Introduction
To counteract DNA damage, specific mechanisms have been evolved, these are activated by specific signal transduction pathways, such as the phosphatidylinositol 3-kinase-like protein kinases (PIKKs) family, ATM, ATR and DNA-PK, and the members of the poly(ADP)ribose polymerase (PARP) family [1–7]. Among the effectors, a key role is played by the tumour suppressor protein p53 [8–15]. Indeed, DNA damage leads to p53 stabilisation by inhibition of interaction with its ubiquitin ligase, MDM2 [9], and as consequence, p53 transcriptionally induces cell cycle arrest, apoptosis or senescence. Among p53 targets in response to DNA damage, there are the CDK inhibitor p21, the pro-apoptotic proteins BAX and PUMA [16]. Moreover, p53 directly activates repair pathways such as nucleotide excision repair (NER) through regulation of the NER factors XPC and DDB2 and induces dNTP synthesis [17].
In addition to its roles in cell death, p53 has also been implicated in cytoskeleton assembly, cell motility and mechanosignaling, as negative regulator of cancer cell mobility, invasion and metastasis [18–20]. Integrin expression and signalling pathways, which play a key role in tumour cell invasion and metastasis, have been reported to be regulated indirectly by p53 [18]. For instance, Nutlin-3a, an MDM2 antagonist that acts as p53 activator, decreases the expression of integrin alpha5 in colorectal cancer and glioma cells [21,22]; also the expression of integrin beta3 decreases upon DNA-damage in wild-type p53 expressing cells [23]. p53 also regulates focal adhesion and Rho signalling pathways by regulating Rho GTPase activity [24] and effector protein genes of RhoA/RhoC and Cdc42 pathways [25–28]. In addition, F-actin formation is negatively or positively regulated by p53 in response to DNA damage depending on the anti-tumour drug used and cell type. For instance, while doxorubicin increases the expression of RhoC and LIM kinase 2 in a p53-dependent manner promoting actin stress fibers formation [29], etoposide and camptothecin attenuate this process through p53-dependent expression of RhoE [30]. It has been also reported that upon etoposide-mediated DNA damage, p53 alters actin cytoskeleton by transcriptionally induction of the expression of the cytoskeleton adaptor protein ankyrin-1 [31]. The relevance of cytoskeleton remodelling and cell mobility in tumours is evidenced by the fact that mutant p53 promotes tumour cell invasion and results in loss of directionality during migration [32]. Cytoskeleton remodelling and cell migration in cancer is a complex process and is controlled by many proteins and pathways, the specific role of p53 in these mechanisms is not yet completely understood.
Here, we describe a novel p53 target gene, ZNF185, which codifies for a Zn-finger protein belonging to LIM-family, activated upon genotoxic stress caused by DNA-damaging drug etoposide. ZNF185 itself is not necessary for p53-dependent cell cycle arrest and apoptosis, yet its silencing affects actin cytoskeleton changes and cell polarity upon etoposide treatment. At mRNA and protein level, ZNF185 is strongly reduced in different types of epithelial tumours, including skin and head and neck squamous cell carcinomas, suggesting that depletion of ZNF185 in cancer cells facilitate cancer cell migration and spreading.
Results
ZNF185 is a p53 target gene
We have previously shown that the p53 family member p63, using a novel promoter region and a specific enhancer, directly regulated ZNF185 expression in keratinocytes [33]. To investigate whether also p53 could regulate ZNF185 expression and expand the p53 target genes involved in cytoskeleton regulation and cell polarity, we further analysed ZNF185 promoter region using UCSC genome browser (Fig 1A). We observed several regions showing high accessibility and conservation between the species, and an enrichment in different transcription factors (TF) binding. Analysis of the publicity available ChIP-seq data for p53 performed in MCF7 cells after p53 stabilization by nutlin (GSE86164 [34],), revealed a strong peak within ZNF185 promoter only in nutlin-treated cells (Fig 1B), suggesting p53 involvement in regulation of ZNF185 transcription. Using previously described bioinformatic tool for p53 binding site (bs) prediction [35], we identified a putative binding site for p53 within the genomic region corresponding to the peak from the ChIP seq shown in Fig 1B (Fig 1C). Interestingly, this region is conserved only in primates and is absent in other species (Fig 1C). To confirm physical binding of p53 on ZNF185 promoter, we performed ChIP assay in p53 Tet-On inducible SaOs-2 cells previously generated in the laboratory [36]. As a positive control, we used the promoter of CDKN1A, the gene coding for p21 (Fig 1D). To confirm if p53 could directly regulate ZNF185 expression binding its promoter, we cloned the genomic locus harbouring p53 bs up-stream of the luciferase reporter gene. Luciferase activity assay showed a strong activation (120-fold, P<0.01) upon p53 overexpression. Interestingly, the overexpression of the two different p53 mutants frequently found in human cancers (R175H and R273H) did not show any strong activation compared to the control (Fig 1E), indicating that ZNF185 is target of wild-type p53. Furthermore, the substitution of cytosines and guanines to adenines within ZNF185 promoter sequence led to dramatic decrease of the luciferase activity (83% reduction, P<0.01) upon p53 overexpression (Fig 1F). To investigate if p53 is able to regulate ZNF185 transcription, we induced p53 expression by doxycycline in SaOs-2 Tet-On cells and measured by RT-qPCR a significant increase of ZNF185 mRNA after p53 induction paralleling CDKN1A increases (15-fold for ZNF185 and 5-fold for CDKN1A at 24 h of induction, P<0.05; Fig 1G). We also confirmed this result changing cellular system. Indeed, overexpression of p53 in H1299 also led to a 20-fold increase of ZNF185 mRNA, meanwhile the overexpression of two p53 mutants didn’t show any significant modulation (Fig 1H). Altogether, these data indicate ZNF185 as a bona fide transcriptional target of wild-type p53.
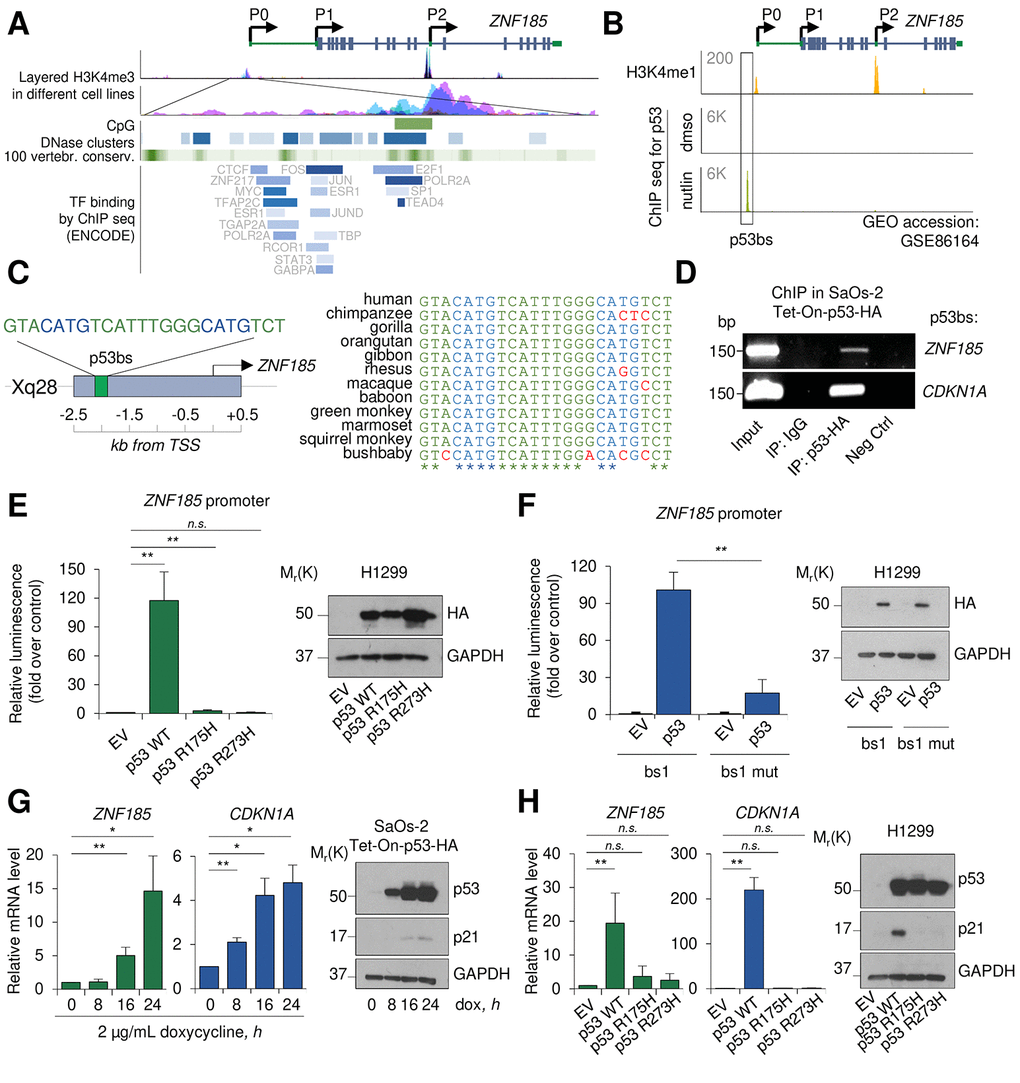
Figure 1. ZNF185 is a transcription target of p53. (A) UCSC genome browser analysis showing layered H3K4me3 mark in different cell lines, CpG islands, DNase clusters, conservation in vertebrates, and TF binding within ZNF185 promoter. (B) Genomic locus of ZNF185 showing the promoter region with H3K4me1 and p53 ChIP-seq signals after MCF7 treatment with either DMSO or nutlin. (C) Identified p53 binding site (p53 bs) within ZNF185 promoter region and conservation analysis among primates. (D) Amplification of specific DNA fragments after ChIP performed in SaOs-2 Tet-On-p53-HA cells using HA antibody. (E) Luciferase activity assay in H1299 after transfection of pGL3-ZNF185 promoter and either empty vector, p53 WT, p53-R175H, or p53-R273H expression vectors. ** P<0.01, n=4. Western blot analysis of cell lysates confirms p53 overexpression. (F) Luciferase activity assay in H1299 after transfection of pGL3-ZNF185 promoter with either WT of mutated p53 bs and either empty vector or p53 WT expression vectors. ** P<0.01, n=3. Western blot analysis of cell lysates to confirm p53 overexpression. (G) RT-qPCR analysis of ZNF185 and CDKN1A mRNA levels in SaOs-2 Tet-On-p53-HA after induction of p53 expression with 2 µg/mL doxycycline. * P<0.05, ** P<0.01, n=3. Western blot shows p53 and p21 levels. (H) RT-qPCR analysis of ZNF185 and CDKN1A mRNA levels in H1299 after transfection with empty vector, p53 WT, p53 R175H, or p53 R273H expression vectors. ** P<0.01, n=3. Western blot shows p53 and p21 levels.
