Breast cancer risk spans a wide range
Individual risk for developing breast cancer varies between 11.6% for women without specific clinical risk factors (i.e., average risk) and 85% for women with pathogenic germline mutations in highly penetrant genes (i.e., BRCA1, BRCA2, TP53, and PTEN) [1–6]. Assessment of individual risk is critical for tailoring screening and prevention strategies appropriate to the severity of risk, and therefore avoid unnecessary screening and over-treatment. Women at average risk can delay initiation of screening as recommended by both the United States Preventative Services Task Force and the American Cancer Society [7,8]. Women at moderate risk can begin annual screening earlier and should consider FDA-approved chemoprevention, such as tamoxifen, raloxifene or aromatase inhibitors [9]. Women at highest risk are candidates for aggressive screening (e.g., with breast MRI) or surgical prevention [10–13].
Limitations of current risk assessment models frequently used in the clinic
A number of models are available for estimation of individual breast cancer risk based on clinical factors such as family history, reproductive profile, history of prior breast biopsy, and breast density (Table 1). The most commonly used clinical models are the Gail [14,15], the Claus [16], and the International Breast Cancer Intervention Study (IBIS) models [17]. For an excellent and comprehensive discussion of all available clinical models (e.g., hereditary, etc.) see the 2017 Cintolo-Gonzalez review [18,19]. The Gail model uses reproductive and biopsy information but only a limited family history (mother or sister with breast cancer) to calculate risk. This model is validated and classifies subsequent breast cancer cases modestly well, with estimates of the area under the receiver-operating characteristic curve (AUC) of 0.45-0.74 [15,20–22]. For risk calculations see https://bcrisktool.cancer.gov. The Claus model uses first- and second-degree family history to calculate risk but does not consider additional family history and other risk factors (such as hormonal factors or biopsy history). This model has an estimated AUC of 0.72 [20]. For risk calculations see CancerGene (https://cagene.com/) [23]. The IBIS model uses reproductive history, biopsy history, family history and body mass index (BMI). The IBIS model also includes a more extensive assessment of family history, characterizing breast cancers in both first- and second-degree relatives and the age at which they were diagnosed. The AUC of the IBIS model ranges between 0.54 – 0.76, depending on the population assessed [20,22,24–28]. For risk calculations see http://www.ems-trials.org/riskevaluator/. See Table 1 for a more complete review of factors included in each model and the discriminatory accuracy in both general and high-risk populations.
Table 1. Comparison of commonly used clinical breast cancer risk assessment models: risk factors considered and discriminatory accuracy in independent datasets.
| Model | Gail | Claus | IBIS (Tyrer-Cuzick) | BCSC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Model version | 2[15,105–108] | 1[16,109] | 6.0.0[17] | 7.0.2 | 8.0 | 2.0[40] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Personal | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age | Xa | X | X | X | X | Xa | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BMI | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Race/ethnicity | X | X | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hormonal | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age at menarche | X | X | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Menopausal status | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Parity, age first birth | X | X | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HRT use | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Benign Breast Disease (BBD) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Num. breast biopsies | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BBD with LCIS | X | X | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BBD with atypia | X | X | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BBD without atypia | X | X | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Family history | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1° female relatives (breast) | Xb | X | X | X | X | Xb | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Extended family hx (breast) | X | Xc | Xc | Xc | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1° male family hx (breast) | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Family hx of ovarian cancer | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Genetic variants | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BRCA status | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Polygenic Risk Score (PRS) | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Breast density | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Breast cancer outcomes | Invasive | Invasive + DCIS | Invasive + DCIS | Invasive | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 5-yr risk | X | X | X | X | X | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| > 10-yr riskd | X | X | X | X | X | X | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General population (AUC) | 0.54-0.67 [26,27,106–108,110–116] | 0.57-0.695 [26,27] | 0.66 [50] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| High-risk women (AUC) | 0.45-0.735 [20,22] | 0.716 [20] | 0.51-0.762 [20,22,28] | 0.54 [24] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| a Model not applicable for women under age 35. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| b Ages of diagnoses not considered. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| c 1° and 2° female relatives, as well as selected 3° relatives (female first cousins), diagnosed with breast cancer. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| d Risk of developing breast cancer outcome by age 90 (Gail model); by age 79 (Claus model); within 10 years and by age 80 (IBIS model 6); to age 85 (IBIS models 7 and 8), and over a 10-yr age interval (BCSC model). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Newer clinical models such as the Breast Cancer Surveillance Consortium (BCSC) model and updated/revised versions of the IBIS model (version 8) have incorporated mammographic density (MD) into assessment of risk. Mammographic density is a strong, independent risk factor for breast cancer development with studies showing a 4-6-fold increased risk for breast cancer for women with the highest breast density category compared with women in the lowest breast density category [29–38]. The BCSC model also incorporates reproductive factors, first-degree family-history, and recently added biopsy history to its set of predictors [39,40]. This model is validated and classifies breast cancer incidence with an AUC of 0.67 [39,41]. Accuracy of the latest version of the IBIS model has not been assessed.
Given that an AUC of 0.5 suggests that the test (or model in this case) performs no better than chance, the fact that none of the above models have an AUC greater than 0.76 leaves room for improvement [22,42,43]. There is, therefore, a significant need for more precise risk assessment. Recent advances in genetics have improved our ability to assess risk at the individual level. Genome-wide association studies have identified >100 single nucleotide polymorphisms (SNPs) associated with breast cancer risk [44–47] and polygenic risk scores (PRS) have been developed by several groups using this information [48,49]. Case-control studies have demonstrated the ability of PRS to accurately categorize risk (with AUC ranging from 0.59 – 0.65) [50–52]. However, risk associated with any of the developed polygenic risk scores needs to be interpreted with caution as their predictive capacity has not been validated outside of the populations in which they were developed. As seen with genetic testing, this may limit generalizability [53]. Several groups have examined whether use of PRS improves accuracy of currently available clinical models and demonstrated AUCs between 0.62 and 0.70 [41,54–59]. The ability of PRS to improve current clinical models is under prospective evaluation in the WISDOM trial [60,61]. Given that many of the SNPs included in polygenic risk scores are likely associated with hereditary risk, caution should be used when adding genetic factors to family history-based models without accounting for joint influences on model fit. The ability of a PRS to improve risk assessment is promising; however, utility in genetically and ethnically diverse populations must be studied.
