reduced expression in the eye. Heads and thoraces were dissected from rpd3def/+ and +/+ flies since the rpd3 is reduced throughout the body in rpd3def/+. RNA was isolated by using the standard Chomczynski protocol and Trizol reagent (Gibco BRL) [44]. cDNA was synthesized from RNA. Using TaqMan primers and the Applied Biosystems Thermal Cycler (AB 7500 System), levels of gene expression were determined. Ankyrin was used as an endogenous control [46].
Statistical analysis
Statistical analysis of Q-PCR results was done using the Student's T-Test from at least three independent experiments and expressed as P values. Error bars represent the standard errors, and P values are specifically indicated in the figures. Life span data were analyzed by Log-Rank tests using the JMP-11 program (SAS). All survivorship data were censored for the first 10 days. Maximum life span was calculated as the mean life span of the longest surviving 10% of the population.
Supplementary Materials
Acknowledgments
We thank Suzanne Kowalski and Luke Piscitelli for their excellent technical help.
Editorial Note
While this paper was in Review, it was reported that RNAi-mediated down regulation of RPD3 mRNA in flies leads to life span extension [47].
Funding
This work was supported by a grant from the National Institutes of Health AG023088 and Institutional support to B. Rogina. B. Rogina is a recipient of a Glenn Award for Research in Biological Mechanisms of Aging. S. Frankel was supported by the Greenberg Junior Faculty and Dean's Research Fund grants from the University of Hartford.
Conflicts of Interest
Authors declare that they have no financial, personal, or professional conflicts of interest.
References
- 1. Kenyon CJ. The genetics of ageing. Nature. 2010; 464: 504 -512. [PubMed] .
- 2. Katewa SD and Kapahi P. Role of Tor signaling in aging and related biologica processes in Drosophila melanogaster. Exp. Gerontol. 2011; 46: 382 -390. .
- 3. Kapahi P, Zid BM, Harper T, Koslover D, Sapin V, Benzer S. Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr. Biol. 2004; 14: 885 -890. [PubMed] .
- 4. Kaeberlein M, Powers RW 3rd, Steffen KK, Westman EA, Hu D, Dang N, Kerr EO, Kirkland KT, Fields S, Kennedy BK. Regulation of yeast replicative life by Tor and Sch9 in response to nutrients. Science. 2005; 310: 1193 -6. [PubMed] .
- 5. Jia K, Chen D, Riddle DL. The Tor pathway interacts with the insulin signaling pathway to regulate C. elegans larval development, metabolism and life span. Development. 2004; 131: 3897 -906. [PubMed] .
- 6. Lamming DW, Ye L, Katajisto P, Goncalves MD, Saitoh M, Stevens DM, Davis JG, Salmon AB, Richardson A, Ahima RS, Guertin DA, Sabatini DM, Baur JA. Rapamycin-induced insulin resistance is mediated by mTORC2 loss and uncoupled from longevity. Science. 2012; 335: 1638 -1643. [PubMed] .
- 7. Haberland M, Montgomery RL, Olson EN. The many roles of histone deacetylases in development and physiology: implication for disease and therapy. Nat. Rev. Genet. 2009; 10: 32 -42. [PubMed] .
- 8. Rogina B, Helfand SL, Frankel S. Longevity regulation by Drosophila Rpd3 deacetylase and caloric restriction. Science. 2002; 298: 1745 [PubMed] .
- 9. Rogina B and Helfand SL. Sir2 mediates longevity in the fly through a pathway related to calorie restriction. Proc. Natl. Acad. Sci. USA. 2004; 101: 15998 -16003. [PubMed] .
- 10. Mottus R, Sobel RE, Grigliatti TA. Mutational analysis of a histone deacetylase in Drosophila melanogaster: missense mutations suppress gene silencing associated with position effect variegation. Genetics. 2000; 154: 657 -668. [PubMed] .
- 11. Frankel S, Ziafazeli T, Rogina B. dSir2 and longevity in Drosophila. Exp. Gerontol. 2011; 46: 391 -6. [PubMed] .
- 12. Whitaker R, Faulkner S, Miyokawa R, Burthenn L, Henriksen M, Wood JG, Helfand SL. Increased expression of Drosophila Sir2 extends life span in a dose-dependent manner. Aging. 2013; 5: 682 -91. [PubMed] .