ZNF185 is up-regulated upon DNA damage
We investigated whether ZNF185 is transcribed as consequence of p53 activation following DNA damage. Using two different carcinoma cell lines harbouring wild-type p53 (HCT116 and MCF7), we analysed ZNF185 expression after 0, 8, 16, and 24 hours of etoposide treatment. In both cases, we saw p53 stabilization as indicated by the western blots (Fig 2 A-B) and, as a consequence of p53 activation, significant up-regulation of ZNF185 mRNA (3-4-fold over control at 24 h of etoposide treatment, P<0.05), and p21 as positive control, both at mRNA and protein levels (Fig 2A-B). Interestingly, analysis of publicity available ChIP seq data (GSE56674 [37],) for p53 performed in keratinocytes showed that p53 binds to the locus within ZNF185 promoter identified by us in this study. Moreover, this binding is observed only upon cisplatin or doxorubicin treatment (Fig 2C). As a model of basal layer keratinocytes, we used the commercial cell line of immortalized keratinocytes, Ker-CT. We confirmed that also in Ker-CT cells etoposide treatment leads to p53 stabilization and ZNF185 up-regulation both at mRNA (3-fold, P<0.01) and protein levels (Fig 2D). To confirm that ZNF185 up-regulation is p53-dependent, we performed siRNA-mediated knock-down of p53 in Ker-CT cells. As expected, depletion of p53 abolished up-regulation of ZNF185 upon etoposide treatment (Fig 2E). Since the major source of DNA damage in the human keratinocytes is UV irradiation, we irradiated Ker-CT cells and analysed ZNF185 level. Also in this case, we saw an up-regulation of ZNF185 at protein level. Altogether, these findings show that upon DNA damage we detected up-regulation of ZNF185 expression in p53-dependent manner both in tumour cell lines and in normal human keratinocytes.
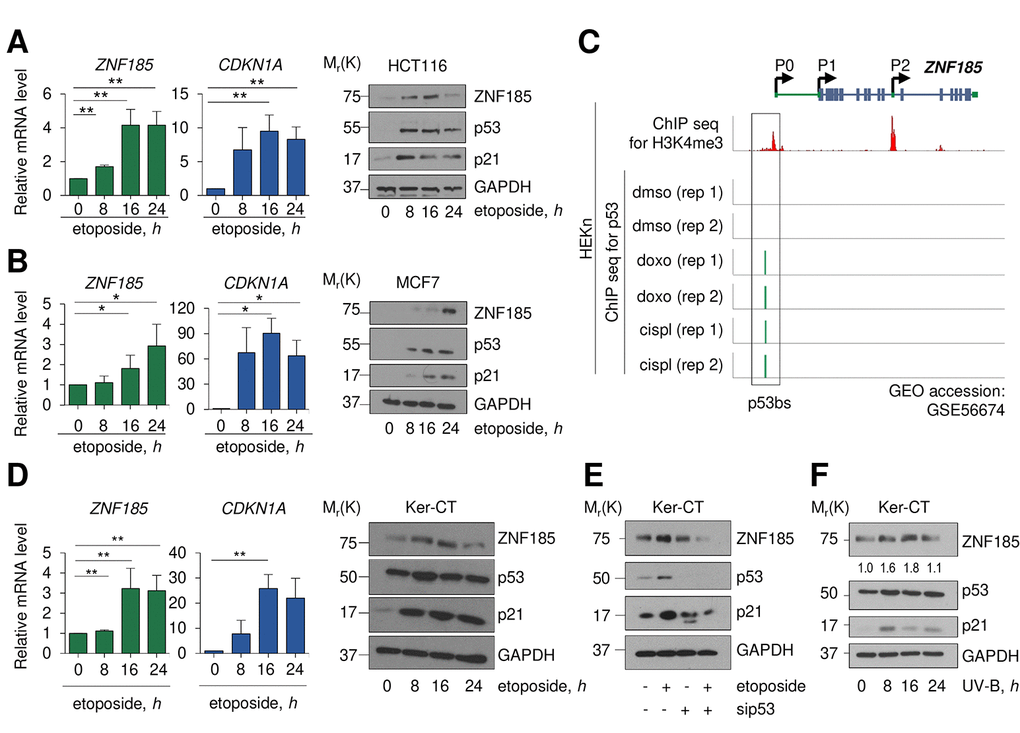
Figure 2. ZNF185 is up-regulated upon DNA damage. (A) qPCR analysis of ZNF185 and CDKN1A mRNA levels in HCT116 after 25 µM etoposide treatment. **P<0.01. Western blot shows ZNF185, p53, and p21 levels. (B) qPCR analysis of ZNF185 and CDKN1A mRNA levels in MCF7 after 25 µM etoposide treatment. Western blot shows ZNF185, p53, and p21 levels. *P<0.05 (C) Genomic locus of ZNF185 showing the promoter region with H3K4me3 and p53 ChIP-seq signals after HEKn treatment with either DMSO, doxorubicin, or cisplatin. (D) qPCR analysis of ZNF185 and CDKN1A mRNA levels in Ker-CT after 100 µM etoposide treatment. **P 0.05, n=3 (for ZNF185) and n=2 (for CDKN1A). Western blot shows ZNF185, p53, and p21 levels. (E) Western blot analysis of ZNF185, p53, and p21 levels after 100 µM etoposide treatment and p53 knock-down in Ker-CT cells. (F) Western blot analysis of ZNF185, p53, and p21 levels after 10 mJ.cm-2 UV-B treatment in Ker-CT cells for indicated times. Densitometry values of ZNF185 expression levels, normalized to GAPDH level, are shown.
ZNF185 is involved in the cytoskeleton remodelling upon DNA damage
Since the major functions of p53 activation upon DNA damage relate to the cell cycle arrest and apoptosis, we asked whether depletion of ZNF185 could alter cell cycle content under this specific stress condition. We performed siRNA mediated knock-down of ZNF185 in Ker-CT cells with two different siRNAs and treated the cells with etoposide. Cytofluorimetric analysis did not reveal any significant modulation in cell cycle distribution and apoptosis respect to the control (Fig 3A). It was previously reported that ZNF185 regulates proliferation of prostate cancer cells [38], to further investigate this point we generated Ker-CT cell line, stably expressing shRNA against ZNF185 (shZNF185). We performed the EdU-incorporation assay to evaluate the number of cells in S-phase, but we did not observe any significant difference in cell proliferation respect to the control (Fig 3B). Given that several LIM-domain Zn-fingers can migrate into the nucleus under stress conditions [39], we asked whether DNA damage can alter ZNF185 localisation. We found that ZNF185 localised in the cytoplasm and at the cell periphery (Fig 3C) also after etoposide treatment. Due to the presence of the actin-interacting domain within ZNF185 protein, we hypothesised that ZNF185 could be involved in cytoskeleton remodelling upon DNA damage. To this aim, we performed immunofluorescence analysis using phalloidin as a marker of filamentous actin and vinculin as a marker of focal adhesion. Under normal conditions, most of the cells had migratory phenotype showing vinculin accumulation on the leading edge. After etoposide treatment, cells lost planar polarity as visualised by homogeneous vinculin distribution on the cell periphery (percentage of polarized cells from 100% to 35%). Surprisingly, this phenotype was abolished in the shZNF185 cells which retained planar polarization also upon etoposide treatment (percentage of polarized cells from 100% to 82%) (Fig 3D). Altogether, these results suggest that ZNF185 is involved in the loss of the planar polarity of cells upon DNA damage.
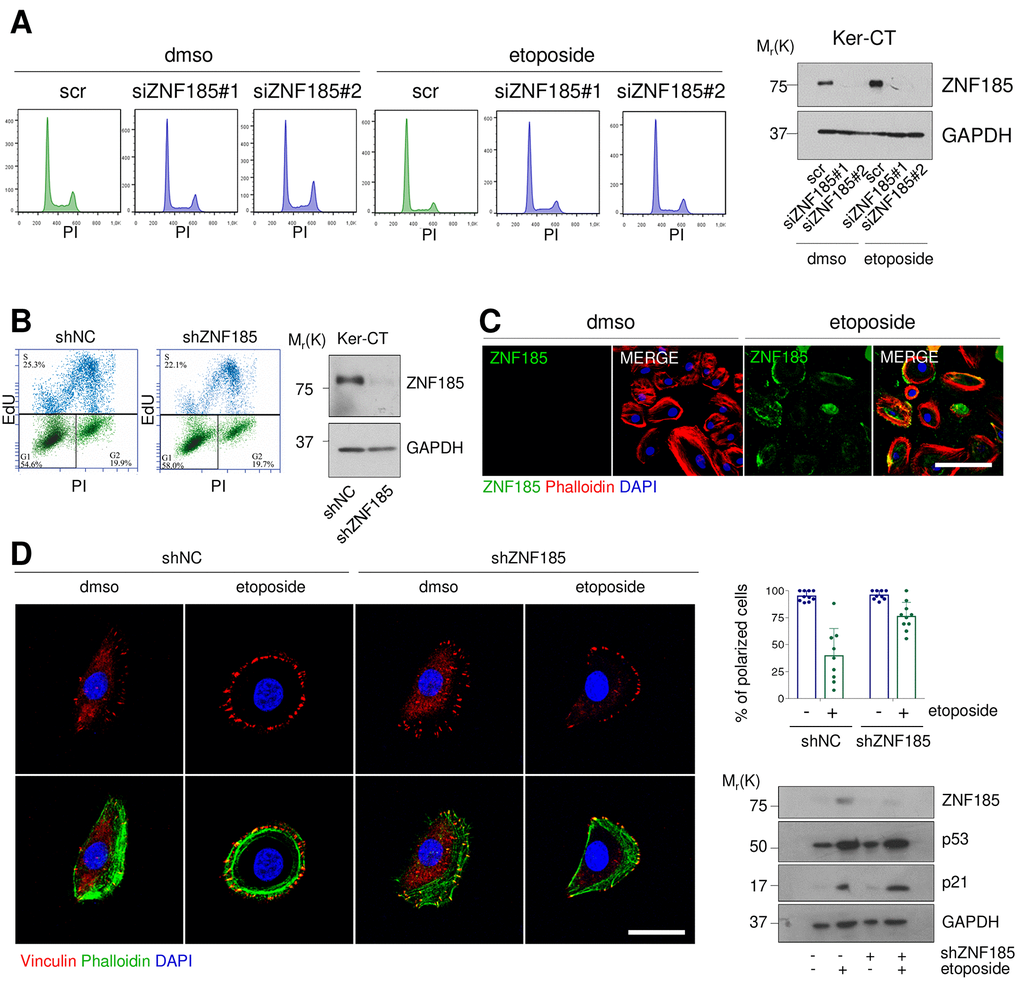
Figure 3. ZNF185 is involved in the cytoskeleton remodelling upon DNA damage. (A) FACS analysis of cell cycle content of the Ker-CT treated with either DMSO or 100 µM etoposide for 24 h after ZNF185 knock-down with two different siRNAs. Western blot confirms ZNF185 silencing. (B) EdU-incorporation assay by FACS showing % of EdU-positive Ker-CT shZNF185 cells. Western blot confirms the ZNF185 knock-down. (C) Immunofluorescence analysis of ZNF185 expression in Ker-CT treated with either DMSO or 100 µM etoposide for 16 h. Phalloidin was used for cytoskeleton staining. Scale bar: 50 µm. (D) Immunofluorescence analysis of vinculin distribution in Ker-CT treated with either DMSO or 100 µM etoposide and knocked-down for ZNF185. Phalloidin was used for cytoskeleton staining. Scale bar: 20 µm. In the right panel is shown the quantification of % of polarized cells in ten random fields. Western blot shows ZNF185, p53, and p21 levels.
