Introduction
The giant panda (Ailuropoda melanoleuca), an endangered species endemic to western China, has long been threatened with extinction due to human population expansion and habitat destruction [1, 2]. Recent research has shown that giant pandas are well adapted to a specialized bamboo diet via enigmatic gut microbiota and low energy expenditure [3–5]. Contrary to previous studies, the latest genome sequencing and resequencing and genome-wide genetic studies have demonstrated that the giant panda has a relatively high genetic diversity [6–8]. Over the years, extensive research has been conducted on genes of the major histocompatibility complex (MHC) in giant pandas and shown that DRB and DQA genes have low diversity but remain positive selection [9–11]. However, most of these studies focused on the adaptations and genetic diversity of giant pandas and little is known about the gene expression of immune-related genes.
Giant pandas are vulnerable to a variety of carnivore-specific and mammalian infectious and parasitic diseases that seriously threaten the giant panda populations [12–16]. For example, Baylisascaris schroederi is the most common parasite in wild and captive giant pandas and was responsible for half of panda deaths between 2001 and 2005 [15]. Likewise, canine distemper virus (CDV) is highly contagious and fatal and affects a wide range of mammals, including the giant panda [17]. In 2014–2015, an outbreak of CDV infection in giant pandas caused the death of five infected animals in two months [18].
The immune system is vulnerable to age-associated alterations, which accumulate to produce a progressive deterioration and lead to an increased incidence of infectious diseases [19]. Although aging is an inevitable biological process and a powerful risk factor for many diseases, the underlying molecular mechanisms that lead to generalized disease susceptibility are largely unknown [20, 21]. Aging has proven difficult to dissect in part due to its interactions with environmental influences, other genetic factors, and a large number of age-related diseases [22]. Immunosenescence, defined as immune changes with ageing, is an unavoidable life process and has been characterized in several species, such as humans [21, 23], mice [24], zebra finches [25] and wolves [26]. A comprehensive meta-analysis of age-related gene expression profiles indicated that signatures of aging most notably involve an overexpression of inflammation and immune response genes and an underexpression of genes associated with energy metabolism [27]. However, the effects of aging on giant panda have not been characterized, especially the age-related changes of immune system. Moreover, understanding the principles of giant panda immune system is vital for the development vaccines that can elicit protective immunity [28].
The aims of this study were to determine which genes and pathways show differential expression with age in giant panda blood tissues and to understand age-related alterations of the immune system. We used RNA-seq technology to identify age-related differentially expressed genes (DEGs) in giant panda blood samples and performed functional enrichment analysis for these DEGs. The collective data generated in this study may represent a valuable resource to enable further advancements in immunological research in giant pandas.
Results
Transcriptome sequencing and assembly
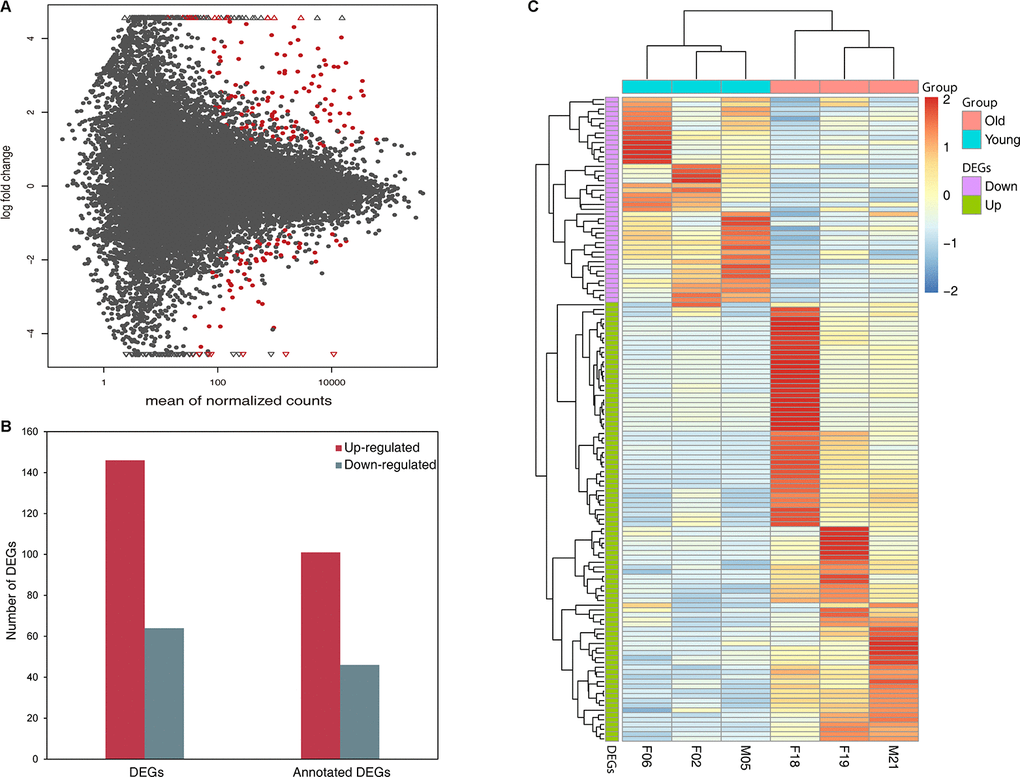
RNA prepared from blood tissues of four giant pandas were subjected to RNA sequencing using Illumina HiSeq™ 2000. Together with the three giant panda transcriptomes from our previous study, we totally acquired approximately 187 million raw paired-end reads and 162 million remained after removing adaptor sequences and discarding low quality reads. The total length of the reads was about 35.8 gigabases (Gb). We aligned each of the seven short-read libraries onto the giant panda Ensembl reference genome (ailMel1) separately and found that an average of 19.86 million high-quality reads (85.68%) could be successfully mapped to the genome per sample. This included a mean of 19.31 million uniquely mapped reads (97.23%) per sample, indicating that the majority of the paired reads aligned correctly (Table 1). These high-quality reads were assembled into 56,543 genes, giving rise to 88,071 transcripts containing 46,091 (52.33%) transcripts that had more than one exon (Supplementary Figure 1). The comparison of similarities between the assembled transcripts and the Ensembl transcripts showed that only 2943 (3.34%) matched exactly with annotated intron chain and a total of 33,600 (38.15%) were identified as potentially novel isoforms of a predicted Ensembl transcript with at least one splice junction shared. The majority of transcripts (34,660, 39.35%) were annotated as intergenic transcripts and a small proportion of transcripts (7824, 8.88%) entirely fell into reference intron (Supplementary Table 1).