- 13. Edwards C, Canfield J, Copes N, Rehan M, Lipps D, Bradshaw PC. D-beta-hydroxybutyrate extends lifespan in C. elegans. Aging. 2014; 6: 621 -44. [PubMed] .
- 14. Kang H-L, Benzer S, Min K-T. Life extension in Drosophila by feeding a drug. Proc. Natl. Acad. Sci. USA. 2002; 99: 838 -843. [PubMed] .
- 15. Zhao Y, Sun H, Lu J, Li X, Chen X, Tao D, Huang W, Huang B. Lifespan extension and elevated hsp gene expression in Drosophila caused by histone deacetylase inhibitors. J. Exp. Biol. 2005; 208: 697 -705. [PubMed] .
- 16. McDonald P, Maizi BM, Arking R. Chemical regulation of mid-and late-life longevities in Drosophila. Exp. Gerontol. 2013; 48: 240 -9. [PubMed] .
- 17. Kim S, Benguria A, Lai C-Y, Jazwinski SM. Modulation of life-span by histone deacetylase genes in Saccharomyces cerevisiae. Mol. Biol. Cell. 1999; 10: 3125 -56. [PubMed] .
- 18. Jiang JC, Wawryn J, Shantha Kumara HMC, Jazwinski SM. Distinct roles of processes modulated by histone deacetylases Rpd3, Hda1p, and Sip2 in life extension by caloric restriction in yeast. Exp. Gerontol. 2002; 37: 1023 -30. [PubMed] .
- 19. McCay CM, Crowell MF, Maynard LA. The effect of retarded growth upon the length of life span and upon the ultimate body size. J. Nutr. 1935; 10: 63 -79. .
- 20. Weindruch RH and Walford RL. The Retardation of Aging and Disease by Dietary Restriction Springfield, IL C. C. Thomas 1988; .
- 21. Alic N and Partridge L. Death and dessert: nutrient signaling pathways and aging. Curr. Opin. Cell Biol. 2011; 23: 738 -743. [PubMed] .
- 22. Mair W, Piper MDW, Partridge L. Calories do not explain extension of life span by dietary restriction in Drosophila. PLoS Biol. 2005; 3: e223 [PubMed] .
- 23. Partridge L, Piper MDW, Mair W. Dietary restriction in Drosophila. Mech Aging Dev. 2005; 126: 938 -950. [PubMed] .
- 24. Tatar M. The plate half-full: Status of research on the mechanisms of dietary restriction in Drosophila melanogaster. Exp. Gerontol. 2011; 46: 363 -368. [PubMed] .
- 25. Pallos J, Bodai L, Lukacsovich T, Purcell JM, Steffan JS, Thompson LM, Marsh JL. Inhibition of specific HDACs and sirtuins suppresses pathogenesis in a Drosophila model of Huntington's disease. Hum. Mol. Genet. 2008; 17: 3767 -3775. [PubMed] .
- 26. Bross TG, Rogina B, Helfand SL. Behavioral, physical, and demographic changes in Drosophila populations through dietary restriction. Aging Cell. 2005; 4: 309 -317. [PubMed] .
- 27. Gershman B, Puig O, Hang L, Peitzch RM, Tatar M, Garofalo RS. High-resolution dynamics of the transcriptional response to nutrition in Drosophila: a key role for dFOXO. Physiol. Genomics. 2007; 29: 24 -34. [PubMed] .
- 28. De Rubertis F, Kadosh D, Henchoz S, Pauli D, Reuter G, Struhl K, Spierer P. The histone deacetylase RPD3 conteracts genomic silencing in Drosophila and yeast. Nature. 1996; 384: 589 -91. [PubMed] .
- 29. Chapman T and Partridge L. Female fitness in Drosophila melanogaster: an interaction between the effects of nutrition and of encounter rate with males. Proc. R. Soc. Lond. B Biol. Sci. 1996; 263: 755 -59. .
- 30. Grandison RC, Wong R, Bass TM, Partridge L, Piper MDW. Effect of a standardized dietary restriction protocol on multiple laboratory strains of Drosophila melanogaster. PLoS ONE. 2009; 4: e4067 [PubMed] .