ZNF185 is down-regulated in epithelial cancers
p53 is frequently mutated in human cancers and we have shown that ZNF185 is positively regulated by wild-type p53, therefore we asked if ZNF185 level is decreased in epithelial cancers and particularly in the skin carcinomas. Firstly, we analysed ZNF185 mRNA expression in different types of epithelial cancers from TCGA collection. Five cancer types - prostate adenocarcinoma (PRAD), chromophobe renal cell carcinoma (KICH), head and neck squamous cell carcinoma (HNSC), oesophageal carcinoma (ESCA), and adenoid cystic carcinoma (ACC) – showed a significant decrease in ZNF185 mRNA level respect to the normal tissues from both TCGA and GTEx database (Fig 4A). Furthermore, we analysed correlation between the expression of ZNF185 and two distinct targets of p53 – PERP and CDKN1A. Interestingly, a strong positive correlation was observed only in the cancers arising from squamous epithelia – oesophageal and head and neck carcinomas (Fig. 4B). Since there are only few datasets of skin cancer with a very low number of samples, we decided to analyse ZNF185 expression in skin cancer by immunohistochemistry using tissue microarray, containing 42 samples of the cutaneous squamous cell carcinoma (cSCC), 14 samples of the cutaneous basal cell carcinoma (cBCC), 12 samples of cutaneous malignant melanoma (cMM), and 10 samples of the normal skin. As a marker of proliferation, we used Ki67. Analysis of ZNF185 expression pattern at protein level in the normal skin confirmed previously published data from our laboratory [33], in which ZNF185 highest expression occurs in the differentiated spinous and granular layers (“SS/SG”) of the epidermis with low expression in the proliferating basal layer (“SB”). Cornified layer (“SC”) and dermis (“D”) were found negative for ZNF185 (Figure 5A). Analysis of skin cancer samples revealed that ZNF185 expression is dramatically down-regulated in the cutaneous squamous and basal cell carcinoma (“cSCC” and “cBCC”) and malignant melanoma (“cMM”) samples (Figure 5B). Furthermore, ZNF185 was found only in well-differentiated subpopulations of squamous cell carcinoma (“WD” of cSCC) in contrast to poorly-differentiated basal-like subpopulations (“PD” of cSCC) (Figure 5C). All the tumour samples showed a significant decrease (P<1x10-5) of ZNF185 H-score respect to the differentiated layers of the normal epidermis (Figure 5D). These findings reveal a dramatic down-regulation of ZNF185 at protein level in the skin cancer and suggest that ZNF185 could be a potential biomarker for epithelial cancer diagnosis and prognosis.
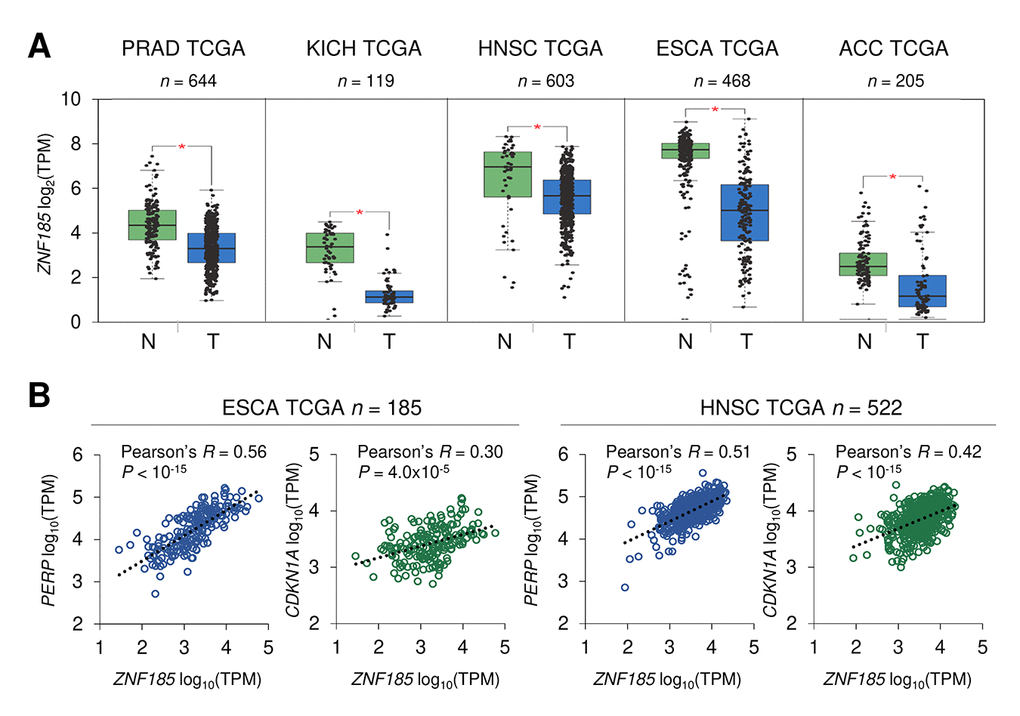
Figure 4. ZNF185 mRNA is down-regulated in epithelial cancer. (A) Box-plot showing expression of ZNF185 mRNA in different types of cancer from the GTEx/TCGA datasets for normal samples (N) and TCGA datasets for tumour samples (T). * P<0.05. (B) Correlation analysis between the expression of ZNF185 and either PERP or CDKN1A in ESCA and HNSC tumour samples from TCGA.
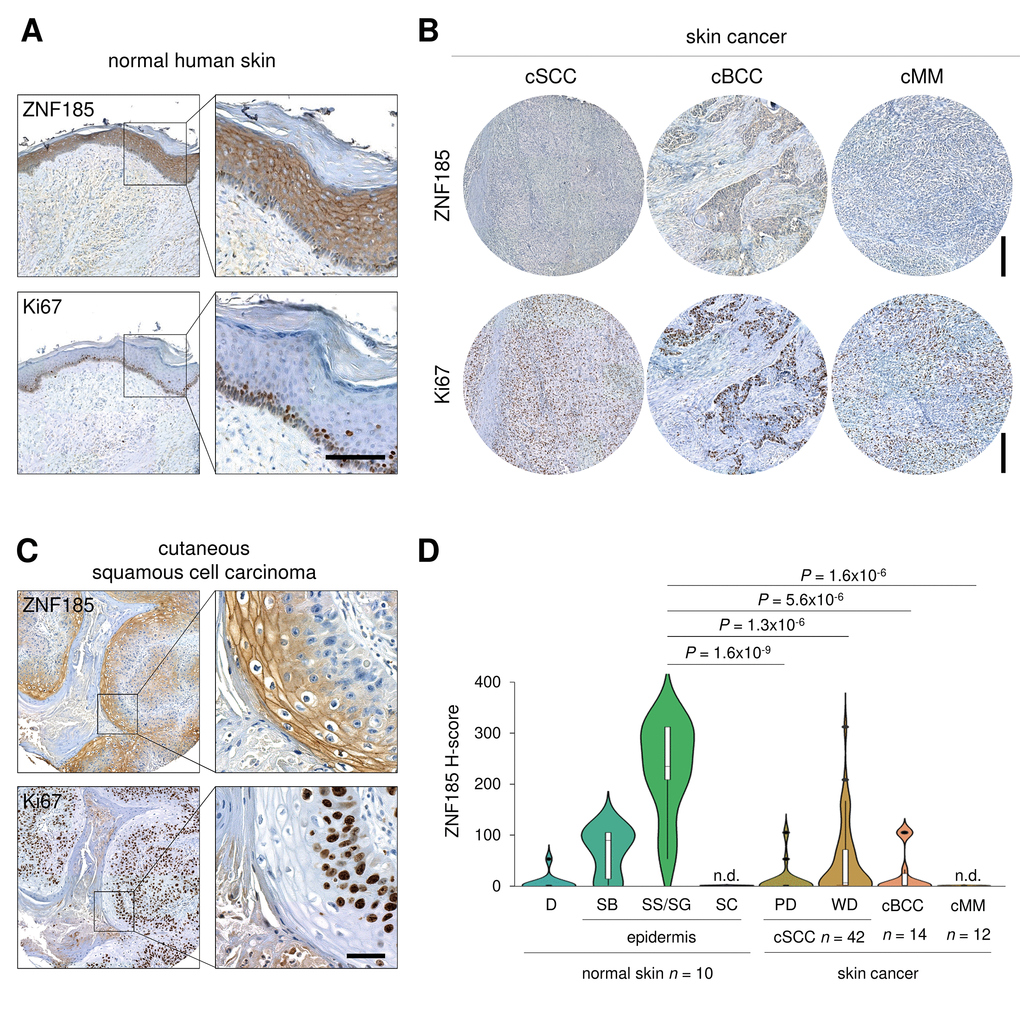
Figure 5. ZNF185 is down-regulated in skin cancer. (A) Immunohistochemical analysis of ZNF185 expression in normal skin. Ki67 was used as a marker of proliferation. Scale bar: 100 µm. (B) Immunohistochemical analysis of ZNF185 expression in skin cancer: cutaneous basal cell (cBCC), squamous cell carcinoma (cSCC), and malignant melanoma (cMM). Ki67 was used as a marker of proliferation. Scale bar: 250 µm. (C) Immunohistochemical analysis of ZNF185 expression in cutaneous squamous cell carcinoma (well differentiated). Ki67 was used as a marker of proliferation. Scale bar: 50 µm. (D) Violin plot showing H-score of protein expression level of ZNF185 in the normal skin (D – dermis, SB – basal layer, SS/SG – spinous and granular layers, SC – cornified layer) and skin cancer (cSCC (cSCC subpopulations: PD – poorly-differentiated cells, WD – well-differentiated cells), cBCC, and cMM).
Discussion
Aging is complex set of genetic [40,41], epigenetic [42–46], immunological [47–51], and metabolic [52–58] rearrangements, involving several cellular signalling pathways [59–63] able to regulate metabolism, ROS formation [64–71] and DNA Damage Response (DDR) [72–74] in all organs [75–77]. Therefore, the identification of novel pathways involving p53-mediated responses [78,79] is of crucial interest. TP53 has been extensively studied in development and differentiation [14,80–82] as well as in cancer progression [83,84] and specifically in DDR [85–87]. The situation involves additional complexity if we consider that also the p53-related family members [88,89], that is p63 [90–92] and p73 [93], may be involved in DDR. TP63 is a gene primarily related to skin development [94], metabolism [36,95,96] and epithelial homeostasis [97,98] however, there is compelling evidence that it is also involved in cancer [27,47,99–103]. Similarly, TP73 is clearly involved in cancer [104], while the knockout studies [105,106] indicate a clear involvement in metabolism [107–109], neuro-development [104,110,111] and cancer progression [112,113]. Therefore, while the potential involvement of p73 with ZNF185 remains to be elucidated, the identification of a novel pathways of p53 regulating DDR via ZNF185 is of relevance.
Recently, the importance of actin-cytoskeleton remodelling and cell polarity during cancer cell spreading and metastasis has emerged [114–117], and p53 is in part involved in counteracting this specific aspect. Indeed, wild-type p53 can influence actin cytoskeleton dynamics controlling integrin and cadherin signalling and extracellular matrix degradation, suppressing EMT via different pathways [20,23,118–120]. Interestingly, also tumour microenvironment influences actin cytoskeleton [121,122], in part repressing wild-type p53 functions [123]. In fact, when p53 is inactivated, cancer cells invasion increases [124]. Here, we demonstrated that p53 wild-type transcriptionally activates ZNF185 in cells upon DNA damage, which could make part of p53 negative regulation of cancer cell mobility, invasion and metastasis. Interestingly, similarly to another p53 target gene Rap2B [125], down-regulation of ZNF185 does not affect cell cycle progression or cell death, but its silencing abolishes the actin cytoskeleton rearrangements and cell polarity changes upon etoposide treatment.
ZNF185 is an actin-cytoskeleton-associated Lin-l 1, Isl-1 and Mec-3 (LIM) domain-containing protein [126]. The domain interacting with actin is located at the N-terminus, and it is necessary to mediate actin-cytoskeleton targeting of ZNF185, while the C-terminus LIM domain is dispensable for actin binding [38]. The LIM domain is a protein-protein interaction domain found in a wide range of proteins whose functions are related to the dynamics of the cytoskeleton [39,127,128]. In keratinocytes and epidermis ZNF185 has been described highly expressed in differentiating conditions, physically interacting with E-cadherin, a component of the adherens junctions, one of the critical cell-cell adhesive complexes crucial in the pluristratified epithelia [33]. ZNF185 involvement in pathologies, such as cancer [38], has not been completely investigated yet. Few studies reported ZNF185 as an unfavourable prognostic marker in ductal carcinoma of pancreas [129]. Its expression was found upregulated in colon cancer and likely correlated with liver metastasis [130]. On the other hand, other studies described epigenetic silencing of ZNF185 associated with high grade and metastatic prostate tumours [131], lung tumours and head and neck squamous cell carcinomas [33,132–134]. Recently, it was reported that ZNF185 expression is negatively correlated with lymph node metastasis of lung adenocarcinoma and its overexpression leads to down-regulation of p-AKT, p-GSK3β, VEGF and MMP-9 expression [135]. These studies suggest a possible tumour-specific contribution of ZNF185 expression in tumour formation.