Table 1. Summary of sequencing and assembly of transcriptome
| Sample | F02 | F06 | F18 | F19 | M05 | M12 | M21 |
|---|---|---|---|---|---|---|---|
| Age | 2 | 6 | 18 | 19 | 5 | 12 | 21 |
| Group | Young | Young | Old | Old | Young | - | Old |
| Gender | Female | Female | Female | Female | Male | Male | Male |
| Read length (bp) | 100 | 90 | 100 | 90 | 100 | 90 | 100 |
| Raw read pairs | 24,547,347 | 26,523,566 | 19,473,310 | 27,373,071 | 26,680,027 | 25,931,660 | 36,467,323 |
| Total bases (Gb) | 4.91 | 4.77 | 3.89 | 4.93 | 5.34 | 4.67 | 7.29 |
| Clean read pairs | 22,419,441 | 22,352,904 | 17,649,156 | 22,931,640 | 24,654,277 | 22,004,856 | 30,218,700 |
| % of clean read pairs | 91.33% | 84.28% | 90.63% | 83.77% | 92.41% | 84.86% | 82.87% |
| Mapped read pairs | 19,423,023 | 20,089,734 | 15,064,083 | 20,316,914 | 21,831,830 | 19,131,977 | 27,080,383 |
| % of mapped reads | 86.63% | 89.88% | 85.35% | 88.60% | 88.55% | 86.94% | 89.61% |
| Uniquely mapped read pairs | 18,975,652 | 19,663,458 | 14,711,801 | 19,892,536 | 21,380,756 | 18,621,171 | 26,492,780 |
| % of uniquely mapped reads | 97.70% | 97.88% | 97.66% | 97.91% | 97.93% | 97.33% | 97.83% |
Gene ontology enrichment analysis of differentially expressed genes
To gain insights into the biological roles of the DEGs, we performed a GO categories enrichment analysis. Only the annotated DEGs were selected and tested against the background set of all genes with GO annotation. We examined the GO categories separately. We found that for up-regulated genes with age, almost all the significantly enriched GO terms in biological process category were associated with the immune system, such as, immune response (GO:0006955), negative regulation of viral genome replication (GO:0045071), and defense response to virus (GO:0051607). MHC class II protein complex (GO:0042613) and GTP binding (GO:0005525) were the most significantly enriched GO term in the cellular component and molecular function categories, respectively (Figure 2, Supplementary Table 3). For down-regulated genes with age, the most significantly