Use of circulating miRNA biomarkers augment clinical tools to provide personalized risk assessment
Novel classes of biomarkers, such as circulating microRNA (C-miRNA) have emerged as promising cancer biomarkers [62–65] and may provide additional risk information. MicroRNAs (miRNA) are short, non-coding RNAs that bind to target mRNA and inhibit protein expression to regulate cellular processes such as proliferation, differentiation, and apoptosis [66–68]. A single miRNA can simultaneously target hundreds of genes, acting as a master regulator of entire biological pathways, with established roles in controlling normal development and tissue homeostasis [69–71]. Aberrant expression of miRNAs has been shown to regulate cancer cell activity by modulating oncogenic or tumor suppressor pathways to promote disease onset and progression [70]. In addition, miRNAs circulate, acting as intercellular signaling molecules, and may function to establish local and systemic environments for initiation and/or progression of cancer. Circulating miRNAs (C-miRNAs) are released from almost all cells in a variety of forms: in microvesicles [72], exosomes [73], bound to protein or lipid particles [74,75] or as free species [69]. Importantly, miRNAs are readily detectable, stable in circulation and found in most body fluids (e.g. blood, urine) [62,76], all characteristics of an ideal biomarker. The importance of standardized analysis of C-miRNA has become increasingly recognized by our group and others as essential for generating reproducible and actionable results [77–79].
In breast cancer patients, the presence of miRNA in circulation correlates with expression of that miRNA in primary breast tumors [80–82]. Additionally, significant differences in specific C-miRNA have been found between cancer patients and healthy controls [65,83–86], suggesting potential clinical utility for cancer detection [64,81,82,87–93]. For cancer risk assessment a biomarker must predict disease status with acceptable specificity and sensitivity [94]. To date, only a handful of studies have evaluated the utility of C-miRNA in cancer risk assessment. For example, several studies have evaluated miRNAs associated with risk for colon cancer and identified miRNAs associated with a pre-neoplastic colon lesion [95–99]. An independent and larger study identified a panel of 3 C-miRNAs as a promising colon cancer risk biomarker [100]. Other studies have discovered a number of miRNAs dysregulated in women <18 months from a breast cancer diagnosis, consistent with early detection [101–103]. Taken together, these data suggest that it is feasible that C-miRNAs can provide a signature of breast cancer risk with actionable lead-time for prevention.
Our group recently identified a C-miRNA-based risk signature predictive of long-term risk in a prospective cohort of women at increased risk for developing breast cancer. This IRB-approved prospective cohort includes over 600 high-risk women (who have signed informed consent) with a median follow-up of 8.9 years. From this cohort we selected 24 invasive breast cancer cases, to whom we matched controls on age, reason for high-risk status (e.g., strong family history of breast cancer or benign breast disease), and follow-up time. The median age at blood draw was 55.4 (range 33.9-77.5) for affected cases and 55.1 (range 32.8-78.4) for cancer-free controls (see Table 1: Subject characteristics in Oncotarget [104] for complete cohort clinical characteristics). RNA was isolated from banked serum, and profiled for over 2500 mature human miRNAs. The full Affymetrix GeneChip miRNA v4 (miRbase v20) microarray expression dataset is freely available in GEO Datasets (GSE98181, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE98181) and the R scripts used for data analysis accompany our open access 2017 Oncotarget manuscript as a supplement [104]. We identified 25 C-miRNAs that were significantly differentially expressed between cases and controls. From these 25 miRNAs, we discovered a group of 6 C-miRNAs that together discriminated cases from non-cases with high accuracy (AUC=0.896) (Figure 1). For the women who developed cancer in this cohort, blood had been banked a median of 3.2 years (range 0.6-8.7) prior to diagnosis, making this clearly a signature associated with risk and not early detection [104]. Refinement and validation of this risk signature is ongoing, using banked samples from previously performed randomized clinical trials. The validation of a sensitive and specific, non-invasive C-miRNA risk assessment tool will arm clinicians with vastly improved individualized risk estimates for patients, relevant to both young and older women. These risk estimates can be used to guide selection of the most appropriate screening and prevention options for a given individual. Information from miRNA expression will also provide valuable insight into the underlying biology of breast cancer initiation and may provide targets for chemoprevention.
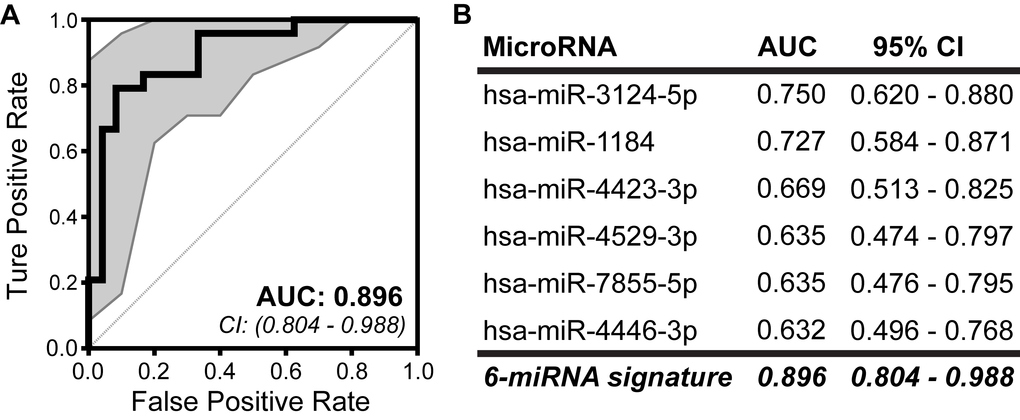
Figure 1. Development of a predictive miRNA signature for breast cancer risk among high-risk women. The predictive ability of A) the 6-miRNA risk signature and B) each individual C-miRNA was assessed by ROC curve and AUC based on calculated risk score. The combined expression of the 6 C-miRNAs discriminate cases from controls with increased accuracy and precision than any single miRNA. 95% confidence intervals (CI) are indicated by gray area around each curve. Modified from our 2017 Oncotarget publication [104].