We confirmed ZNF185 down-regulation in different epithelial tumours and, by analysing the expression of ZNF185 at protein level, we found a significant decrease of ZNF185 in all the tumour samples analysed. Moreover, we found ZNF185 positive signal only in well-differentiated subpopulations of squamous cell carcinoma in contrast to poorly-differentiated basal-like aggressive subpopulations, suggesting a tumour-suppressor role of ZNF185. The possible involvement of ZNF185 in cytoskeleton remodelling upon DNA damage suggests its role in the metastasis promotion which is in line with previous reports [135]. The identification of p53-ZNF185 axis could contribute to determine how p53 controls cell spreading by actin cytoskeletal remodelling, in which both the mechanical properties of the cytoskeleton of the cell as well as the microenvironment of the tumour cells seem to play an important role. Further investigation on the mechanisms by which p53 controls actin cytoskeleton reorganization and cell polarity, including the identification of novel target genes and pathways, would possibly be useful in developing new anti-cancer strategies and therapies.
Materials and Methods
Cell culture, transfection, and treatments
Immortalized human epidermal keratinocytes Ker-CT (ATCC, Manassas, VA, USA) were cultured in EpiLife medium with addition of Human Keratinocyte Growth Supplements (HKGS, LifeTechnologies, Carlsbad, CA, USA). Human Non-Small-Cell Lung Carcinoma cells H1299 (ATCC), Human Osteosarcoma cells SaOs-2 inducible for p53 expression (SaOs-2 Tet-on p53), Human Colorectal Carcinoma cells HCT116 (ATCC), and mammary gland adenocarcinoma cells MCF7 (ATCC) were grown in DMEM medium with the addition of 10% FBS, 100 U penicillin, and 100 μg/mL streptomycin (Gibco, LifeTechnologies). For siRNA-mediated knock-down experiments, 2.5x105 of cells were seeded on 60 mm culture dishes and the day after transfected with 80 pmol of specific siRNAs (Supplementary Table 1) by Lipofectamine RNAiMAX transfection reagent (Invitrogen, Carlsbad, CA, USA). Cells were collected 48h posttransfection. For overexpression experiments, 1x106 of cells were seeded in 100 mm culture dishes and the day after were transfected with 5 µg of plasmidic DNA using Lipofectamine 2000 transfection reagent (Invitrogen). Cells were collected 24 h later. For shRNA-mediated knock-down, 2.5x105 of Ker-CT were infected at m.o.i. 10 with lentiviral particles carrying either scramble control shRNA (NC, Cat. No. VSC7078, Dharmacon, Lafayette, CO, USA) or specific shRNA for ZNF185 (shZNF185, Cat. No. V3SVHSHC_5107270, Dharmacon). Transduced cells were selected by puromycin addiction to culture medium (1.5 µg/mL, Sigma, St. Louis, MO, USA). To induce DNA damage, cells were treated with either 25 µM or 100 µM etoposide (Sigma) or 0.02-0.40% DMSO (Sigma) for the indicated times or for 16 h if not otherwise indicated. Ker-CT cells were irradiated for indicated times with 10 mJ.cm-2 UV-B rays. p53 expression was induced in SaOs-2 Tet-On p53 cells by adding of 2 µg/mL doxycycline (Sigma) for the indicated times.
Western blotting
The cells were collected by trypsinization, washed in PBS and lysed in RIPA buffer (50 mM Tris-cl pH 7.4, 150 mM NaCl, 1% NP40, 0.25% Na-deoxycholate, 1 mM AEBSF, 1 mM DTT). 20-50 µg of total protein extracts were resolved in SDS polyacrylamide gel using the Mini-PROTEAN Tetra cell System (Bio-Rad, Hercules, CA, USA) and blotted onto a Hybond PVDF membrane (GE Healthcare, Chicago, IL, USA) using the Bio-Rad Mini Trans-Blot Cell system Bio-Rad). Membranes were blocked with 5% non-fat dry milk (Bio-Rad) in PBS/0.1% Tween-20 buffer, for 1 h at room temperature in agitation. Membranes were incubated with primary antibodies over night at +4 °C, washed and hybridized for 1 h at room temperature with the appropriate horseradish peroxidase-conjugated secondary antibodies (goat anti-rabbit and goat anti-mouse antibodies, Bio-Rad). Detection was performed with the ECL chemiluminescence kit (Perkin Elmer, Waltham, MA, USA). The following antibodies were used: anti-ZNF185 (1:300, Cat. No. HPA000400, Sigma), anti-GAPDH (1:15000, Cat. No. G8795, Sigma), anti-p53 (1:500, Cat. No. SC-126, Santa Cruz, Dallas, TX, USA), anti-p21 (1:300, Cat. No. SC-756, Santa Cruz), anti-HA (1:1000, Cat. No. 901502, BioLegend, San Diego, CA, USA).
RNA extraction and RT-qPCR analysis
Total RNA was isolated using the RNeasy Mini Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. Total RNA (1 µg) was used for cDNA synthesis by GoScript Reverse Transcription System kit (Promega, Madison, WI, USA). RT-qPCRs were performed using the GoTaq Real-Time PCR System (Promega) in Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) using appropriate qPCR primers (Supplementary Table 1). TBP was used as housekeeping gene for normalization. The expression of each gene was defined from the threshold cycle (Ct), and relative expression levels were calculated using the 2−ΔΔCt method. All reactions were run in triplicate.
Analysis of ZNF185 genomic locus
To analyse ZNF185 genomic locus, different publicity accessible high-throughput sequencing data from ENCODE database (ChIP seq for H3K4me3 in different cell lines, CpG islands, DNase clusters, vertebrate conservation, TF binding) were visualised in the UCSC Genome Browser. Several ChIP-seq data from NCBI GEO database were analysed to assess p53 binding to the ZNF185 promoter locus (GSE56674 ([37]) and GSE86164 ([34])). To identify putative p53 binding sites was used the “p53 scan” software [136]. The conservation analysis of ZNF185 promoter locus was performed within UCSC genome browser.
Chromatin immunoprecipitation assay
1x106 of SaOs Tet-On p53 cells, induced to overexpress p53 for 16 h, were used for ChIP assay. Cells were collected, fixed in 1% formaldehyde, and subjected to sonication for DNA shearing. The chromatin immunoprecipitation was performed with HA antibody (BioLegend) or unspecific immunoglobulin G (IgG, Invitrogen) using the MAGnify ChIP Kit (Invitrogen). Specific primers were used to amplify the putative p53 response element identified within ZNF185 promoter region (Supplementary Table 1).
Luciferase activity assay
Promoter region of ZNF185 containing the putative p53 binding site was amplified from human genomic DNA using specific primers (Supplementary Table 1). PCR products were digested by Kpn1/Nhe1 restriction enzymes (New England Biolabs, Ipswich, MA, USA) and subcloned into the pGL3-Promoter reporter vector (Promega). The constructs were completely sequenced. For luciferase activity assay, a total of 1.2x105 H1299 cells were seeded in 12-well dishes 24 h before transfection. 100 ng of pGL3 reporter vector, 2 ng of pRL-CMV-Renilla luciferase vector (Promega) and 300 ng of either pcDNA-HA-p53, pcDNA-HA-p53-R175H, pcDNA-HA-p53-R273H, or empty pcDNA-HA vector (as a control) were cotransfected using Effectene transfection reagent according to the manufacturer’s instructions (Qiagen). The luciferase activity was measured 24 h after transfection using a Dual Luciferase Reporter Assay System (Promega). The light emission was measured over 10 sec using a Lumat LB9507 luminometer (EG&GBerthold, Bad Wildbad, Germany). The transfection efficiency was normalized to Renilla luciferase activity. The overexpression of p53 was confirmed by western blotting.
Mutagenesis
For mutagenesis of p53 binding site was performed a PCR on 100 ng of pGL3 vector carrying p53 binding site using specific primers (Supplementary Table 1). PCR product was digested with DpnI restriction enzyme (New England Biolabs). The presence of mutated site was confirmed by sequencing.
Immunohistochemical staining and TMA
The immunohistochemical staining of the skin cancer tissue microarray sections (US Biomax, Rockville, MD, USA) was performed using the BenchMark ULTRA slide staining system (Roche Diagnostics, Risch-Rotkreuz, Switzerland). The staining for Ki67 was performed using anti-Ki67 antibody (Cat. No. 790-4286, Ventana) following manufacturer’s indications. For ZNF185 staining, samples were incubated at 95°C for 76 min in the Cell Conditioning solution CC1 (Roche) and stained with anti-ZNF185 antibody (1:100, Sigma) for 40 min. Sections were counterstained with Mayer’s haematoxylin, dehydrated and mounted. Samples were scored using a semi-quantitative method. Cases were analysed for staining intensity, which was scored as 0 (not detected), 1+ (weak), 2+ (intermediate), and 3+ (strong). For each case, the histological “H-score” (0-300) was calculated by multiplying the percentage of positive cells (0%-100%) by the intensity (0-3).
Immunofluorescence
Ker-CT cells were seeded on 5 mm coverslips, fixed for 10 min in 10% formalin buffered solution, washed with PBS and permeabilized in 0.2% triton X-100 solution in PBS for 10 min. Sections were incubated for 1 h in 10% goat serum in PBS at room temperature and overnight at 4 ºC with primary antibodies. Following antibodies were used: anti-ZNF185 (1:50, Sigma) and anti-vinculin (1:300, BD Biosciences, Franklin Lakes, NJ, USA). Sections were incubated for 1 h at room temperature with secondary anti-mouse and anti-rabbit 488- or 568-AlexaFluor conjugated antibodies (Invitrogen, 1:1000) together with 1 μg/mL DAPI (Sigma) for nuclear DNA staining. For cytoskeleton staining, 488- or 568-AlexaFluor conjugated phalloidin was used (1:1000, Thermo Fisher Scientific, Waltham, MA, USA). Images were acquired by Nikon A1 confocal laser microscope using NIS elements software (Nikon, Tokyo, Japan).
Cell proliferation
The incorporation of EdU during DNA synthesis was evaluated using the Click-iT EdU flow cytometry assay kit according to the manufacturer's protocol (Thermo Fisher Scientific). The cell cycle was analysed using an Accuri C6 flow cytometer (BD Biosciences). Fifteen thousand events were evaluated using the Accuri C6 (BD Biosciences) software. For cell cycle analysis, cells were fixed in 50% methanol/acetone 4:1 mix for 30 min at +4 °C, then treated with 13 Kunitz U/mL RNase for 15 min and stained with 50 µg/mL of propidium iodide for 20 min. Twelve thousand events were acquired using FACScalibur (BD Biosciences). Cell cycle distribution was calculated using FloxJo software.
Bioinformatic analysis
Analysis of ZNF185 expression in normal and tumour samples from TCGA/GTEx databases was performed using GEPIA [137]. ZNF185, PERP, and CDKN1A expression data in ESCA and HNSC samples from TCGA collection were obtained using R2: Genomics Analysis and Visualization Platform (http://r2.amc.nl/).