enriched biological process GO terms were complement activation, classical pathway (GO:0006958), B cell receptor signaling pathway (GO:0050853) and Fc-epsilon receptor signaling pathway (GO:0038095) (Figure 3, Supplementary Table 4). Interestingly, several up-regulated genes were enriched in retina homeostasis (GO:0001895) and regulation of blood pressure (GO:0008217), which might be related to age.
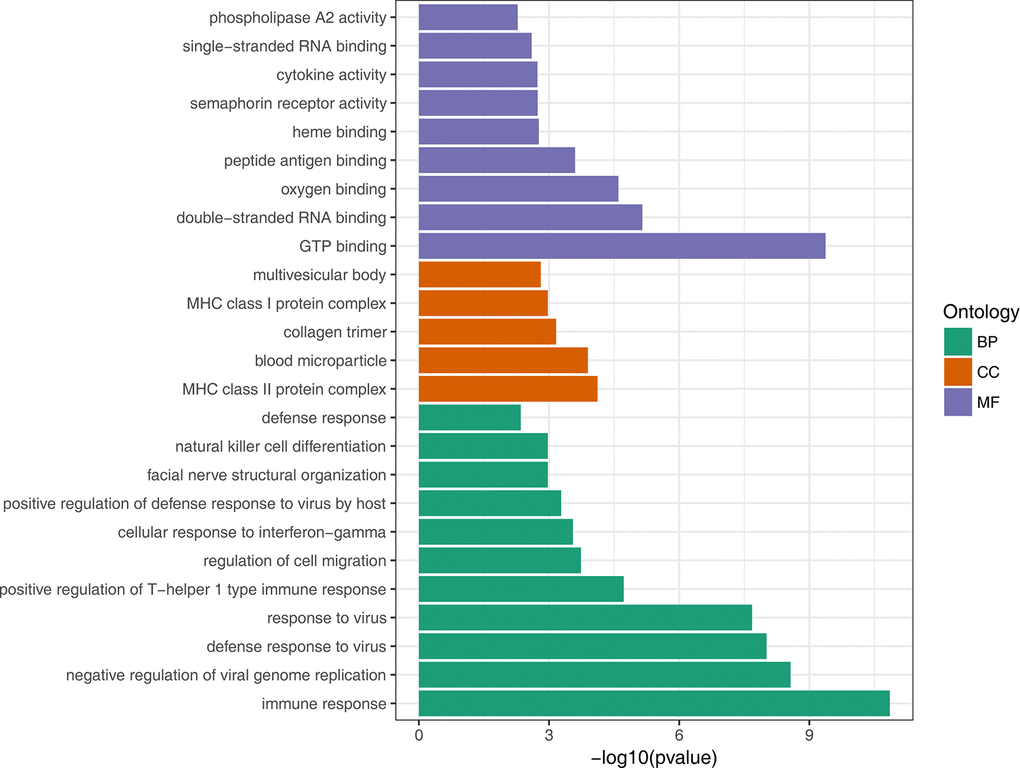
Figure 2. GO enrichment analysis of up-regulated DEGs
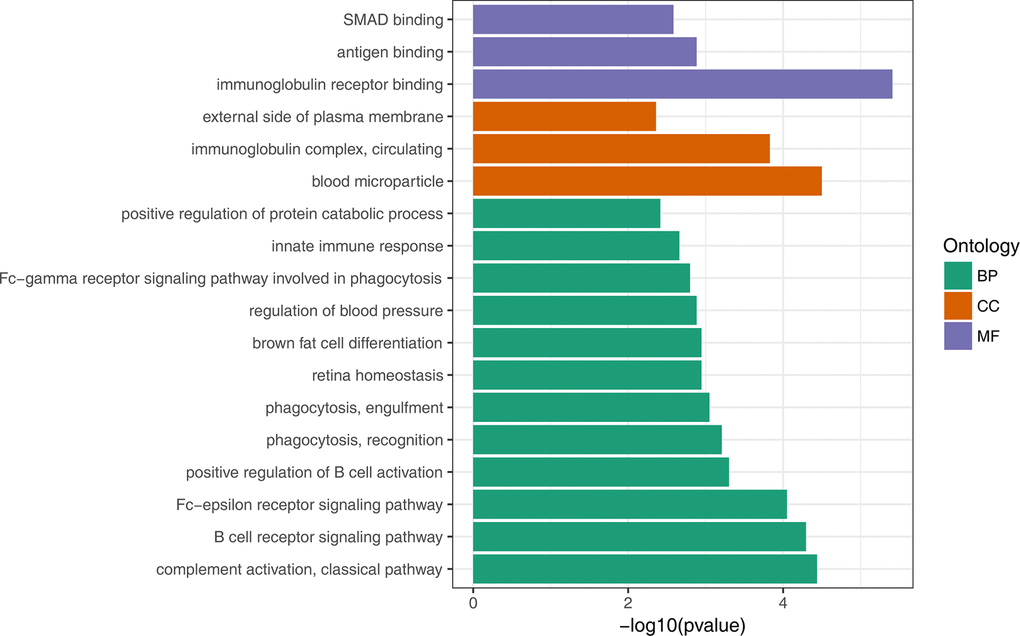
Figure 3. GO enrichment analysis of down-regulated DEGs
Pathway enrichment analysis of differentially expressed genes
To further evaluate the biological significance of the DEGs, we also performed KEGG pathway enrichment analysis. Hypergeometric tests with a P value cutoff of 0.05 was used as the criteria for pathway detection. We found that up-regulated genes were significantly enriched in 35 pathways, among which 22 pathways were disease related and 7 pathways were immune related. However, down-regulated genes were not significantly enriched to any pathways and this may be because many down-regulated genes were not annotated in KEGG pathways. Figure 4 illustrates the result of pathway enrichment of up-regulated genes, in which antigen processing and presentation (ko04612), cytosolic DNA-sensing pathway (ko04623) and RIG-I-like receptor signaling pathway (ko04622) were most significantly enriched.
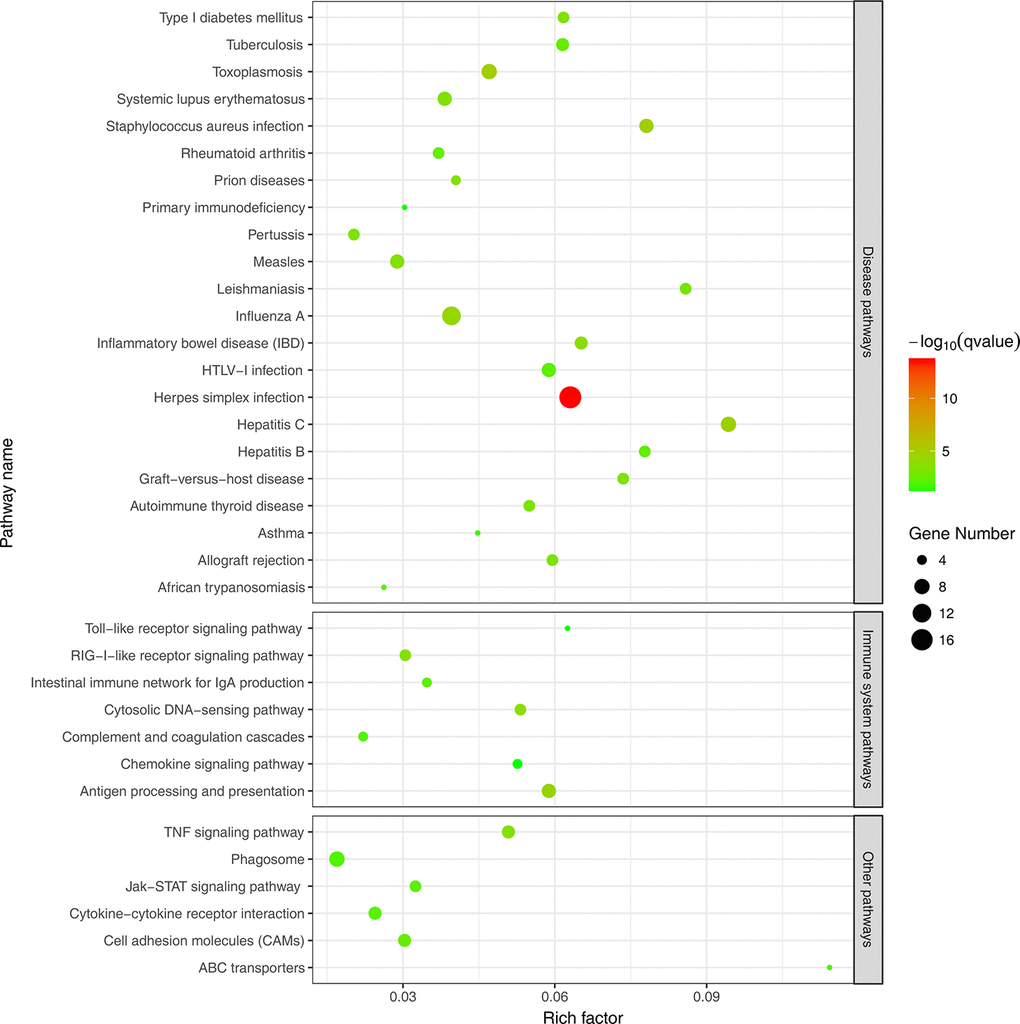
Figure 4. KEGG pathway enrichment analysis of up-regulated DEGs
Protein-protein interaction network of differentially expressed genes
We have performed a protein-protein interaction analysis of all DEGs. A total of 122 interactions between 36 DEGs were extracted from the STRING database. Interferon-stimulated gene 15 (ISG15), signal transducer and activator of transcription 1 (STAT1), interferon regulatory factor 7 (IRF7) and DExD/H-Box Helicase 58 (DDX58) that play important roles in response to pathogen invasion were at key position of the interaction network (Figure 5).