Personalized and precise risk assessment can identify those women at greatest risk to develop breast cancer, thus avoiding overtreatment of women at lower/average risk and identifying women at high risk who would be candidates for high risk screening, chemoprevention or surgical prevention. Progress has been made towards personalized risk assessment and some promising new markers have been identified. However, rigorous validation of the most promising markers, and the predictive models they contribute to, in relevant populations is necessary before deployment for clinical use.
Acknowledgments
The authors appreciate the support of the following University of Vermont Cancer Center Cores: Clinical and Translational Research Laboratory (Jon Ramsey) and Vermont Integrated Genomics Resource (Julie Dragon, Nelson Vila-Santana, Meghann Palermo, Tim Hunter). We would like to express our gratitude to the women who contributed clinical data and samples.
Conflicts of Interest
The authors declare no potential conflict of interests.
Funding
This publication was supported by a University of Vermont Cancer Center pilot award to MEW and JBL and a J Walter Juckett Fellowship to NHF from the Lake Champlain Cancer Research Organization. Support was also provided by the following grants: National Cancer Institute (P01 CA082834 and U01 CA196383), Department of Defense (W81XWH-14-0468), National Institute of General Medical Sciences (P20 GM103644 and U54 GM115516 (Northern New England Clinical and Translational Research pilot award)), and the Charlotte Perelman Fund for Cancer Research.
References
- 1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492 [PubMed]
- 2. Kuchenbaecker KB, Hopper JL, Barnes DR, Phillips KA, Mooij TM, Roos-Blom MJ, Jervis S, van Leeuwen FE, Milne RL, Andrieu N, Goldgar DE, Terry MB, Rookus MA, et al, and BRCA1 and BRCA2 Cohort Consortium. Risks of Breast, Ovarian, and Contralateral Breast Cancer for BRCA1 and BRCA2 Mutation Carriers. JAMA. 2017; 317:2402–16. https://doi.org/10.1001/jama.2017.7112 [PubMed]
- 3. Tan MH, Mester JL, Ngeow J, Rybicki LA, Orloff MS, Eng C. Lifetime cancer risks in individuals with germline PTEN mutations. Clin Cancer Res. 2012; 18:400–07. https://doi.org/10.1158/1078-0432.CCR-11-2283 [PubMed]
- 4. Gonzalez KD, Noltner KA, Buzin CH, Gu D, Wen-Fong CY, Nguyen VQ, Han JH, Lowstuter K, Longmate J, Sommer SS, Weitzel JN. Beyond Li Fraumeni Syndrome: clinical characteristics of families with p53 germline mutations. J Clin Oncol. 2009; 27:1250–56. https://doi.org/10.1200/JCO.2008.16.6959 [PubMed]
- 5. Hearle N, Schumacher V, Menko FH, Olschwang S, Boardman LA, Gille JJ, Keller JJ, Westerman AM, Scott RJ, Lim W, Trimbath JD, Giardiello FM, Gruber SB, et al. Frequency and spectrum of cancers in the Peutz-Jeghers syndrome. Clin Cancer Res. 2006; 12:3209–15. https://doi.org/10.1158/1078-0432.CCR-06-0083 [PubMed]
- 6. Pharoah PD, Guilford P, Caldas C, and International Gastric Cancer Linkage Consortium. Incidence of gastric cancer and breast cancer in CDH1 (E-cadherin) mutation carriers from hereditary diffuse gastric cancer families. Gastroenterology. 2001; 121:1348–53. https://doi.org/10.1053/gast.2001.29611 [PubMed]
- 7. Nelson HD, Pappas M, Zakher B, Mitchell JP, Okinaka-Hu L, Fu R. Risk assessment, genetic counseling, and genetic testing for BRCA-related cancer in women: a systematic review to update the U.S. Preventive Services Task Force recommendation. Ann Intern Med. 2014; 160:255–66. https://doi.org/10.7326/M13-1684 [PubMed]
- 8. Oeffinger KC, Fontham ET, Etzioni R, Herzig A, Michaelson JS, Shih YC, Walter LC, Church TR, Flowers CR, LaMonte SJ, Wolf AM, DeSantis C, Lortet-Tieulent J, et al, and American Cancer Society. Breast Cancer Screening for Women at Average Risk: 2015 Guideline Update From the American Cancer Society. JAMA. 2015; 314:1599–614. https://doi.org/10.1001/jama.2015.12783 [PubMed]
- 9. Cuzick J, DeCensi A, Arun B, Brown PH, Castiglione M, Dunn B, Forbes JF, Glaus A, Howell A, von Minckwitz G, Vogel V, Zwierzina H. Preventive therapy for breast cancer: a consensus statement. Lancet Oncol. 2011; 12:496–503. https://doi.org/10.1016/S1470-2045(11)70030-4 [PubMed]
- 10. Hartmann LC, Lindor NM. The Role of Risk-Reducing Surgery in Hereditary Breast and Ovarian Cancer. N Engl J Med. 2016; 374:454–68. https://doi.org/10.1056/NEJMra1503523 [PubMed]
- 11. Saslow D, Boetes C, Burke W, Harms S, Leach MO, Lehman CD, Morris E, Pisano E, Schnall M, Sener S, Smith RA, Warner E, Yaffe M, et al, and American Cancer Society Breast Cancer Advisory Group. American Cancer Society guidelines for breast screening with MRI as an adjunct to mammography. CA Cancer J Clin. 2007; 57:75–89. https://doi.org/10.3322/canjclin.57.2.75 [PubMed]