Statistical analysis
The significance of differences between two experimental groups was calculated using unpaired, two-tailed Student’s t-test. Values of P < 0.05 were considered significant. For RT-qPCR and luciferase assay, values reported are the mean ± SD. For statistical analysis of TMA scoring was used Mann–Whitney U test. All statistical analyses were performed using GraphPad Prism 7.0 Software. Violin plots were generated in R using ggplot2 package.
Supplementary Materials
Author Contributions
AS, AC, AML and LA performed the research, EC designed the research, EC, AM, NDD, MAP and GM analysed the data, EC wrote the paper and all the authors read the paper and made comments.
Conflicts of Interest
The authors declare no conflict of interest.
Funding
This work has been supported by grants from Associazione Italiana per la Ricerca contro il Cancro (AIRC): IG15653 (to G.M.), and by Istituto Dermopatico dell’Immacolata, IRCCS, RC to EC and joint research program Italia-Cina. This work has been also partially supported by the Medical Research Council, UK.
References
- 1. Harper JW, Elledge SJ. The DNA damage response: ten years after. Mol Cell. 2007; 28:739–45. https://doi.org/10.1016/j.molcel.2007.11.015 [PubMed]
- 2. Jackson SP, Bartek J. The DNA-damage response in human biology and disease. Nature. 2009; 461:1071–78. https://doi.org/10.1038/nature08467 [PubMed]
- 3. Houtgraaf JH, Versmissen J, van der Giessen WJ. A concise review of DNA damage checkpoints and repair in mammalian cells. Cardiovasc Revasc Med. 2006; 7:165–72. https://doi.org/10.1016/j.carrev.2006.02.002 [PubMed]
- 4. Ciccia A, Elledge SJ. The DNA damage response: making it safe to play with knives. Mol Cell. 2010; 40:179–204. https://doi.org/10.1016/j.molcel.2010.09.019 [PubMed]
- 5. Hawley BR, Lu WT, Wilczynska A, Bushell M. The emerging role of RNAs in DNA damage repair. Cell Death Differ. 2017; 24:1989. https://doi.org/10.1038/cdd.2017.146 [PubMed]
- 6. Baran K, Yang M, Dillon CP, Samson LL, Green DR. The proline rich domain of p53 is dispensable for MGMT-dependent DNA repair and cell survival following alkylation damage. Cell Death Differ. 2017; 24:1925–36. https://doi.org/10.1038/cdd.2017.116 [PubMed]
- 7. Delbridge AR, Grabow S, Strasser A. Loss of BIM augments resistance of ATM-deficient thymocytes to DNA damage-induced apoptosis but does not accelerate lymphoma development. Cell Death Differ. 2017; 24:1987–88. https://doi.org/10.1038/cdd.2017.138 [PubMed]
- 8. Zhou BB, Elledge SJ. The DNA damage response: putting checkpoints in perspective. Nature. 2000; 408:433–39. https://doi.org/10.1038/35044005 [PubMed]
- 9. Lakin ND, Jackson SP. Regulation of p53 in response to DNA damage. Oncogene. 1999; 18:7644–55. https://doi.org/10.1038/sj.onc.1203015 [PubMed]
- 10. Kruiswijk F, Labuschagne CF, Vousden KH. p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol. 2015; 16:393–405. https://doi.org/10.1038/nrm4007 [PubMed]
- 11. Kubbutat MH, Jones SN, Vousden KH. Regulation of p53 stability by Mdm2. Nature. 1997; 387:299–303. https://doi.org/10.1038/387299a0 [PubMed]
- 12. Gurley KE, Ashley AK, Moser RD, Kemp CJ. Synergy between Prkdc and Trp53 regulates stem cell proliferation and GI-ARS after irradiation. Cell Death Differ. 2017; 24:1853–60. https://doi.org/10.1038/cdd.2017.107 [PubMed]
- 13. Nayak D, Kumar A, Chakraborty S, Rasool RU, Amin H, Katoch A, Gopinath V, Mahajan V, Zilla MK, Rah B, Gandhi SG, Ali A, Kumar LD, Goswami A. Inhibition of Twist1-mediated invasion by Chk2 promotes premature senescence in p53-defective cancer cells. Cell Death Differ. 2017; 24:1275–87. https://doi.org/10.1038/cdd.2017.70 [PubMed]
- 14. Van Nostrand JL, Bowen ME, Vogel H, Barna M, Attardi LD. The p53 family members have distinct roles during mammalian embryonic development. Cell Death Differ. 2017; 24:575–79. https://doi.org/10.1038/cdd.2016.128 [PubMed]
- 15. Solomon H, Bräuning B, Fainer I, Ben-Nissan G, Rabani S, Goldfinger N, Moscovitz O, Shakked Z, Rotter V, Sharon M. Post-translational regulation of p53 function through 20S proteasome-mediated cleavage. Cell Death Differ. 2017; 24:2187–98. https://doi.org/10.1038/cdd.2017.139 [PubMed]
- 16. Riley T, Sontag E, Chen P, Levine A. Transcriptional control of human p53-regulated genes. Nat Rev Mol Cell Biol. 2008; 9:402–12. https://doi.org/10.1038/nrm2395 [PubMed]
- 17. Ford JM. Regulation of DNA damage recognition and nucleotide excision repair: another role for p53. Mutat Res. 2005; 577:195–202. https://doi.org/10.1016/j.mrfmmm.2005.04.005 [PubMed]
- 18. Ebata T, Hirata H, Kawauchi K. Functions of the Tumor Suppressors p53 and Rb in Actin Cytoskeleton Remodeling. BioMed Res Int. 2016; 2016:9231057. https://doi.org/10.1155/2016/9231057 [PubMed]
- 19. Roger L, Gadea G, Roux P. Control of cell migration: a tumour suppressor function for p53? Biol Cell. 2006; 98:141–52. https://doi.org/10.1042/BC20050058 [PubMed]
- 20. Araki K, Ebata T, Guo AK, Tobiume K, Wolf SJ, Kawauchi K. p53 regulates cytoskeleton remodeling to suppress tumor progression. Cell Mol Life Sci. 2015; 72:4077–94. https://doi.org/10.1007/s00018-015-1989-9 [PubMed]
- 21. Janouskova H, Maglott A, Leger DY, Bossert C, Noulet F, Guerin E, Guenot D, Pinel S, Chastagner P, Plenat F, Entz-Werle N, Lehmann-Che J, Godet J, et al. Integrin α5β1 plays a critical role in resistance to temozolomide by interfering with the p53 pathway in high-grade glioma. Cancer Res. 2012; 72:3463–70. https://doi.org/10.1158/0008-5472.CAN-11-4199 [PubMed]
- 22. Janouskova H, Ray AM, Noulet F, Lelong-Rebel I, Choulier L, Schaffner F, Lehmann M, Martin S, Teisinger J, Dontenwill M. Activation of p53 pathway by Nutlin-3a inhibits the expression of the therapeutic target α5 integrin in colon cancer cells. Cancer Lett. 2013; 336:307–18. https://doi.org/10.1016/j.canlet.2013.03.018 [PubMed]
- 23. Bon G, Di Carlo SE, Folgiero V, Avetrani P, Lazzari C, D’Orazi G, Brizzi MF, Sacchi A, Soddu S, Blandino G, Mottolese M, Falcioni R. Negative regulation of beta4 integrin transcription by homeodomain-interacting protein kinase 2 and p53 impairs tumor progression. Cancer Res. 2009; 69:5978–86. https://doi.org/10.1158/0008-5472.CAN-09-0244 [PubMed]
- 24. Muller PA, Vousden KH, Norman JC. p53 and its mutants in tumor cell migration and invasion. J Cell Biol. 2011; 192:209–18. https://doi.org/10.1083/jcb.201009059 [PubMed]
- 25. Gadea G, de Toledo M, Anguille C, Roux P. Loss of p53 promotes RhoA-ROCK-dependent cell migration and invasion in 3D matrices. J Cell Biol. 2007; 178:23–30. https://doi.org/10.1083/jcb.200701120 [PubMed]
- 26. Lefort K, Mandinova A, Ostano P, Kolev V, Calpini V, Kolfschoten I, Devgan V, Lieb J, Raffoul W, Hohl D, Neel V, Garlick J, Chiorino G, Dotto GP. Notch1 is a p53 target gene involved in human keratinocyte tumor suppression through negative regulation of ROCK1/2 and MRCKalpha kinases. Genes Dev. 2007; 21:562–77. https://doi.org/10.1101/gad.1484707 [PubMed]
- 27. Guo F, Gao Y, Wang L, Zheng Y. p19Arf-p53 tumor suppressor pathway regulates cell motility by suppression of phosphoinositide 3-kinase and Rac1 GTPase activities. J Biol Chem. 2003; 278:14414–19. https://doi.org/10.1074/jbc.M300341200 [PubMed]
- 28. Gadéa G, Lapasset L, Gauthier-Rouvière C, Roux P. Regulation of Cdc42-mediated morphological effects: a novel function for p53. EMBO J. 2002; 21:2373–82. https://doi.org/10.1093/emboj/21.10.2373 [PubMed]
- 29. Croft DR, Crighton D, Samuel MS, Lourenco FC, Munro J, Wood J, Bensaad K, Vousden KH, Sansom OJ, Ryan KM, Olson MF. p53-mediated transcriptional regulation and activation of the actin cytoskeleton regulatory RhoC to LIMK2 signaling pathway promotes cell survival. Cell Res. 2011; 21:666–82. https://doi.org/10.1038/cr.2010.154 [PubMed]
- 30. Ongusaha PP, Kim HG, Boswell SA, Ridley AJ, Der CJ, Dotto GP, Kim YB, Aaronson SA, Lee SW. RhoE is a pro-survival p53 target gene that inhibits ROCK I-mediated apoptosis in response to genotoxic stress. Curr Biol. 2006; 16:2466–72. https://doi.org/10.1016/j.cub.2006.10.056 [PubMed]
- 31. Hall AE, Lu WT, Godfrey JD, Antonov AV, Paicu C, Moxon S, Dalmay T, Wilczynska A, Muller PA, Bushell M. The cytoskeleton adaptor protein ankyrin-1 is upregulated by p53 following DNA damage and alters cell migration. Cell Death Dis. 2016; 7:e2184. https://doi.org/10.1038/cddis.2016.91 [PubMed]
- 32. Muller PA, Caswell PT, Doyle B, Iwanicki MP, Tan EH, Karim S, Lukashchuk N, Gillespie DA, Ludwig RL, Gosselin P, Cromer A, Brugge JS, Sansom OJ, et al. Mutant p53 drives invasion by promoting integrin recycling. Cell. 2009; 139:1327–41. https://doi.org/10.1016/j.cell.2009.11.026 [PubMed]
- 33. Smirnov A, Lena AM, Cappello A, Panatta E, Anemona L, Bischetti S, Annicchiarico-Petruzzelli M, Mauriello A, Melino G, Candi E. ZNF185 is a p63 target gene critical for epidermal differentiation and squamous cell carcinoma development. Oncogene. Epub ahead of print. https://doi.org/10.1038/s41388-018-0509-4 [PubMed]
- 34. Andrysik Z, Galbraith MD, Guarnieri AL, Zaccara S, Sullivan KD, Pandey A, MacBeth M, Inga A, Espinosa JM. Identification of a core TP53 transcriptional program with highly distributed tumor suppressive activity. Genome Res. 2017; 27:1645–57. https://doi.org/10.1101/gr.220533.117 [PubMed]