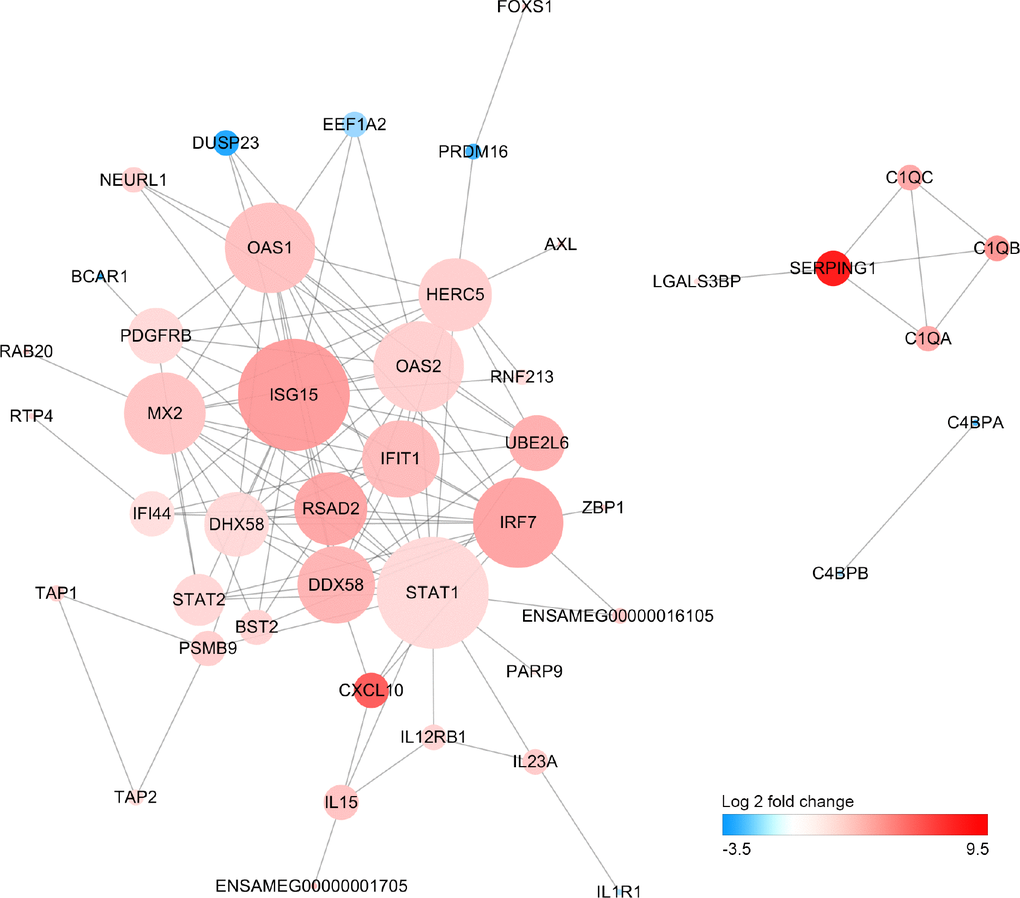
Figure 5. Protein-protein interaction network of differentially expressed genes Size of the node is proportional to the number of DEGs interacted with it, and color of node represents Log2FoldChange in expression levels of DEGs between old and young giant pandas.
Discussion
In a previous study, we have characterized giant panda blood transcriptome and identified 15 immune pathways where more than 70% of the total known genes were mapped by assembled transcripts [30]. In the present study, we still applied the widely used RNA-seq approach to highlight significant DEGs in blood between young and old giant pandas. The previous work has proved that age had a broad impact on gene expression levels, whereas sex had very minimal effects on gene expression patterns [26]. Therefore, it is appropriate to investigate age-related changes in giant panda by sequencing the blood transcriptome. To the best of our knowledge, this is the first study to assess the impact of age on immune-related gene expression.
The assembly results unfortunately underlined the incompleteness of the giant panda genome annotation of current version, and the need for further studies to decipher its complexity. We have provided a valuable resource for improvement of genome assembly and annotation. For example, a recent study has improved the genome assembly and identified novel transcripts by using transcriptomes from 12 giant panda tissues, excluding blood [31]. Another major outcome of transcriptome sequencing is the ability to identify DEGs between different conditions [32]. We identified 210 DEGs, which is much lower than the 625 and 1497 DEGs found in wolves [26] and humans [21], respectively. This may suggest that age-related alterations of gene expression are species-specific [33]. The most significantly up-regulated gene SERPING1, as a signature of mammalian aging, encodes the complement component 1 (C1) inhibitor [27]. The C1 inhibitor’s main function is the inhibition of several different proteases in the complement, contact, coagulation and fibrinolytic systems and has been reported to be related to an increased risk of several age-related human diseases [34–36]. The most significantly down-regulated gene C4BPA encodes the alpha-subunit of complement 4 binding protein. C4BPA is a key regulator of the complement system and has been considered a major factor in the age-related decline of the immune response due to decreased expression [37].
We identified age-related DEGs that may be critical in the progression of the giant panda aging process, such as ISG15, STAT1, IRF7. ISG15 is an innate immune and interferon-induced protein that plays a central role in response to viral infection [38]. ISG15 is also a stress-response gene that has been implicated as a tumor suppressor and contributor to inflammatory responses [39]. ISG15 expression was regulated by telomere length and increased with shortening telomeres caused by aging [40]. STAT1 carries tumor suppressor functions and regulates various cellular activities, such as apoptosis, angiogenesis, invasion and evasion of the immune system [41, 42]. Previous studies indicated that loss of STAT1 expression has been implicated in the pathobiology of various types of human cancer [41]. IRF7 has been discovered as the crucial regulator of type I interferons against pathogenic infections and plays important roles in immune modulation and regulation of oncogenesis and apoptosis [43]. Our results showed ISG15 can interact with both STAT1 and IRF7 and other antiviral related genes. It is possible that giant panda aging increased expression of ISG15 result in regulating innate immunity by interacting with genes involved in anti-viral responses. Furthermore, the invasive viruses can lead to activation of IRFs and induction of interferons (IFNs) resulting in antiviral IFNs signal through the JAK/STAT pathway to induce ISG production [44].
Aging is associated with functional integrity of the immune system declines leading to increased susceptibility to aging diseases [45]. Both GO and KEGG enrichment analysis showed that up-regulated genes were mainly involved in the immune response, particularly defense against viruses. The involved genes, such as IFIT5, BST2 and OAS1, were closely related to the innate immunity response [46–48]. In contrast, the down-regulated genes were implicated in the B cell receptor signaling pathway and positive regulation of B cell activation. In accordance with observations in wolf blood transcriptome with age, down-regulated genes were mainly associated with B cells and up-regulated genes were significantly associated with innate immunity [26]. Although, both innate and adaptive immune responses are affected by the aging process, the adaptive response seems to be more affected by age-related changes [49]. However, contrary to the previous results observed in the wolf [26], MHC class I and II genes were up-regulated with age in our study. In addition to their central role in adaptive immunity, both MHC class I and II genes can regulate Toll-Like Receptor to mediate the innate immune response [50, 51]. Taken together, these suggest the possibility that the aging immune system maintains a persistently activated innate immune response to compensate for the decline in the adaptive immune function [52].