- 12. Kauff ND, Satagopan JM, Robson ME, Scheuer L, Hensley M, Hudis CA, Ellis NA, Boyd J, Borgen PI, Barakat RR, Norton L, Castiel M, Nafa K, Offit K. Risk-reducing salpingo-oophorectomy in women with a BRCA1 or BRCA2 mutation. N Engl J Med. 2002; 346:1609–15. https://doi.org/10.1056/NEJMoa020119 [PubMed]
- 13. Rebbeck TR, Levin AM, Eisen A, Snyder C, Watson P, Cannon-Albright L, Isaacs C, Olopade O, Garber JE, Godwin AK, Daly MB, Narod SA, Neuhausen SL, et al. Breast cancer risk after bilateral prophylactic oophorectomy in BRCA1 mutation carriers. J Natl Cancer Inst. 1999; 91:1475–79. https://doi.org/10.1093/jnci/91.17.1475 [PubMed]
- 14. Gail MH, Brinton LA, Byar DP, Corle DK, Green SB, Schairer C, Mulvihill JJ. Projecting individualized probabilities of developing breast cancer for white females who are being examined annually. J Natl Cancer Inst. 1989; 81:1879–86. https://doi.org/10.1093/jnci/81.24.1879 [PubMed]
- 15. Costantino JP, Gail MH, Pee D, Anderson S, Redmond CK, Benichou J, Wieand HS. Validation studies for models projecting the risk of invasive and total breast cancer incidence. J Natl Cancer Inst. 1999; 91:1541–48. https://doi.org/10.1093/jnci/91.18.1541 [PubMed]
- 16. Claus EB, Risch N, Thompson WD. Autosomal dominant inheritance of early-onset breast cancer. Implications for risk prediction. Cancer. 1994; 73:643–51. https://doi.org/10.1002/1097-0142(19940201)73:3<643::AID-CNCR2820730323>3.0.CO;2-5 [PubMed]
- 17. Tyrer J, Duffy SW, Cuzick J. A breast cancer prediction model incorporating familial and personal risk factors. Stat Med. 2004; 23:1111–30. https://doi.org/10.1002/sim.1668 [PubMed]
- 18. Cintolo-Gonzalez JA, Braun D, Blackford AL, Mazzola E, Acar A, Plichta JK, Griffin M, Hughes KS. Erratum to: Breast cancer risk models: a comprehensive overview of existing models, validation, and clinical applications. Breast Cancer Res Treat. 2017; 164:745. https://doi.org/10.1007/s10549-017-4349-7 [PubMed]
- 19. Cintolo-Gonzalez JA, Braun D, Blackford AL, Mazzola E, Acar A, Plichta JK, Griffin M, Hughes KS. Breast cancer risk models: a comprehensive overview of existing models, validation, and clinical applications. Breast Cancer Res Treat. 2017; 164:263–84. https://doi.org/10.1007/s10549-017-4247-z [PubMed]
- 20. Amir E, Evans DG, Shenton A, Lalloo F, Moran A, Boggis C, Wilson M, Howell A. Evaluation of breast cancer risk assessment packages in the family history evaluation and screening programme. J Med Genet. 2003; 40:807–14. https://doi.org/10.1136/jmg.40.11.807 [PubMed]
- 21. Meads C, Ahmed I, Riley RD. A systematic review of breast cancer incidence risk prediction models with meta-analysis of their performance. Breast Cancer Res Treat. 2012; 132:365–77. https://doi.org/10.1007/s10549-011-1818-2 [PubMed]
- 22. Boughey JC, Hartmann LC, Anderson SS, Degnim AC, Vierkant RA, Reynolds CA, Frost MH, Pankratz VS. Evaluation of the Tyrer-Cuzick (International Breast Cancer Intervention Study) model for breast cancer risk prediction in women with atypical hyperplasia. J Clin Oncol. 2010; 28:3591–96. https://doi.org/10.1200/JCO.2010.28.0784 [PubMed]
- 23. Euhus DM. Understanding mathematical models for breast cancer risk assessment and counseling. Breast J. 2001; 7:224–32. https://doi.org/10.1046/j.1524-4741.2001.20012.x [PubMed]
- 24. Lo LL, Milne RL, Liao Y, Cuzick J, Terry MB, Phillips KA. Validation of the IBIS breast cancer risk evaluator for women with lobular carcinoma in-situ. Br J Cancer. 2018; 119:36–39. https://doi.org/10.1038/s41416-018-0120-z [PubMed]
- 25. Jacobi CE, de Bock GH, Siegerink B, van Asperen CJ. Differences and similarities in breast cancer risk assessment models in clinical practice: which model to choose? Breast Cancer Res Treat. 2009; 115:381–90. https://doi.org/10.1007/s10549-008-0070-x [PubMed]
- 26. Quante AS, Whittemore AS, Shriver T, Strauch K, Terry MB. Breast cancer risk assessment across the risk continuum: genetic and nongenetic risk factors contributing to differential model performance. Breast Cancer Res. 2012; 14:R144. https://doi.org/10.1186/bcr3352 [PubMed]
- 27. Brentnall AR, Harkness EF, Astley SM, Donnelly LS, Stavrinos P, Sampson S, Fox L, Sergeant JC, Harvie MN, Wilson M, Beetles U, Gadde S, Lim Y, et al. Mammographic density adds accuracy to both the Tyrer-Cuzick and Gail breast cancer risk models in a prospective UK screening cohort. Breast Cancer Res. 2015; 17:147. https://doi.org/10.1186/s13058-015-0653-5 [PubMed]