- 35. Kouwenhoven EN, van Heeringen SJ, Tena JJ, Oti M, Dutilh BE, Alonso ME, de la Calle-Mustienes E, Smeenk L, Rinne T, Parsaulian L, Bolat E, Jurgelenaite R, Huynen MA, et al. Genome-wide profiling of p63 DNA-binding sites identifies an element that regulates gene expression during limb development in the 7q21 SHFM1 locus. PLoS Genet. 2010; 6:e1001065. https://doi.org/10.1371/journal.pgen.1001065 [PubMed]
- 36. Gressner O, Schilling T, Lorenz K, Schulze Schleithoff E, Koch A, Schulze-Bergkamen H, Lena AM, Candi E, Terrinoni A, Catani MV, Oren M, Melino G, Krammer PH, et al. TAp63α induces apoptosis by activating signaling via death receptors and mitochondria. EMBO J. 2005; 24:2458–71. https://doi.org/10.1038/sj.emboj.7600708 [PubMed]
- 37. McDade SS, Patel D, Moran M, Campbell J, Fenwick K, Kozarewa I, Orr NJ, Lord CJ, Ashworth AA, McCance DJ. Genome-wide characterization reveals complex interplay between TP53 and TP63 in response to genotoxic stress. Nucleic Acids Res. 2014; 42:6270–85. https://doi.org/10.1093/nar/gku299 [PubMed]
- 38. Zhang JS, Gong A, Young CY. ZNF185, an actin-cytoskeleton-associated growth inhibitory LIM protein in prostate cancer. Oncogene. 2007; 26:111–22. https://doi.org/10.1038/sj.onc.1209769 [PubMed]
- 39. Kadrmas JL, Beckerle MC. The LIM domain: from the cytoskeleton to the nucleus. Nat Rev Mol Cell Biol. 2004; 5:920–31. https://doi.org/10.1038/nrm1499 [PubMed]
- 40. Stoll EA, Karapavlovic N, Rosa H, Woodmass M, Rygiel K, White K, Turnbull DM, Faulkes CG. Naked mole-rats maintain healthy skeletal muscle and Complex IV mitochondrial enzyme function into old age. Aging (Albany NY). 2016; 8:3468–85. https://doi.org/10.18632/aging.101140 [PubMed]
- 41. Ticinesi A, Tana C, Nouvenne A, Prati B, Lauretani F, Meschi T. Gut microbiota, cognitive frailty and dementia in older individuals: a systematic review. Clin Interv Aging. 2018; 13:1497–511. https://doi.org/10.2147/CIA.S139163 [PubMed]
- 42. Kaestner L, Minetti G. The potential of erythrocytes as cellular aging models. Cell Death Differ. 2017; 24:1475–77. https://doi.org/10.1038/cdd.2017.100 [PubMed]
- 43. Kuro-O M. Molecular Mechanisms Underlying Accelerated Aging by Defects in the FGF23-Klotho System. Int J Nephrol. 2018; 2018:9679841. https://doi.org/10.1155/2018/9679841 [PubMed]
- 44. Pollard AK, Ortori CA, Stöger R, Barrett DA, Chakrabarti L. Mouse mitochondrial lipid composition is defined by age in brain and muscle. Aging (Albany NY). 2017; 9:986–98. https://doi.org/10.18632/aging.101204 [PubMed]
- 45. Regina C, Compagnone M, Peschiaroli A, Lena AM, Melino G, Candi E. ΔNp63α modulates histone methyl transferase SETDB1 to transcriptionally repress target genes in cancers. Cell Death Dis. 2016; 2:16015. https://doi.org/10.1038/cddiscovery.2016.15 [PubMed]
- 46. Smirnov A, Panatta E, Lena A, Castiglia D, Di Daniele N, Melino G, Candi E. FOXM1 regulates proliferation, senescence and oxidative stress in keratinocytes and cancer cells. Aging (Albany NY). 2016; 8:1384–97. https://doi.org/10.18632/aging.100988 [PubMed]
- 47. Xu-Monette ZY, Zhang S, Li X, Manyam GC, Wang XX, Xia Y, Visco C, Tzankov A, Zhang L, Montes-Moreno S, Dybkaer K, Chiu A, Orazi A, et al. p63 expression confers significantly better survival outcomes in high-risk diffuse large B-cell lymphoma and demonstrates p53-like and p53-independent tumor suppressor function. Aging (Albany NY). 2016; 8:345–65. https://doi.org/10.18632/aging.100898 [PubMed]
- 48. Lareau CA, DeWeese CF, Adrianto I, Lessard CJ, Gaffney PM, Iannuzzi MC, Rybicki BA, Levin AM, Montgomery CG. Polygenic risk assessment reveals pleiotropy between sarcoidosis and inflammatory disorders in the context of genetic ancestry. Genes Immun. 2017; 18:88–94. https://doi.org/10.1038/gene.2017.3 [PubMed]
- 49. Messemaker TC, Mikkers HM, Huizinga TW, Toes RE, van der Helm-van Mil AH, Kurreeman F. Inflammatory genes TNFα and IL6 display no signs of increased H3K4me3 in circulating monocytes from untreated rheumatoid arthritis patients. Genes Immun. 2017; 18:191–96. https://doi.org/10.1038/gene.2017.20 [PubMed]
- 50. Nikolic T, Woittiez NJ, van der Slik A, Laban S, Joosten A, Gysemans C, Mathieu C, Zwaginga JJ, Koeleman B, Roep BO. Differential transcriptome of tolerogenic versus inflammatory dendritic cells points to modulated T1D genetic risk and enriched immune regulation. Genes Immun. 2017; 18:176–83. https://doi.org/10.1038/gene.2017.18 [PubMed]
- 51. Liu S, Hou XL, Sui WG, Lu QJ, Hu YL, Dai Y. Direct measurement of B-cell receptor repertoire’s composition and variation in systemic lupus erythematosus. Genes Immun. 2017; 18:22–27. https://doi.org/10.1038/gene.2016.45 [PubMed]
- 52. Uzhachenko R, Boyd K, Olivares-Villagomez D, Zhu Y, Goodwin JS, Rana T, Shanker A, Tan WJ, Bondar T, Medzhitov R, Ivanova AV. Mitochondrial protein Fus1/Tusc2 in premature aging and age-related pathologies: critical roles of calcium and energy homeostasis. Aging (Albany NY). 2017; 9:627–49. https://doi.org/10.18632/aging.101213 [PubMed]
- 53. Baraibar MA, Hyzewicz J, Rogowska-Wrzesinska A, Bulteau AL, Prip-Buus C, Butler-Browne G, Friguet B. Impaired energy metabolism of senescent muscle satellite cells is associated with oxidative modifications of glycolytic enzymes. Aging (Albany NY). 2016; 8:3375–89. https://doi.org/10.18632/aging.101126 [PubMed]
- 54. Honrath B, Matschke L, Meyer T, Magerhans L, Perocchi F, Ganjam GK, Zischka H, Krasel C, Gerding A, Bakker BM, Bünemann M, Strack S, Decher N, et al. SK2 channels regulate mitochondrial respiration and mitochondrial Ca2+ uptake. Cell Death Differ. 2017; 24:761–73. https://doi.org/10.1038/cdd.2017.2 [PubMed]
- 55. Ingram T, Chakrabarti L. Proteomic profiling of mitochondria: what does it tell us about the ageing brain? Aging (Albany NY). 2016; 8:3161–79. https://doi.org/10.18632/aging.101131 [PubMed]
- 56. Kagan VE, Jiang J, Huang Z, Tyurina YY, Desbourdes C, Cottet-Rousselle C, Dar HH, Verma M, Tyurin VA, Kapralov AA, Cheikhi A, Mao G, Stolz D, et al. NDPK-D (NM23-H4)-mediated externalization of cardiolipin enables elimination of depolarized mitochondria by mitophagy. Cell Death Differ. 2016; 23:1140–51. https://doi.org/10.1038/cdd.2015.160 [PubMed]
- 57. Kaufman DM, Wu X, Scott BA, Itani OA, Van Gilst MR, Bruce JE, Crowder CM. Ageing and hypoxia cause protein aggregation in mitochondria. Cell Death Differ. 2017; 24:1730–38. https://doi.org/10.1038/cdd.2017.101 [PubMed]
- 58. Giacomello M, Pellegrini L. The coming of age of the mitochondria-ER contact: a matter of thickness. Cell Death Differ. 2016; 23:1417–27. https://doi.org/10.1038/cdd.2016.52 [PubMed]
- 59. Choi JY, Hwang CY, Lee B, Lee SM, Bahn YJ, Lee KP, Kang M, Kim YS, Woo SH, Lim JY, Kim E, Kwon KS. Age-associated repression of type 1 inositol 1, 4, 5-triphosphate receptor impairs muscle regeneration. Aging (Albany NY). 2016; 8:2062–80. https://doi.org/10.18632/aging.101039 [PubMed]
- 60. Jęśko H, Stępień A, Lukiw WJ, Strosznajder RP. The Cross-Talk Between Sphingolipids and Insulin-Like Growth Factor Signaling: Significance for Aging and Neurodegeneration. Mol Neurobiol. 2018. https://doi.org/10.1007/s12035-018-1286-3 [PubMed]
- 61. Lee SJ, Hwang J, Jeong HJ, Yoo M, Go GY, Lee JR, Leem YE, Park JW, Seo DW, Kim YK, Hahn MJ, Han JW, Kang JS, Bae GU. PKN2 and Cdo interact to activate AKT and promote myoblast differentiation. Cell Death Dis. 2016; 7:e2431. https://doi.org/10.1038/cddis.2016.296 [PubMed]
- 62. Oh J, Sinha I, Tan KY, Rosner B, Dreyfuss JM, Gjata O, Tran P, Shoelson SE, Wagers AJ. Age-associated NF-κB signaling in myofibers alters the satellite cell niche and re-strains muscle stem cell function. Aging (Albany NY). 2016; 8:2871–96. https://doi.org/10.18632/aging.101098 [PubMed]
- 63. Shao AW, Sun H, Geng Y, Peng Q, Wang P, Chen J, Xiong T, Cao R, Tang J. Bclaf1 is an important NF-κB signaling transducer and C/EBPβ regulator in DNA damage-induced senescence. Cell Death Differ. 2016; 23:865–75. https://doi.org/10.1038/cdd.2015.150 [PubMed]
- 64. Knupp J, Martinez-Montañés F, Van Den Bergh F, Cottier S, Schneiter R, Beard D, Chang A. Sphingolipid accumulation causes mitochondrial dysregulation and cell death. Cell Death Differ. 2017; 24:2044–53. https://doi.org/10.1038/cdd.2017.128 [PubMed]
- 65. Lalia AZ, Dasari S, Robinson MM, Abid H, Morse DM, Klaus KA, Lanza IR. Influence of omega-3 fatty acids on skeletal muscle protein metabolism and mitochondrial bioenergetics in older adults. Aging (Albany NY). 2017; 9:1096–129. https://doi.org/10.18632/aging.101210 [PubMed]
- 66. Lang A, Anand R, Altinoluk-Hambüchen S, Ezzahoini H, Stefanski A, Iram A, Bergmann L, Urbach J, Böhler P, Hänsel J, Franke M, Stühler K, Krutmann J, et al. SIRT4 interacts with OPA1 and regulates mitochondrial quality control and mitophagy. Aging (Albany NY). 2017; 9:2163–89. https://doi.org/10.18632/aging.101307 [PubMed]
- 67. Marini A, Rotblat B, Sbarrato T Niklison-Chirou MV, Knight JR, Dudek K, Jones C, Bushell M, Knight RA, Amelio I, Willis AE, Melino G. TAp73 contributes to the oxidative stress response by regulating protein synthesis. Proc Natl Acad Sci USA. 2018; 115:6219–24. https://doi.org/10.1073/pnas.1718531115 [PubMed]