Vaccines appear as one of the most efficient medical interventions against infectious diseases [53] and have been also used to assist giant pandas to defend against CDV infections [54, 55]. However, the vaccines for giant pandas do not elicit consistent antibody titers [56]. One explanation may be that the attenuated vaccine is inadequate to stimulate giant pandas to produce a high level of antibodies against CDV [55]. Another possible explanation might be aging-associated declines in immune system function. It is clear that the decline in adaptive immunity leads to dramatically reduced vaccine responses and vaccine longevity in older individuals [57]. We have noted that the immune response genes, such as IGHV3-23 and IGKC, that participate in the antigen recognition and B cell activation were down-regulated with ageing. This may lead to a decline of adaptive immunity resulting in decreased response to vaccines in giant pandas. In addition, the decreased ability of the adaptive immune system is closely related with the decline in production of naive lymphocytes and the accumulation of functionally impaired memory lymphocytes during aging [58].
In conclusion, we identified 210 genes which differentially expressed between old and young giant pandas. The up-regulated genes with age were mainly involved in the innate immune response, particularly in defense against virus. Instead, the down-regulated genes with age were mainly related to B cell activation. This suggests that innate immunity is relatively well preserved in aged giant pandas, although adaptive immunity deteriorates due to a decline of naive lymphocytes and the expansion of incompetent memory lymphocytes. Moreover, ISG15, STAT1, IRF7 and DDX58 were identified as the hub genes that play a pivotal role in response to viral infection and their expression were up-regulated with ageing. This work provides insight into gene expression changes associated with aging and will provide a foundation for future studies on the molecular mechanisms underlying immune changes associated with ageing.
Materials and Methods
Sample collection
Blood samples were collected during a routine examination from four captive giant pandas at the Chengdu Research Base of Giant Panda Breeding in Chengdu, China. The other three giant panda blood transcriptomes were obtained from our previous study in NCBI SRA (project Accession no. SRP041998). Samples collection and utility protocols were approved by the Chengdu Institute of Biology Animal Use Ethics, which is responsible for the Chengdu Research Base of Giant Panda Breeding. The fresh blood samples were immediately stored in RNAlater (Ambion Inc., Austin, TX, USA). Total RNA was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol and treated with RNase-free DNase I.
Library preparation and sequencing
A total amount of 3 μg RNA per sample was used as input material for the RNA sample preparations. Sequencing libraries were generated using NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA) following manufacturer’s recommendations and index codes were added to attribute sequences to each sample. Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature in NEBNext First Strand Synthesis Reaction Buffer (5X). First strand cDNA was synthesized using random hexamer primer and M-MuLV Reverse Transcriptase (RNase H−). Second strand cDNA synthesis was subsequently performed using DNA Polymerase I and RNase H. Remaining overhangs were converted into blunt ends via exonuclease/polymerase activities. After adenylation of 3’ ends of DNA fragments, NEBNext Adaptor with hairpin loop structure were ligated to prepare for hybridization. In order to select cDNA fragments of 150~200 bp in length, the library fragments were purified with AMPure XP system (Beckman Coulter, Beverly, USA). Then 3 μl USER Enzyme (NEB, USA) was used with size-selected, adaptor-ligated cDNA at 37° C for 15 min followed by 5 min at 95° C before PCR. Then PCR was performed with Phusion High-Fidelity DNA polymerase, Universal PCR primers and Index (X) Primer. Lastly, PCR products were purified (AMPure XP system) and library quality was assessed on the Agilent Bioanalyzer 2100 system. The library preparations were sequenced on an Illumina Hiseq 2000 platform and 100 bp paired-end reads were generated.
Quality control and transcriptome assembly
The giant panda genome assembly ailMel1 and reference annotation were downloaded from Ensembl v83 (www.ensembl.org). In order to obtain high quality reads, we employed NGS QC Toolkit [59] with stringent criteria (high-quality paired reads with more than 90% of bases with Q-value ≥ 20 were retained) to remove the low-quality paired-end reads or reads containing adaptors. High-quality paired-end reads from seven libraries were mapped on giant panda reference genome using HISAT [60], separately. Each of the alignment output files were assembled into separate transcriptomes using StringTie [61], which produces a transcript GTF file. The transcript GTF files produced after assembling were merged together to generate a complete transcript annotation file using StringTie with–merge option.
Functional annotation
The merged assemblies were compared using the Cufflinks [62] inclusive utility Cuffcompare with the Ensembl annotations to find known and novel genes and isoforms, as well as transcripts expressed from intergenic regions. Only the genes with “=” and “j” class code were considered to be known genes. The GO annotations of these genes were extracted from Ensembl BioMart (http://www.ensembl.org/biomart/martview). The unknown gene sequences were searched against Swiss-Prot and NCBI non-redundant (NR) database using BLASTX [63] with a typical cutoff E-value of 1E-5. Gene ontology (GO) was applied with the Blast2GO [64] to obtain annotation of unknown genes.
Gene expression estimation and differentially expressed genes analysis
We used the merged GTF file as the reference annotation file to guide the assembly process. StringTie was performed to estimate the genes expression level with -G, -b and -e options. The expression value of TPM (transcripts per million) and raw read counts for each gene were extracted from above result. Differential expression analysis was performed using DESeq2 [65] that takes the raw read count as the input. The results of all statistical tests were corrected for multiple testing with the Benjamini-Hochberg false discovery rate (FDR ≤ 0.05) and an absolute value of log 2 fold change ≥ 1 was used to determine significant differences in gene expression.
Gene enrichment analysis
To functional classify the DEGs, we performed GO enrichment analysis using clusterProfiler [66] and searched for over-represented GO terms in three categories, namely biological process, molecular function and cellular component. We also performed KEGG pathway enrichment analysis using KOBAS [67]. An FDR cut-off of 0.05 was considered to be significantly enriched.
Supplementary Materials
Acknowledgements
We thank the Chengdu Research Base of Giant Panda Breeding for their help in collecting the samples for this study. We also appreciate Dr. Megan Price for critical reading and English editing on the manuscript.
Conflicts of Interest
The authors disclose no potential conflicts of interest.