- 28. Warwick J, Birke H, Stone J, Warren RM, Pinney E, Brentnall AR, Duffy SW, Howell A, Cuzick J. Mammographic breast density refines Tyrer-Cuzick estimates of breast cancer risk in high-risk women: findings from the placebo arm of the International Breast Cancer Intervention Study I. Breast Cancer Res. 2014; 16:451. https://doi.org/10.1186/s13058-014-0451-5 [PubMed]
- 29. Harvey JA, Bovbjerg VE. Quantitative assessment of mammographic breast density: relationship with breast cancer risk. Radiology. 2004; 230:29–41. https://doi.org/10.1148/radiol.2301020870 [PubMed]
- 30. McCormack VA, dos Santos Silva I. Breast density and parenchymal patterns as markers of breast cancer risk: a meta-analysis. Cancer Epidemiol Biomarkers Prev. 2006; 15:1159–69. https://doi.org/10.1158/1055-9965.EPI-06-0034 [PubMed]
- 31. Boyd NF, Byng JW, Jong RA, Fishell EK, Little LE, Miller AB, Lockwood GA, Tritchler DL, Yaffe MJ. Quantitative classification of mammographic densities and breast cancer risk: results from the Canadian National Breast Screening Study. J Natl Cancer Inst. 1995; 87:670–75. https://doi.org/10.1093/jnci/87.9.670 [PubMed]
- 32. Vacek PM, Geller BM. A prospective study of breast cancer risk using routine mammographic breast density measurements. Cancer Epidemiol Biomarkers Prev. 2004; 13:715–22. [PubMed]
- 33. Yaffe MJ, Boyd NF, Byng JW, Jong RA, Fishell E, Lockwood GA, Little LE, Tritchler DL. Breast cancer risk and measured mammographic density. Eur J Cancer Prev. 1998 (Suppl 1); 7:S47–55. https://doi.org/10.1097/00008469-199802001-00010 [PubMed]
- 34. Lam PB, Vacek PM, Geller BM, Muss HB. The association of increased weight, body mass index, and tissue density with the risk of breast carcinoma in Vermont. Cancer. 2000; 89:369–75. https://doi.org/10.1002/1097-0142(20000715)89:2<369::AID-CNCR23>3.0.CO;2-J [PubMed]
- 35. Ursin G, Ma H, Wu AH, Bernstein L, Salane M, Parisky YR, Astrahan M, Siozon CC, Pike MC. Mammographic density and breast cancer in three ethnic groups. Cancer Epidemiol Biomarkers Prev. 2003; 12:332–38. [PubMed]
- 36. Torres-Mejía G, De Stavola B, Allen DS, Pérez-Gavilán JJ, Ferreira JM, Fentiman IS, Dos Santos Silva I. Mammographic features and subsequent risk of breast cancer: a comparison of qualitative and quantitative evaluations in the Guernsey prospective studies. Cancer Epidemiol Biomarkers Prev. 2005; 14:1052–59. https://doi.org/10.1158/1055-9965.EPI-04-0717 [PubMed]
- 37. Maskarinec G, Pagano I, Lurie G, Kolonel LN. A longitudinal investigation of mammographic density: the multiethnic cohort. Cancer Epidemiol Biomarkers Prev. 2006; 15:732–39. https://doi.org/10.1158/1055-9965.EPI-05-0798 [PubMed]
- 38. Warwick J, Pinney E, Warren RM, Duffy SW, Howell A, Wilson M, Cuzick J. Breast density and breast cancer risk factors in a high-risk population. Breast. 2003; 12:10–16. https://doi.org/10.1016/S0960-9776(02)00212-6 [PubMed]
- 39. Tice JA, Cummings SR, Smith-Bindman R, Ichikawa L, Barlow WE, Kerlikowske K. Using clinical factors and mammographic breast density to estimate breast cancer risk: development and validation of a new predictive model. Ann Intern Med. 2008; 148:337–47. https://doi.org/10.7326/0003-4819-148-5-200803040-00004 [PubMed]
- 40. Tice JA, Miglioretti DL, Li CS, Vachon CM, Gard CC, Kerlikowske K. Breast Density and Benign Breast Disease: Risk Assessment to Identify Women at High Risk of Breast Cancer. J Clin Oncol. 2015; 33:3137–43. https://doi.org/10.1200/JCO.2015.60.8869 [PubMed]
- 41. Vachon CM, Pankratz VS, Scott CG, Haeberle L, Ziv E, Jensen MR, Brandt KR, Whaley DH, Olson JE, Heusinger K, Hack CC, Jud SM, Beckmann MW, et al. The contributions of breast density and common genetic variation to breast cancer risk. J Natl Cancer Inst. 2015; 107. https://doi.org/10.1093/jnci/dju397 [PubMed]
- 42. Euhus DM, Leitch AM, Huth JF, Peters GN. Limitations of the Gail model in the specialized breast cancer risk assessment clinic. Breast J. 2002; 8:23–27. https://doi.org/10.1046/j.1524-4741.2002.08005.x [PubMed]
- 43. Sakorafas GH, Krespis E, Pavlakis G. Risk estimation for breast cancer development; a clinical perspective. Surg Oncol. 2002; 10:183–92. https://doi.org/10.1016/S0960-7404(02)00016-6 [PubMed]
- 44. Michailidou K, Beesley J, Lindstrom S, Canisius S, Dennis J, Lush MJ, Maranian MJ, Bolla MK, Wang Q, Shah M, Perkins BJ, Czene K, Eriksson M, et al, and BOCS, and kConFab Investigators, and AOCS Group, and NBCS, and GENICA Network. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat Genet. 2015; 47:373–80. https://doi.org/10.1038/ng.3242 [PubMed]