- 68. Pinto M, Pickrell AM, Wang X, Bacman SR, Yu A, Hida A, Dillon LM, Morton PD, Malek TR, Williams SL, Moraes CT. Transient mitochondrial DNA double strand breaks in mice cause accelerated aging phenotypes in a ROS-dependent but p53/p21-independent manner. Cell Death Differ. 2017; 24:288–99. https://doi.org/10.1038/cdd.2016.123 [PubMed]
- 69. Qi Y, Liu H, Daniels MP, Zhang G, Xu H. Loss of Drosophila i-AAA protease, dYME1L, causes abnormal mitochondria and apoptotic degeneration. Cell Death Differ. 2016; 23:291–302. https://doi.org/10.1038/cdd.2015.94 [PubMed]
- 70. Ohkusu-Tsukada K, Yamashita T, Tsukada T, Takahashi K. Low expression of a Ddm7/Ldm7-hybrid mutant (D/Ldm7) in the novel haplotype H-2nc identified in atopic dermatitis model NC/Nga mice. Genes Immun. 2017. https://doi.org/10.1038/s41435-017-0003-y [PubMed]
- 71. Tak H, Eun JW, Kim J, Park SJ, Kim C, Ji E, Lee H, Kang H, Cho DH, Lee K, Kim W, Nam SW, Lee EK. T-cell-restricted intracellular antigen 1 facilitates mitochondrial fragmentation by enhancing the expression of mitochondrial fission factor. Cell Death Differ. 2017; 24:49–58. https://doi.org/10.1038/cdd.2016.90 [PubMed]
- 72. Galluzzi L, Vitale I, Aaronson SA, Abrams JM, Adam D, Agostinis P, Alnemri ES, Altucci L, Amelio I, Andrews DW, Annicchiarico-Petruzzelli M, Antonov AV, Arama E, et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 2018; 25:486–541. https://doi.org/10.1038/s41418-017-0012-4 [PubMed]
- 73. Gross A, Zaltsman Y, Maryanovich M. The ATM-BID pathway plays a critical role in the DNA damage response by regulating mitochondria metabolism. Cell Death Differ. 2016; 23:182. https://doi.org/10.1038/cdd.2015.154 [PubMed]
- 74. Lovat PE, Ranalli M, Annichiarrico-Petruzzelli M, Bernassola F, Piacentini M, Malcolm AJ, Pearson AD, Melino G, Redfern CP. Effector mechanisms of fenretinide-induced apoptosis in neuroblastoma. Exp Cell Res. 2000; 260:50–60. https://doi.org/10.1006/excr.2000.4988 [PubMed]
- 75. Belle JI, Petrov JC, Langlais D, Robert F, Cencic R, Shen S, Pelletier J, Gros P, Nijnik A. Repression of p53-target gene Bbc3/PUMA by MYSM1 is essential for the survival of hematopoietic multipotent progenitors and contributes to stem cell maintenance. Cell Death Differ. 2016; 23:759–75. https://doi.org/10.1038/cdd.2015.140 [PubMed]
- 76. Brzeszczyńska J, Johns N, Schilb A, Degen S, Degen M, Langen R, Schols A, Glass DJ, Roubenoff R, Greig CA, Jacobi C, Fearon KC, Ross JA. Loss of oxidative defense and potential blockade of satellite cell maturation in the skeletal muscle of patients with cancer but not in the healthy elderly. Aging (Albany NY). 2016; 8:1690–702. https://doi.org/10.18632/aging.101006 [PubMed]
- 77. Greschik H, Duteil D, Messaddeq N, Willmann D, Arrigoni L, Sum M, Jung M, Metzger D, Manke T, Günther T, Schüle R. The histone code reader Spin1 controls skeletal muscle development. Cell Death Dis. 2017; 8:e3173. https://doi.org/10.1038/cddis.2017.468 [PubMed]
- 78. Aggarwal M, Saxena R, Sinclair E, Fu Y, Jacobs A, Dyba M, Wang X, Cruz I, Berry D, Kallakury B, Mueller SC, Agostino SD, Blandino G, et al. Reactivation of mutant p53 by a dietary-related compound phenethyl isothiocyanate inhibits tumor growth. Cell Death Differ. 2016; 23:1615–27. https://doi.org/10.1038/cdd.2016.48 [PubMed]
- 79. Alexandrova EM, Moll UM. Depleting stabilized GOF mutant p53 proteins by inhibiting molecular folding chaperones: a new promise in cancer therapy. Cell Death Differ. 2017; 24:3–5. https://doi.org/10.1038/cdd.2016.145 [PubMed]
- 80. Artigas N, Gámez B, Cubillos-Rojas M, Sánchez-de Diego C, Valer JA, Pons G, Rosa JL, Ventura F. p53 inhibits SP7/Osterix activity in the transcriptional program of osteoblast differentiation. Cell Death Differ. 2017; 24:2022–31. https://doi.org/10.1038/cdd.2017.113 [PubMed]
- 81. Baldelli S, Ciriolo MR. Altered S-nitrosylation of p53 is responsible for impaired antioxidant response in skeletal muscle during aging. Aging (Albany NY). 2016; 8:3450–67. https://doi.org/10.18632/aging.101139 [PubMed]
- 82. Cam H, Griesmann H, Beitzinger M, Hofmann L, Beinoraviciute-Kellner R, Sauer M, Hüttinger-Kirchhof N, Oswald C, Friedl P, Gattenlöhner S, Burek C, Rosenwald A, Stiewe T. p53 family members in myogenic differentiation and rhabdomyosarcoma development. Cancer Cell. 2006; 10:281–93. https://doi.org/10.1016/j.ccr.2006.08.024 [PubMed]
- 83. Ghosh S, Salot S, Sengupta S, Navalkar A, Ghosh D, Jacob R, Das S, Kumar R, Jha NN, Sahay S, Mehra S, Mohite GM, Ghosh SK, et al. p53 amyloid formation leading to its loss of function: implications in cancer pathogenesis. Cell Death Differ. 2017; 24:1784–98. https://doi.org/10.1038/cdd.2017.105 [PubMed]
- 84. Nicolai S, Rossi A, Di Daniele N, Melino G, Annicchiarico-Petruzzelli M, Raschellà G. DNA repair and aging: the impact of the p53 family. Aging (Albany NY). 2015; 7:1050–65. https://doi.org/10.18632/aging.100858 [PubMed]
- 85. Seitz SJ, Schleithoff ES, Koch A, Schuster A, Teufel A, Staib F, Stremmel W, Melino G, Krammer PH, Schilling T, Müller M. Chemotherapy-induced apoptosis in hepatocellular carcinoma involves the p53 family and is mediated via the extrinsic and the intrinsic pathway. Int J Cancer. 2010; 126:2049–66. [PubMed]
- 86. Charni M, Aloni-Grinstein R, Molchadsky A, Rotter V. p53 on the crossroad between regeneration and cancer. Cell Death Differ. 2017; 24:8–14. https://doi.org/10.1038/cdd.2016.117 [PubMed]
- 87. Amelio I, Knight RA, Lisitsa A, Melino G, Antonov AV. p53MutaGene: an online tool to estimate the effect of p53 mutational status on gene regulation in cancer. Cell Death Dis. 2016; 7:e2148. https://doi.org/10.1038/cddis.2016.42 [PubMed]
- 88. Levine AJ, Tomasini R, McKeon FD, Mak TW, Melino G. The p53 family: guardians of maternal reproduction. Nat Rev Mol Cell Biol. 2011; 12:259–65. https://doi.org/10.1038/nrm3086 [PubMed]
- 89. De Laurenzi V, Melino G. Evolution of functions within the p53/p63/p73 family. Ann N Y Acad Sci. 2000; 926:90–100. https://doi.org/10.1111/j.1749-6632.2000.tb05602.x [PubMed]
- 90. Memmi EM, Sanarico AG, Giacobbe A, Peschiaroli A, Frezza V, Cicalese A, Pisati F, Tosoni D, Zhou H, Tonon G, Antonov A, Melino G, Pelicci PG, Bernassola F. p63 Sustains self-renewal of mammary cancer stem cells through regulation of Sonic Hedgehog signaling. Proc Natl Acad Sci USA. 2015; 112:3499–504. https://doi.org/10.1073/pnas.1500762112 [PubMed]
- 91. Novelli F, Lena AM, Panatta E, Nasser W, Shalom-Feuerstein R, Candi E, Melino G. Allele-specific silencing of EEC p63 mutant R304W restores p63 transcriptional activity. Cell Death Dis. 2016; 7:e2227. https://doi.org/10.1038/cddis.2016.118 [PubMed]
- 92. Rivetti di Val Cervo P, Lena AM, Nicoloso M, Rossi S, Mancini M, Zhou H, Saintigny G, Dellambra E, Odorisio T, Mahé C, Calin GA, Candi E, Melino G. p63-microRNA feedback in keratinocyte senescence. Proc Natl Acad Sci USA. 2012; 109:1133–38. https://doi.org/10.1073/pnas.1112257109 [PubMed]
- 93. Melino G, De Laurenzi V, Vousden KH. p73: Friend or foe in tumorigenesis. Nat Rev Cancer. 2002; 2:605–15. https://doi.org/10.1038/nrc861 [PubMed]
- 94. Candi E, Schmidt R, Melino G. The cornified envelope: a model of cell death in the skin. Nat Rev Mol Cell Biol. 2005; 6:328–40. https://doi.org/10.1038/nrm1619 [PubMed]
- 95. Viticchiè G, Agostini M, Lena AM, Mancini M, Zhou H, Zolla L, Dinsdale D, Saintigny G, Melino G, Candi E. p63 supports aerobic respiration through hexokinase II. Proc Natl Acad Sci USA. 2015; 112:11577–82. https://doi.org/10.1073/pnas.1508871112 [PubMed]
- 96. Candi E, Smirnov A, Panatta E, Lena AM, Novelli F, Mancini M, Viticchiè G, Piro MC, Di Daniele N, Annicchiarico-Petruzzelli M, Melino G. Metabolic pathways regulated by p63. Biochem Biophys Res Commun. 2017; 482:440–44. https://doi.org/10.1016/j.bbrc.2016.10.094 [PubMed]
- 97. Candi E, Terrinoni A, Rufini A, Chikh A, Lena AM, Suzuki Y, Sayan BS, Knight RA, Melino G. p63 is upstream of IKK alpha in epidermal development. J Cell Sci. 2006; 119:4617–22. . http://jcs.biologists.org/cgi/doi/10.1242/jcs.03265 https://doi.org/10.1242/jcs.03265 [PubMed]
- 98. Martin SE, Temm CJ, Goheen MP, Ulbright TM, Hattab EM. Cytoplasmic p63 immunohistochemistry is a useful marker for muscle differentiation: an immunohistochemical and immunoelectron microscopic study. Mod Pathol. 2011; 24:1320–26. https://doi.org/10.1038/modpathol.2011.89 [PubMed]
- 99. Compagnone M, Gatti V, Presutti D, Ruberti G, Fierro C, Markert EK, Vousden KH, Zhou H, Mauriello A, Anemone L, Bongiorno-Borbone L, Melino G, Peschiaroli A. ΔNp63-mediated regulation of hyaluronic acid metabolism and signaling supports HNSCC tumorigenesis. Proc Natl Acad Sci USA. 2017; 114:13254–59. https://doi.org/10.1073/pnas.1711777114 [PubMed]
- 100. Di Franco S, Sala G, Todaro M. p63 role in breast cancer. Aging (Albany NY). 2016; 8:2256–57. https://doi.org/10.18632/aging.101042 [PubMed]
- 101. Flores ER, Lozano G. The p53 family grows old. Genes Dev. 2012; 26:1997–2000. https://doi.org/10.1101/gad.202648.112 [PubMed]
- 102. Flores ER, Sengupta S, Miller JB, Newman JJ, Bronson R, Crowley D, Yang A, McKeon F, Jacks T. Tumor predisposition in mice mutant for p63 and p73: evidence for broader tumor suppressor functions for the p53 family. Cancer Cell. 2005; 7:363–73. https://doi.org/10.1016/j.ccr.2005.02.019 [PubMed]