Funding
This work was supported by the National Natural Science Foundation of China (31570534) and Chengdu Panda Breeding Research Foundation (CPF2017-22).
References
- 1. Huang J, Li Y, Du L, Yang B, Shen F, Zhang H, Zhang Z, Zhang X, Yue B. Genome-wide survey and analysis of microsatellites in giant panda (Ailuropoda melanoleuca), with a focus on the applications of a novel microsatellite marker system. BMC Genomics. 2015; 16:61. https://doi.org/10.1186/s12864-015-1268-z [PubMed]
- 2. Wei F, Hu Y, Zhu L, Bruford MW, Zhan X, Zhang L. Black and white and read all over: the past, present and future of giant panda genetics. Mol Ecol. 2012; 21:5660–5674. https://doi.org/10.1111/mec.12096 [PubMed]
- 3. Nie Y, Speakman JR, Wu Q, Zhang C, Hu Y, Xia M, Yan L, Hambly C, Wang L, Wei W, Zhang J, Wei F. Exceptionally low daily energy expenditure in the bamboo-eating giant panda. Science. 2015; 349:171–174. https://doi.org/10.1126/science.aab2413 [PubMed]
- 4. Xue Z, Zhang W, Wang L, Hou R, Zhang M, Fei L, Zhang X, Huang H, Bridgewater LC, Jiang Y, Jiang C, Zhao L, Pang X, et al. The bamboo-eating giant panda harbors a carnivore-like gut microbiota, with excessive seasonal variations. mBio. 2015; 6:e00022–15. https://doi.org/10.1128/mBio.00022-15 [PubMed]
- 5. Wei F, Wang X, Wu Q. The giant panda gut microbiome. Trends Microbiol. 2015; 23:450–452. https://doi.org/10.1016/j.tim.2015.06.004 [PubMed]
- 6. Li R, Fan W, Tian G, Zhu H, He L, Cai J, Huang Q, Cai Q, Li B, Bai Y, Zhang Z, Zhang Y, Wang W, et al. The sequence and de novo assembly of the giant panda genome. Nature. 2010; 463:311–317. https://doi.org/10.1038/nature08696 [PubMed]
- 7. Zhao S, Zheng P, Dong S, Zhan X, Wu Q, Guo X, Hu Y, He W, Zhang S, Fan W, Zhu L, Li D, Zhang X, et al. Whole genome sequencing of giant pandas provides insights into demographic history and local adaptation. Nat Genet. 2013; 45:57–71. https://doi.org/10.1038/ng.2494 [PubMed]
- 8. Wei F, Hu Y, Yan L, Nie Y, Wu Q, Zhang Z. Giant pandas are not an evolutionary cul-de-sac: evidence from multidisciplinary research. Mol Biol Evol. 2015; 34:4–12. https://doi.org/10.1093/molbev/msu278 [PubMed]
- 9. Wan Q, Zhu L, Wu H, Fang S. Major histocompatibility complex class II variation in the giant panda (Ailuropoda melanoleuca). Mol Ecol. 2006; 15:2441–2450. https://doi.org/10.1111/j.1365-294X.2006.02966.x [PubMed]
- 10. Zhu L, Ruan X, Ge Y, Wan H, Fang S. Low major histocompatibility complex class II DQA diversity in the giant panda (Ailuropoda melanoleuca). BMC Genet. 2007; 8:29. https://doi.org/10.1186/1471-2156-8-29 [PubMed]
- 11. Zhang L, Wu Q, Hu Y, Wei F. Major histocompatibility complex alleles associated with parasite susceptibility in wild giant pandas. Heredity. 2015; 114:85–93. https://doi.org/10.1038/hdy.2014.73 [PubMed]
- 12. Qin Q, Li D, Zhang H, Hou R, Zhang Z, Zhang C, Zhang J, Wei F. Serosurvey of selected viruses in captive giant pandas (Ailuropoda melanoleuca) in China. Vet Microbiol. 2010; 142:199–204. https://doi.org/10.1016/j.vetmic.2009.09.062 [PubMed]
- 13. Li D, Zhu L, Cui H, Ling S, Fan S, Yu Z, Zhou Y, Wang T, Qian J, Xia X, Xu Z, Gao Y, Wang C. Influenza A(H1N1)pdm09 virus infection in giant pandas, China. Emerg Infect Dis. 2014; 20:480–483. https://doi.org/10.3201/eid2003.131531 [PubMed]
- 14. Wang X, Yan Q, Xia X, Zhang Y, Li D, Wang C, Chen S, Hou R. Serotypes, virulence factors, and antimicrobial susceptibilities of vaginal and fecal isolates of Escherichia coli from giant pandas. Appl Environ Microbiol. 2013; 79:5146–5150. https://doi.org/10.1128/AEM.01367-13 [PubMed]
- 15. Xie Y, Zhou X, Zhang Z, Wang C, Sun Y, Liu T, Gu X, Wang T, Peng X, Yang G. Absence of genetic structure in Baylisascaris schroederi populations, a giant panda parasite, determined by mitochondrial sequencing. Parasit Vectors. 2014; 7:606. https://doi.org/10.1186/s13071-014-0606-3 [PubMed]
- 16. Wang T, Chen Z, Xie Y, Hou R, Wu Q, Gu X, Lai W, Peng X, Yang G. Prevalence and molecular characterization of Cryptosporidium in giant panda (Ailuropoda melanoleuca) in Sichuan province, China. Parasit Vectors. 2015; 8:344. https://doi.org/10.1186/s13071-015-0953-8 [PubMed]
- 17. Guo L, Yang S, Wang C, Hou R, Chen S, Yang X, Liu J, Pan H, Hao Z, Zhang M, Cao S, Yan Q. Phylogenetic analysis of the haemagglutinin gene of canine distemper virus strains detected from giant panda and raccoon dogs in China. Virol J. 2013; 10:109. https://doi.org/10.1186/1743-422X-10-109 [PubMed]
- 18. Feng N, Yu Y, Wang T, Wilker P, Wang J, Li Y, Sun Z, Gao Y, Xia X. Fatal canine distemper virus infection of giant pandas in China. Sci Rep. 2016; 6:17518. https://doi.org/10.1038/srep27518 [PubMed]
- 19. Aw D, Silva AB, Palmer DB. Immunosenescence: emerging challenges for an aging population. Immunology. 2007; 120:435–446. https://doi.org/10.1111/j.1365-2567.2007.02555.x [PubMed]
- 20. Niccoli T and Partridge L. Aging as a risk factor for disease. Curr Biol. 2012; 22:R741–R752. https://doi.org/10.1016/j.cub.2012.07.024 [PubMed]
- 21. Peters MJ, Joehanes R, Pilling LC, Schurmann C, Conneely KN, Powell J, Reinmaa E, Sutphin GL, Zhernakova A, Schramm K, Wilson YA, Kobes S, Tukiainen T, et al. The transcriptional landscape of age in human peripheral blood. Nat Commun. 2015; 6:8570. https://doi.org/10.1038/ncomms9570 [PubMed]