- 45. Michailidou K, Hall P, Gonzalez-Neira A, Ghoussaini M, Dennis J, Milne RL, Schmidt MK, Chang-Claude J, Bojesen SE, Bolla MK, Wang Q, Dicks E, Lee A, et al, and Breast and Ovarian Cancer Susceptibility Collaboration, and Hereditary Breast and Ovarian Cancer Research Group Netherlands (HEBON), and kConFab Investigators, and Australian Ovarian Cancer Study Group, and GENICA (Gene Environment Interaction and Breast Cancer in Germany) Network. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat Genet. 2013; 45:353–61, e1–2. https://doi.org/10.1038/ng.2563 [PubMed]
- 46. Ghoussaini M, Fletcher O, Michailidou K, Turnbull C, Schmidt MK, Dicks E, Dennis J, Wang Q, Humphreys MK, Luccarini C, Baynes C, Conroy D, Maranian M, et al, and Netherlands Collaborative Group on Hereditary Breast and Ovarian Cancer (HEBON), and Familial Breast Cancer Study (FBCS), and Gene Environment Interaction of Breast Cancer in Germany (GENICA) Network, and kConFab Investigators, and Australian Ovarian Cancer Study Group. Genome-wide association analysis identifies three new breast cancer susceptibility loci. Nat Genet. 2012; 44:312–18. https://doi.org/10.1038/ng.1049 [PubMed]
- 47. Turnbull C, Ahmed S, Morrison J, Pernet D, Renwick A, Maranian M, Seal S, Ghoussaini M, Hines S, Healey CS, Hughes D, Warren-Perry M, Tapper W, et al, and Breast Cancer Susceptibility Collaboration (UK). Genome-wide association study identifies five new breast cancer susceptibility loci. Nat Genet. 2010; 42:504–07. https://doi.org/10.1038/ng.586 [PubMed]
- 48. Rudolph A, Chang-Claude J, Schmidt MK. Gene-environment interaction and risk of breast cancer. Br J Cancer. 2016; 114:125–33. https://doi.org/10.1038/bjc.2015.439 [PubMed]
- 49. Campa D, Kaaks R, Le Marchand L, Haiman CA, Travis RC, Berg CD, Buring JE, Chanock SJ, Diver WR, Dostal L, Fournier A, Hankinson SE, Henderson BE, et al. Interactions between genetic variants and breast cancer risk factors in the breast and prostate cancer cohort consortium. J Natl Cancer Inst. 2011; 103:1252–63. https://doi.org/10.1093/jnci/djr265 [PubMed]
- 50. Mavaddat N, Pharoah PD, Michailidou K, Tyrer J, Brook MN, Bolla MK, Wang Q, Dennis J, Dunning AM, Shah M, Luben R, Brown J, Bojesen SE, et al. Prediction of breast cancer risk based on profiling with common genetic variants. J Natl Cancer Inst. 2015; 107. https://doi.org/10.1093/jnci/djv036 [PubMed]
- 51. Sawyer S, Mitchell G, McKinley J, Chenevix-Trench G, Beesley J, Chen XQ, Bowtell D, Trainer AH, Harris M, Lindeman GJ, James PA. A role for common genomic variants in the assessment of familial breast cancer. J Clin Oncol. 2012; 30:4330–36. https://doi.org/10.1200/JCO.2012.41.7469 [PubMed]
- 52. Li H, Feng B, Miron A, Chen X, Beesley J, Bimeh E, Barrowdale D, John EM, Daly MB, Andrulis IL, Buys SS, Kraft P, Thorne H, et al, and kConFab investigators. Breast cancer risk prediction using a polygenic risk score in the familial setting: a prospective study from the Breast Cancer Family Registry and kConFab. Genet Med. 2017; 19:30–35. https://doi.org/10.1038/gim.2016.43 [PubMed]
- 53. Manrai AK, Funke BH, Rehm HL, Olesen MS, Maron BA, Szolovits P, Margulies DM, Loscalzo J, Kohane IS. Genetic Misdiagnoses and the Potential for Health Disparities. N Engl J Med. 2016; 375:655–65. https://doi.org/10.1056/NEJMsa1507092 [PubMed]
- 54. Vachon CM, Schaid DJ, Ingle JN, Wickerham DL, Kubo M, Mushiroda T, Goetz MP, Carlson EE, Paik S, Wolmark N, Nakamura Y, Wang L, Weinshilboum R, Couch FJ. A polygenic risk score for breast cancer in women receiving tamoxifen or raloxifene on NSABP P-1 and P-2. Breast Cancer Res Treat. 2015; 149:517–23. https://doi.org/10.1007/s10549-014-3175-4 [PubMed]
- 55. Dite GS, MacInnis RJ, Bickerstaffe A, Dowty JG, Allman R, Apicella C, Milne RL, Tsimiklis H, Phillips KA, Giles GG, Terry MB, Southey MC, Hopper JL. Breast Cancer Risk Prediction Using Clinical Models and 77 Independent Risk-Associated SNPs for Women Aged Under 50 Years: Australian Breast Cancer Family Registry. Cancer Epidemiol Biomarkers Prev. 2016; 25:359–65. https://doi.org/10.1158/1055-9965.EPI-15-0838 [PubMed]
- 56. Ziv E, Tice JA, Sprague B, Vachon CM, Cummings SR, Kerlikowske K. Using Breast Cancer Risk Associated Polymorphisms to Identify Women for Breast Cancer Chemoprevention. PLoS One. 2017; 12:e0168601. https://doi.org/10.1371/journal.pone.0168601 [PubMed]
- 57. Gail MH. Discriminatory accuracy from single-nucleotide polymorphisms in models to predict breast cancer risk. J Natl Cancer Inst. 2008; 100:1037–41. https://doi.org/10.1093/jnci/djn180 [PubMed]
- 58. Darabi H, Czene K, Zhao W, Liu J, Hall P, Humphreys K. Breast cancer risk prediction and individualised screening based on common genetic variation and breast density measurement. Breast Cancer Res. 2012; 14:R25. https://doi.org/10.1186/bcr3110 [PubMed]