- 31. Lasko P and Sonenberg N. Coordinated transcriptional and translational control in metabolic homeostasis in flies. Genes Dev. 2007; 21: 235 -237. [PubMed] .
- 32. Zid BM, Rogers AN, Katewa SD, Vargas MA, Kolipinski MC, Lu TA, Benzer S, Kapahi P. 4E-BP extends lifespan upon dietary restriction by enhancing mitochondrial activity in Drosophila. Cell. 2009; 139: 149 -160. [PubMed] .
- 33. Bauer J, Chang C, Bae G, Morris SNS, Helfand SL. Dominant-negative Dmp53 extends life span through the dTOR pathway in D. melanogaster. Mech. Ageing Dev. 2010; 131: 193 -201. [PubMed] .
- 34. Partridge L, Alic N, Bjedov I, Piper MDW. Aging in Drosophila. The role of the insulin/Igf and Tor signaling network. Exp. Gerontol. 2011; 46: 376 -381. [PubMed] .
- 35. Lu J-Y, Lin Y-Y, Sheu J-C, Wu J-T, Lee F-J, Chen Y, Lin M-I, Chiang F-T, Tai T-Y, Berger SL, Zhao Y, Tsai K-S, Zhu H, Chuang L-M, Boeke JD. Acetylation of yeast AMPK controls intrinsic aging independently of caloric restriction. Cell. 2011; 146: 969 -979. [PubMed] .
- 36. Burger JMS, Hwangbo DS, Corby-Harris V, Promislow DEL. The functional cost and benefits of dietary restriction in Drosophila. Aging Cell. 2007; 6: 63 -71. [PubMed] .
- 37. Chippindale AK, Leroi AM, Kim SB, Rose MR. Phenotypic plasticity and selection in Drosophila life-history evolution. I. Nutrition and cost of reproduction. J. Evol. Biol. 1993; 6: 171 -193. .
- 38. Katewa SD, Demontis F, Kolipinski M, Hubbard A, Gill MS, Perrimon N, Meloc S, Kapahi P. Intramyocellular fatty-acid metabolism plays a critical role in medicting responses to dietary restriction in Drosophila melanogaster. Cell Metab. 2012; 16: 97 -103. [PubMed] .
- 39. Wang PY, Neretti N, Whitaker R, Hosier S, Chang C, Lu D, Rogina B, Helfand SL. Long-lived Indy and calorie restriction interact to extend life span. Proc Natl Acad Sci USA. 2009; 106: 9262 -9267. [PubMed] .
- 40. Teleman AA, Chen Y-W, Cohen SM. 4E-BP functions as a metabolic brake used under stress conditions but not during normal growth. Genes Dev. 2005; 19: 1844 -1848. [PubMed] .
- 41. Tettweiler G, Miron M, Jenkins M, Sonenberg N, Lasko PF. Starvation and oxidative stress resistance in Drosophila are mediated through the eIF4E-binding protein, d4E-BP. Genes Dev. 2005; 19: 1840 -1843. [PubMed] .
- 42. Bai H, Post S, Kang P, Tatar M. Drosophila longevity assurance conferred by reduced insulin receptor substrate Chico partially requires d4eBP. PLOS One. 2015; e:0134415 [PubMed] .
- 43. Rogina B and Helfand SL. Indy mutations and Drosophila longevity. Front. Genet. 2013; 4: 47 [PubMed] .
- 44. Woods J, Kowalski S, Rogina B. Determination of the spontaneous locomotor activity in Drosophila melanogaster. J Vis. Exp. 2014; 86 https://doi.org/10.3791/51449 .
- 45. Chomczynski P and Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 1987; 162: 156 -9. [PubMed] .
- 46. Rogers RP and Rogina B. Increased mitochondrial biogenesis preserves intestinal stem cell homeostasis and contributes to longevity in Indy mutant flies. Aging. 2014; 4: 335 -50. [PubMed] .
- 47. Kopp ZA, et al. Heart-specific Rpd3 downregulation enhances cardiac function and longevity. Aging. 7: 648 -663. [PubMed] .