- 22. Sebastiani P, Solovieff N, Dewan AT, Walsh KM, Puca A, Hartley SW, Melista E, Andersen S, Dworkis DA, Wilk JB, Myers RH, Steinberg MH, Montano M, et al. Genetic signatures of exceptional longevity in humans. PLoS ONE. 2012; 7:e29848. https://doi.org/10.1371/journal.pone.0029848 [PubMed]
- 23. Gayoso I, Sanchez-Correa B, Campos C, Alonso C, Pera A, Casado JG, Morgado S, Tarazona R, Solana R. Immunosenescence of human natural killer cells. J Innate Immun. 2011; 3:337–343. https://doi.org/10.1159/000328005 [PubMed]
- 24. Maue AC, Yager EJ, Swain SL, Woodland DL, Blackman MA, Haynes L. T-cell immunosenescence: lessons learned from mouse models of aging. Trends Immunol. 2009; 30:301–305. https://doi.org/10.1016/j.it.2009.04.007 [PubMed]
- 25. Noreen E, Bourgeon S, Bech C. Growing old with the immune system: a study of immunosenescence in the zebra finch (Taeniopygia guttata). J Comp Physiol B. 2011; 181:649–656. https://doi.org/10.1007/s00360-011-0553-7 [PubMed]
- 26. Charruau P, Johnston RA, Stahler DR, Lea A, Snyder-Mackler N, Smith DW, vonHoldt BM, Cole SW, Tung J, Wayne RK. Pervasive effects of aging on gene expression in wild wolves. Mol Biol Evol. 2016; 33:1967–1978. https://doi.org/10.1093/molbev/msw072 [PubMed]
- 27. de Magalhães JP, Curado J, Church GM. Meta-analysis of age-related gene expression profiles identifies common signatures of aging. Bioinformatics. 2009; 25:875–881. https://doi.org/10.1093/bioinformatics/btp073 [PubMed]
- 28. Medzhitov R. Recognition of microorganisms and activation of the immune response. Nature. 2007; 449:819–826. https://doi.org/10.1038/nature06246 [PubMed]
- 29. Hiroshi K, Shigeo O, Hideo K, Shigehisa T, Tsuneo S, Kiichi K, In-Jen P, Toshi W, Zhang Z, Hou R, Wang C, Shen F, Zhang L, et al. Seroprevalence of susceptibility to morbillivirus infection on giant panda in captivity. Chin J Vet Sci. 2008; 28:1167–1170.
- 30. Du L, Li W, Fan Z, Shen F, Yang M, Wang Z, Jian Z, Hou R, Yue B, Zhang X. First insights into the giant panda (Ailuropoda melanoleuca) blood transcriptome: a resource for novel gene loci and immunogenetics. Mol Ecol Resour. 2015; 15:1001–1013. https://doi.org/10.1111/1755-0998.12367 [PubMed]
- 31. Chen M, Hu Y, Liu J, Wu Q, Zhang C, Yu J, Xiao J, Wei F, Wu J. Improvement of genome assembly completeness and identification of novel full-length protein-coding genes by RNA-seq in the giant panda genome. Sci Rep. 2015; 5:18019. https://doi.org/10.1038/srep18019 [PubMed]
- 32. Li M, Wang F, Jiang Q, Ma J, Xiong A. Identification of SSRs and differentially expressed genes in two cultivars of celery (Apium graveolens L.) by deep transcriptome sequencing. Hortic Res. 2014; 1:10. https://doi.org/10.1038/hortres.2014.10 [PubMed]
- 33. Yang J, Huang T, Petralia F, Long Q, Zhang B, Argmann C, Zhao Y, Mobbs CV, GTEx Consortium, Schadt EE, Zhu J, Tu Z. Synchronized age-related gene expression changes across multiple tissues in human and the link to complex diseases. Sci Rep. 2015; 5:15145. https://doi.org/10.1038/srep15145 [PubMed]
- 34. Cicardi M, Zingale L, Zanichelli A, Pappalardo E, Cicardi B. C1 inhibitor: molecular and clinical aspects. Semin Immunopathol. 2005; 27:286–298. https://doi.org/10.1007/s00281-005-0001-4 [PubMed]
- 35. Ennis S, Jomary C, Mullins R, Cree A, Chen X, Macleod A, Jones S, Collins A, Stone E, Lotery A. Association between the SERPING1 gene and age-related macular degeneration: a two-stage case–control study. Lancet. 2008; 372:1828–1834. https://doi.org/10.1016/S0140-6736(08)61348-3 [PubMed]
- 36. Firinu D, Colomba P, Manconi PE, Barca MP, Fenu L, Piseddu G, Zizzo C, Del Giacco SR, Duro G. Identification of a novel and recurrent mutation in the SERPING1 gene in patients with hereditary angioedema. Clin Immunol. 2013; 147:129–132. https://doi.org/10.1016/j.clim.2013.03.007 [PubMed]
- 37. Lavery LW and Goyns MH. Decline in the expression of C4 binding protein alpha-chain gene during aging of the rat liver. Biogerontology. 2002; 3:207–211. https://doi.org/10.1023/A:1016251929165 [PubMed]
- 38. Perng YC and Lenschow DJ. ISG15 in antiviral immunity and beyond. Nat Rev Microbiol. 2018; 16:423–439. https://doi.org/10.1038/s41579-018-0020-5 [PubMed]
- 39. Andersen JB and Hassel BA. The interferon regulated ubiquitin-like protein, ISG15, in tumorigenesis: friend or foe? Cytokine Growth Factor Rev. 2006; 17:411–421. https://doi.org/10.1016/j.cytogfr.2006.10.001 [PubMed]
- 40. Lou Z, Wei J, Riethman H, Baur JA, Voglauer R, Shay JW, Wright WE. Telomere length regulates ISG15 expression in human cells. Aging (Albany NY). 2009; 1:608–621. https://doi.org/10.18632/aging.100066 [PubMed]
- 41. Zhang Y, Molavi O, Su M, Lai R. The clinical and biological significance of STAT1 in esophageal squamous cell carcinoma. BMC Cancer. 2014; 14:791. https://doi.org/10.1186/1471-2407-14-791 [PubMed]
- 42. Wu C, Molavi O, Zhang H, Gupta N, Alshareef A, Bone KM, Gopal K, Wu F, Lewis JT, Douglas DN, Kneteman NM, Lai R. STAT1 is phosphorylated and downregulated by the oncogenic tyrosine kinase NPM-ALK in ALK-positive anaplastic large-cell lymphoma. Blood. 2015; 126:336–345. https://doi.org/10.1182/blood-2014-10-603738 [PubMed]