- 59. Garcia-Closas M, Gunsoy NB, Chatterjee N. Combined associations of genetic and environmental risk factors: implications for prevention of breast cancer. J Natl Cancer Inst. 2014; 106:106. https://doi.org/10.1093/jnci/dju305 [PubMed]
- 60. Shieh Y, Hu D, Ma L, Huntsman S, Gard CC, Leung JW, Tice JA, Vachon CM, Cummings SR, Kerlikowske K, Ziv E. Breast cancer risk prediction using a clinical risk model and polygenic risk score. Breast Cancer Res Treat. 2016; 159:513–25. https://doi.org/10.1007/s10549-016-3953-2 [PubMed]
- 61. Shieh Y, Eklund M, Madlensky L, Sawyer SD, Thompson CK, Stover Fiscalini A, Ziv E, Van’t Veer LJ, Esserman LJ, Tice JA. Athena Breast Health Network I. Breast Cancer Screening in the Precision Medicine Era: Risk-Based Screening in a Population-Based Trial. J Natl Cancer Inst. 2017; 109. https://doi.org/10.1093/jnci/djw290 [PubMed]
- 62. Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, Galas DJ, Wang K. The microRNA spectrum in 12 body fluids. Clin Chem. 2010; 56:1733–41. https://doi.org/10.1373/clinchem.2010.147405 [PubMed]
- 63. He Y, Lin J, Kong D, Huang M, Xu C, Kim TK, Etheridge A, Luo Y, Ding Y, Wang K. Current State of Circulating MicroRNAs as Cancer Biomarkers. Clin Chem. 2015; 61:1138–55. https://doi.org/10.1373/clinchem.2015.241190 [PubMed]
- 64. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A, Lin DW, Urban N, Drescher CW, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008; 105:10513–18. https://doi.org/10.1073/pnas.0804549105 [PubMed]
- 65. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008; 18:997–1006. https://doi.org/10.1038/cr.2008.282 [PubMed]
- 66. Garber JE, Offit K. Hereditary cancer predisposition syndromes. J Clin Oncol. 2005; 23:276–92. https://doi.org/10.1200/JCO.2005.10.042 [PubMed]
- 67. Taft RJ, Pang KC, Mercer TR, Dinger M, Mattick JS. Non-coding RNAs: regulators of disease. J Pathol. 2010; 220:126–39. https://doi.org/10.1002/path.2638 [PubMed]
- 68. Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009; 10:704–14. https://doi.org/10.1038/nrg2634 [PubMed]
- 69. Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA. MicroRNAs in body fluids--the mix of hormones and biomarkers. Nat Rev Clin Oncol. 2011; 8:467–77. https://doi.org/10.1038/nrclinonc.2011.76 [PubMed]
- 70. Shah MY, Ferrajoli A, Sood AK, Lopez-Berestein G, Calin GA. microRNA Therapeutics in Cancer - An Emerging Concept. EBioMedicine. 2016; 12:34–42. https://doi.org/10.1016/j.ebiom.2016.09.017 [PubMed]
- 71. Kunej T, Godnic I, Horvat S, Zorc M, Calin GA. Cross talk between microRNA and coding cancer genes. Cancer J. 2012; 18:223–31. https://doi.org/10.1097/PPO.0b013e318258b771 [PubMed]
- 72. Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem. 2010; 285:17442–52. https://doi.org/10.1074/jbc.M110.107821 [PubMed]
- 73. Denzer K, Kleijmeer MJ, Heijnen HF, Stoorvogel W, Geuze HJ. Exosome: from internal vesicle of the multivesicular body to intercellular signaling device. J Cell Sci. 2000; 113:3365–74. [PubMed]
- 74. Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol. 2011; 13:423–33. https://doi.org/10.1038/ncb2210 [PubMed]
- 75. Wang K, Zhang S, Weber J, Baxter D, Galas DJ. Export of microRNAs and microRNA-protective protein by mammalian cells. Nucleic Acids Res. 2010; 38:7248–59. https://doi.org/10.1093/nar/gkq601 [PubMed]
- 76. Tiberio P, Callari M, Angeloni V, Daidone MG, Appierto V. Challenges in using circulating miRNAs as cancer biomarkers. BioMed Res Int. 2015; 2015:731479. https://doi.org/10.1155/2015/731479 [PubMed]
- 77. Farina NH, Wood ME, Perrapato SD, Francklyn CS, Stein GS, Stein JL, Lian JB. Standardizing analysis of circulating microRNA: clinical and biological relevance. J Cell Biochem. 2014; 115:805–11. https://doi.org/10.1002/jcb.24745 [PubMed]
- 78. Moldovan L, Batte KE, Trgovcich J, Wisler J, Marsh CB, Piper M. Methodological challenges in utilizing miRNAs as circulating biomarkers. J Cell Mol Med. 2014; 18:371–90. https://doi.org/10.1111/jcmm.12236 [PubMed]
- 79. Vigneron N, Meryet-Figuière M, Guttin A, Issartel JP, Lambert B, Briand M, Louis MH, Vernon M, Lebailly P, Lecluse Y, Joly F, Krieger S, Lheureux S, et al. Towards a new standardized method for circulating miRNAs profiling in clinical studies: interest of the exogenous normalization to improve miRNA signature accuracy. Mol Oncol. 2016; 10:981–92. https://doi.org/10.1016/j.molonc.2016.03.005 [PubMed]
- 80. Das SG, Romagnoli M, Mineva ND, Barillé-Nion S, Jézéquel P, Campone M, Sonenshein GE. miR-720 is a downstream target of an ADAM8-induced ERK signaling cascade that promotes the migratory and invasive phenotype of triple-negative breast cancer cells. Breast Cancer Res. 2016; 18:40. https://doi.org/10.1186/s13058-016-0699-z [PubMed]