- 103. Latina A, Viticchiè G, Lena AM, Piro MC, Annicchiarico-Petruzzelli M, Melino G, Candi E. ΔNp63 targets cytoglobin to inhibit oxidative stress-induced apoptosis in keratinocytes and lung cancer. Oncogene. 2016; 35:1493–503. https://doi.org/10.1038/onc.2015.222 [PubMed]
- 104. Niklison-Chirou MV, Steinert JR, Agostini M, Knight RA, Dinsdale D, Cattaneo A, Mak TW, Melino G. TAp73 knockout mice show morphological and functional nervous system defects associated with loss of p75 neurotrophin receptor. Proc Natl Acad Sci USA. 2013; 110:18952–57. https://doi.org/10.1073/pnas.1221172110 [PubMed]
- 105. Rufini A, Niklison-Chirou MV, Inoue S, Tomasini R, Harris IS, Marino A, Federici M, Dinsdale D, Knight RA, Melino G, Mak TW. TAp73 depletion accelerates aging through metabolic dysregulation. Genes Dev. 2012; 26:2009–14. https://doi.org/10.1101/gad.197640.112 [PubMed]
- 106. Tomasini R, Tsuchihara K, Wilhelm M, Fujitani M, Rufini A, Cheung CC, Khan F, Itie-Youten A, Wakeham A, Tsao MS, Iovanna JL, Squire J, Jurisica I, et al. TAp73 knockout shows genomic instability with infertility and tumor suppressor functions. Genes Dev. 2008; 22:2677–91. https://doi.org/10.1101/gad.1695308 [PubMed]
- 107. Amelio I, Inoue S, Markert EK, Levine AJ, Knight RA, Mak TW, Melino G. TAp73 opposes tumor angiogenesis by promoting hypoxia-inducible factor 1α degradation. Proc Natl Acad Sci USA. 2015; 112:226–31. https://doi.org/10.1073/pnas.1410609111 [PubMed]
- 108. Agostini M, Romeo F, Inoue S, Niklison-Chirou MV, Elia AJ, Dinsdale D, Morone N, Knight RA, Mak TW, Melino G. Metabolic reprogramming during neuronal differentiation. Cell Death Differ. 2016; 23:1502–14. https://doi.org/10.1038/cdd.2016.36 [PubMed]
- 109. Tóth B, Garabuczi E, Sarang Z, Vereb G, Vámosi G, Aeschlimann D, Blaskó B, Bécsi B, Erdõdi F, Lacy-Hulbert A, Zhang A, Falasca L, Birge RB, et al. Transglutaminase 2 is needed for the formation of an efficient phagocyte portal in macrophages engulfing apoptotic cells. J Immunol. 2009; 182:2084–92. https://doi.org/10.4049/jimmunol.0803444 [PubMed]
- 110. Agostini M, Tucci P, Steinert JR, Shalom-Feuerstein R, Rouleau M, Aberdam D, Forsythe ID, Young KW, Ventura A, Concepcion CP, Han YC, Candi E, Knight RA, et al. microRNA-34a regulates neurite outgrowth, spinal morphology, and function. Proc Natl Acad Sci USA. 2011; 108:21099–104. https://doi.org/10.1073/pnas.1112063108 [PubMed]
- 111. Agostini M, Tucci P, Killick R, Candi E, Sayan BS, Rivetti di Val Cervo P, Nicotera P, McKeon F, Knight RA, Mak TW, Melino G. Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci USA. 2011; 108:21093–98. https://doi.org/10.1073/pnas.1112061109 [PubMed]
- 112. Martin-Lopez M, Maeso-Alonso L, Fuertes-Alvarez S, Balboa D, Rodríguez-Cortez V, Weltner J, Diez-Prieto I, Davis A, Wu Y, Otonkoski T, Flores ER, Menéndez P, Marques MM, Marin MC. p73 is required for appropriate BMP-induced mesenchymal-to-epithelial transition during somatic cell reprogramming. Cell Death Dis. 2017; 8:e3034. https://doi.org/10.1038/cddis.2017.432 [PubMed]
- 113. Matin RN, Chikh A, Chong SL, Mesher D, Graf M, Sanza’ P, Senatore V, Scatolini M, Moretti F, Leigh IM, Proby CM, Costanzo A, Chiorino G, et al. p63 is an alternative p53 repressor in melanoma that confers chemoresistance and a poor prognosis. J Exp Med. 2013; 210:581–603. https://doi.org/10.1084/jem.20121439 [PubMed]
- 114. Schmitz AA, Govek EE, Böttner B, Van Aelst L. Rho GTPases: signaling, migration, and invasion. Exp Cell Res. 2000; 261:1–12. https://doi.org/10.1006/excr.2000.5049 [PubMed]
- 115. Nagano M, Hoshino D, Koshikawa N, Akizawa T, Seiki M. Turnover of focal adhesions and cancer cell migration. Int J Cell Biol. 2012; 2012:310616. https://doi.org/10.1155/2012/310616 [PubMed]
- 116. Halaoui R, McCaffrey L. Rewiring cell polarity signaling in cancer. Oncogene. 2015; 34:939–50. https://doi.org/10.1038/onc.2014.59 [PubMed]
- 117. Sgrò F, Bianchi FT, Falcone M, Pallavicini G, Gai M, Chiotto AM, Berto GE, Turco E, Chang YJ, Huttner WB, Di Cunto F. Tissue-specific control of midbody microtubule stability by Citron kinase through modulation of TUBB3 phosphorylation. Cell Death Differ. 2016; 23:801–13. https://doi.org/10.1038/cdd.2015.142 [PubMed]
- 118. Wu DW, Lee MC, Wang J, Chen CY, Cheng YW, Lee H. DDX3 loss by p53 inactivation promotes tumor malignancy via the MDM2/Slug/E-cadherin pathway and poor patient outcome in non-small-cell lung cancer. Oncogene. 2014; 33:1515–26. https://doi.org/10.1038/onc.2013.107 [PubMed]
- 119. Schubert J, Brabletz T. p53 Spreads out further: suppression of EMT and stemness by activating miR-200c expression. Cell Res. 2011; 21:705–07. https://doi.org/10.1038/cr.2011.62 [PubMed]
- 120. Kim T, Veronese A, Pichiorri F, Lee TJ, Jeon YJ, Volinia S, Pineau P, Marchio A, Palatini J, Suh SS, Alder H, Liu CG, Dejean A, Croce CM. p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J Exp Med. 2011; 208:875–83. https://doi.org/10.1084/jem.20110235 [PubMed]
- 121. Cianfrocca R, Rosanò L, Tocci P, Sestito R, Caprara V, Di Castro V, De Maria R, Bagnato A. Blocking endothelin-1-receptor/β-catenin circuit sensitizes to chemotherapy in colorectal cancer. Cell Death Differ. 2017; 24:1811–20. https://doi.org/10.1038/cdd.2017.121 [PubMed]
- 122. Delaidelli A, Negri GL, Jan A, Jansonius B, El-Naggar A, Lim JK, Khan D, Zarni Oo H, Carnie CJ, Remke M, Maris JM, Leprivier G, Sorensen PH. MYCN amplified neuroblastoma requires the mRNA translation regulator eEF2 kinase to adapt to nutrient deprivation. Cell Death Differ. 2017; 24:1564–76. https://doi.org/10.1038/cdd.2017.79 [PubMed]
- 123. Bao W, Strömblad S. Integrin alphav-mediated inactivation of p53 controls a MEK1-dependent melanoma cell survival pathway in three-dimensional collagen. J Cell Biol. 2004; 167:745–56. https://doi.org/10.1083/jcb.200404018 [PubMed]
- 124. Mak AS. p53 in cell invasion, podosomes, and invadopodia. Cell Adhes Migr. 2014; 8:205–14. https://doi.org/10.4161/cam.27841 [PubMed]
- 125. Di J, Huang H, Wang Y, Qu D, Tang J, Cheng Q, Lu Z, Zhang Y, Zheng J. p53 target gene Rap2B regulates the cytoskeleton and inhibits cell spreading. J Cancer Res Clin Oncol. 2015; 141:1791–98. https://doi.org/10.1007/s00432-015-1948-8 [PubMed]
- 126. Heiss NS, Gloeckner G, Bächner D, Kioschis P, Klauck SM, Hinzmann B, Rosenthal A, Herman GE, Poustka A. Genomic structure of a novel LIM domain gene (ZNF185) in Xq28 and comparisons with the orthologous murine transcript. Genomics. 1997; 43:329–38. https://doi.org/10.1006/geno.1997.4810 [PubMed]
- 127. Dawid IB, Breen JJ, Toyama R. LIM domains: multiple roles as adapters and functional modifiers in protein interactions. Trends Genet. 1998; 14:156–62. https://doi.org/10.1016/S0168-9525(98)01424-3 [PubMed]
- 128. Bach I. The LIM domain: regulation by association. Mech Dev. 2000; 91:5–17. https://doi.org/10.1016/S0925-4773(99)00314-7 [PubMed]
- 129. Furukawa D, Chijiwa T, Matsuyama M, Mukai M, Matsuo EI, Nishimura O, Kawai K, Suemizu H, Nakagohri T, Ozawa S, Shimada K, Hiraoka N, Nakamura M. Plasma membrane expression of ZNF185 is a prognostic factor in pancreatic ductal carcinoma. Oncol Lett. 2017; 14:3633–40. https://doi.org/10.3892/ol.2017.6633 [PubMed]
- 130. Furukawa D, Chijiwa T, Matsuyama M, Mukai M, Matsuo EI, Nishimura O, Kawai K, Suemizu H, Hiraoka N, Nakagohri T, Yasuda S, Nakamura M. Zinc finger protein 185 is a liver metastasis-associated factor in colon cancer patients. Mol Clin Oncol. 2014; 2:709–13. https://doi.org/10.3892/mco.2014.298 [PubMed]
- 131. Vanaja DK, Cheville JC, Iturria SJ, Young CY. Transcriptional silencing of zinc finger protein 185 identified by expression profiling is associated with prostate cancer progression. Cancer Res. 2003; 63:3877–82. [PubMed]
- 132. Gonzalez HE, Gujrati M, Frederick M, Henderson Y, Arumugam J, Spring PW, Mitsudo K, Kim HW, Clayman GL. Identification of 9 genes differentially expressed in head and neck squamous cell carcinoma. Arch Otolaryngol Head Neck Surg. 2003; 129:754–59. https://doi.org/10.1001/archotol.129.7.754 [PubMed]
- 133. Medina PP, Carretero J, Ballestar E, Angulo B, Lopez-Rios F, Esteller M, Sanchez-Cespedes M. Transcriptional targets of the chromatin-remodelling factor SMARCA4/BRG1 in lung cancer cells. Hum Mol Genet. 2005; 14:973–82. https://doi.org/10.1093/hmg/ddi091 [PubMed]
- 134. Krøigård AB, Larsen MJ, Lænkholm AV, Knoop AS, Jensen JD, Bak M, Mollenhauer J, Thomassen M, Kruse TA. Identification of metastasis driver genes by massive parallel sequencing of successive steps of breast cancer progression. PLoS One. 2018; 13:e0189887. https://doi.org/10.1371/journal.pone.0189887 [PubMed]
- 135. Wang J, Huang HH, Liu FB. ZNF185 inhibits growth and invasion of lung adenocarcinoma cells through inhibition of the akt/gsk3β pathway. J Biol Regul Homeost Agents. 2016; 30:683–91. [PubMed]
- 136. Oti M, Kouwenhoven EN, Zhou H. Genome-wide p63-regulated gene expression in differentiating epidermal keratinocytes. Genom Data. 2015; 5:159–63. https://doi.org/10.1016/j.gdata.2015.06.002 [PubMed]
- 137. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017; 45:W98–102. https://doi.org/10.1093/nar/gkx247 [PubMed]