- 43. Ning S, Pagano JS, Barber GN. IRF7: activation, regulation, modification and function. Genes Immun. 2011; 12:399–414. https://doi.org/10.1038/gene.2011.21 [PubMed]
- 44. Schoggins JW and Rice CM. Interferon-stimulated genes and their antiviral effector functions. Curr Opin Virol. 2011; 1:519–525. https://doi.org/10.1016/j.coviro.2011.10.008 [PubMed]
- 45. Frenk S and Houseley J. Gene expression hallmarks of cellular ageing. Biogerontology. 2018; 19:547–566. https://doi.org/10.1007/s10522-018-9750-z [PubMed]
- 46. Zhang B, Liu X, Chen W, Chen L. IFIT5 potentiates anti-viral response through enhancing innate immune signaling pathways. Acta Biochim Biophys Sin. 2013; 45:867–874. https://doi.org/10.1093/abbs/gmt088 [PubMed]
- 47. Evans DT, Serra-Moreno R, Singh RK, Guatelli JC. BST-2/tetherin: a new component of the innate immune response to enveloped viruses. Trends Microbiol. 2010; 18:388–396. https://doi.org/10.1016/j.tim.2010.06.010 [PubMed]
- 48. Fagone P, Nunnari G, Lazzara F, Longo A, Cambria D, Distefano G, Palumbo M, Nicoletti F, Malaguarnera L, Rosa MD. Induction of OAS gene family in HIV monocyte infected patients with high and low viral load. Antivir Res. 2016; 131:66–73. https://doi.org/10.1016/j.antiviral.2016.04.009 [PubMed]
- 49. Castelo-Branco C and Soveral I. The immune system and aging: a review. Gynecol Endocrinol. 2014; 30:16–22. https://doi.org/10.3109/09513590.2013.852531 [PubMed]
- 50. Xu S, Liu X, Bao Y, Zhu X, Han C, Zhang P, Zhang X, Li W, Cao X. Constitutive MHC class I molecules negatively regulate TLR-triggered inflammatory responses via the Fps-SHP-2 pathway. Nat Immunol. 2012; 13:551–559. https://doi.org/10.1038/ni.2283 [PubMed]
- 51. Liu X, Zhan Z, Li D, Xu L, Ma F, Zhang P, Yao H, Cao X. Intracellular MHC class II molecules promote TLR-triggered innate immune responses by maintaining activation of the kinase Btk. Nat Immunol. 2011; 12:416–424. https://doi.org/10.1038/ni.2015 [PubMed]
- 52. Sharma G, Hanania NA, Shim YM. The aging immune system and its relationship to the development of chronic obstructive pulmonary disease. Proc Am Thorac Soc. 2009; 6:573–580. https://doi.org/10.1513/pats.200904-022RM [PubMed]
- 53. Rappuoli R, Pizza M, Del Giudice G, De Gregorio E. Vaccines, new opportunities for a new society. Proc Natl Acad Sci USA. 2014; 111:12288–12293. https://doi.org/10.1073/pnas.1402981111 [PubMed]
- 54. Bronson E, Deem SL, Sanchez C, Murray S. Serologic response to a canarypox-vectored canine distemper virus vaccine in the giant panda (Ailuropoda melanoleuca). J Zoo Wildl Med. 2007; 38:363–366. https://doi.org/10.1638/1042-7260(2007)038[0363:SRTACC]2.0.CO;2 [PubMed]
- 55. Wang C, Yang S, Wu K, Gao Y, Zhang Z, Luo L, Wang C, Wang T, Yan Y, Hu J, Yang Z, Lan J. Serological evaluation of the efficacy of the multivalent canine distemper attenuated live vaccines on giant pandas (Ailuropoda melanoleuca). ACTA Theriol Sin. 2008; 28:212–216.
- 56. Qin Q, Li D, Zhang H, Hou R, Zhang Z, Zhang C, Zhang J, Wei F. Serosurvey of selected viruses in captive giant pandas (Ailuropoda melanoleuca) in China. Vet Microbiol. 2010; 142:199–204. https://doi.org/10.1016/j.vetmic.2009.09.062 [PubMed]
- 57. Lord J. The effect of aging of the immune system on vaccination responses. Hum Vaccin Immunother. 2013; 9:1364–1367. https://doi.org/10.4161/hv.24696 [PubMed]
- 58. Weng N. Aging of the immune system: how much can the adaptive immune system adapt? Immunity. 2006; 24:495–499. https://doi.org/10.1016/j.immuni.2006.05.001 [PubMed]
- 59. Patel RK and Jain M. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS ONE. 2012; 7:e30619. https://doi.org/10.1371/journal.pone.0030619 [PubMed]
- 60. Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015; 12:357–360. https://doi.org/10.1038/nmeth.3317 [PubMed]
- 61. Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol. 2015; 33:290–295. https://doi.org/10.1038/nbt.3122 [PubMed]
- 62. Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012; 7:562–578. https://doi.org/10.1038/nprot.2012.016 [PubMed]
- 63. Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997; 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389 [PubMed]
- 64. Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics. 2005; 21:3674–3676. https://doi.org/10.1093/bioinformatics/bti610 [PubMed]
- 65. Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014; 15:550. https://doi.org/10.1186/s13059-014-0550-8 [PubMed]
- 66. Yu G, Wang L, Han Y, He Q. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012; 16:284–287. https://doi.org/10.1089/omi.2011.0118 [PubMed]
- 67. Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li C, Wei L. KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011; 39:W316–W322. https://doi.org/10.1093/nar/gkr483 [PubMed]
- 68. Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, von Mering C. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017; 45:D362–D368. https://doi.org/10.1093/nar/gkw937 [PubMed]
- 69. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003; 13:2498–2504. https://doi.org/10.1101/gr.1239303 [PubMed]