- 81. Schrauder MG, Strick R, Schulz-Wendtland R, Strissel PL, Kahmann L, Loehberg CR, Lux MP, Jud SM, Hartmann A, Hein A, Bayer CM, Bani MR, Richter S, et al. Circulating micro-RNAs as potential blood-based markers for early stage breast cancer detection. PLoS One. 2012; 7:e29770. https://doi.org/10.1371/journal.pone.0029770 [PubMed]
- 82. Zhao H, Shen J, Medico L, Wang D, Ambrosone CB, Liu S. A pilot study of circulating miRNAs as potential biomarkers of early stage breast cancer. PLoS One. 2010; 5:e13735. https://doi.org/10.1371/journal.pone.0013735 [PubMed]
- 83. van Schooneveld E, Wouters MC, Van der Auwera I, Peeters DJ, Wildiers H, Van Dam PA, Vergote I, Vermeulen PB, Dirix LY, Van Laere SJ. Expression profiling of cancerous and normal breast tissues identifies microRNAs that are differentially expressed in serum from patients with (metastatic) breast cancer and healthy volunteers. Breast Cancer Res. 2012; 14:R34. https://doi.org/10.1186/bcr3127 [PubMed]
- 84. Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011; 57:84–91. https://doi.org/10.1373/clinchem.2010.151845 [PubMed]
- 85. Kodahl AR, Lyng MB, Binder H, Cold S, Gravgaard K, Knoop AS, Ditzel HJ. Novel circulating microRNA signature as a potential non-invasive multi-marker test in ER-positive early-stage breast cancer: a case control study. Mol Oncol. 2014; 8:874–83. https://doi.org/10.1016/j.molonc.2014.03.002 [PubMed]
- 86. Di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol. 2014; 9:287–314. https://doi.org/10.1146/annurev-pathol-012513-104715 [PubMed]
- 87. Healy NA, Heneghan HM, Miller N, Osborne CK, Schiff R, Kerin MJ. Systemic mirnas as potential biomarkers for malignancy. Int J Cancer. 2012; 131:2215–22. https://doi.org/10.1002/ijc.27642 [PubMed]
- 88. Madhavan D, Peng C, Wallwiener M, Zucknick M, Nees J, Schott S, Rudolph A, Riethdorf S, Trumpp A, Pantel K, Sohn C, Chang-Claude J, Schneeweiss A, Burwinkel B. Circulating miRNAs with prognostic value in metastatic breast cancer and for early detection of metastasis. Carcinogenesis. 2016; 37:461–70. https://doi.org/10.1093/carcin/bgw008 [PubMed]
- 89. Tozzoli R, D’Aurizio F, Falcomer F, Basso SM, Lumachi F. Serum Tumor Markers in Stage I-II Breast Cancer. Med Chem. 2016; 12:285–89. https://doi.org/10.2174/1573406412666151116144520 [PubMed]
- 90. Sochor M, Basova P, Pesta M, Dusilkova N, Bartos J, Burda P, Pospisil V, Stopka T. Oncogenic microRNAs: miR-155, miR-19a, miR-181b, and miR-24 enable monitoring of early breast cancer in serum. BMC Cancer. 2014; 14:448. https://doi.org/10.1186/1471-2407-14-448 [PubMed]
- 91. Mishra S, Srivastava AK, Suman S, Kumar V, Shukla Y. Circulating miRNAs revealed as surrogate molecular signatures for the early detection of breast cancer. Cancer Lett. 2015; 369:67–75. https://doi.org/10.1016/j.canlet.2015.07.045 [PubMed]
- 92. Lv M, Zhu X, Chen W, Zhao J, Tang J. Searching for candidate microRNA biomarkers in detection of breast cancer: a meta-analysis. Cancer Biomark. 2013; 13:395–401. https://doi.org/10.3233/CBM-130379 [PubMed]
- 93. Nassar FJ, Nasr R, Talhouk R. MicroRNAs as biomarkers for early breast cancer diagnosis, prognosis and therapy prediction. Pharmacol Ther. 2017; 172:34–49. https://doi.org/10.1016/j.pharmthera.2016.11.012 [PubMed]
- 94. Etheridge A, Lee I, Hood L, Galas D, Wang K. Extracellular microRNA: a new source of biomarkers. Mutat Res. 2011; 717:85–90. https://doi.org/10.1016/j.mrfmmm.2011.03.004 [PubMed]
- 95. Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010; 127:118–26. https://doi.org/10.1002/ijc.25007 [PubMed]
- 96. Wang Q, Huang Z, Ni S, Xiao X, Xu Q, Wang L, Huang D, Tan C, Sheng W, Du X. Plasma miR-601 and miR-760 are novel biomarkers for the early detection of colorectal cancer. PLoS One. 2012; 7:e44398. https://doi.org/10.1371/journal.pone.0044398 [PubMed]
- 97. Luo X, Stock C, Burwinkel B, Brenner H. Identification and evaluation of plasma microRNAs for early detection of colorectal cancer. PLoS One. 2013; 8:e62880. https://doi.org/10.1371/journal.pone.0062880 [PubMed]
- 98. Wang S, Xiang J, Li Z, Lu S, Hu J, Gao X, Yu L, Wang L, Wang J, Wu Y, Chen Z, Zhu H. A plasma microRNA panel for early detection of colorectal cancer. Int J Cancer. 2015; 136:152–61. https://doi.org/10.1002/ijc.28136 [PubMed]
- 99. Roberts BS, Hardigan AA, Moore DE, Ramaker RC, Jones AL, Fitz-Gerald MB, Cooper GM, Wilcox CM, Kimberly RP, Myers RM. Discovery and Validation of Circulating Biomarkers of Colorectal Adenoma by High-Depth Small RNA Sequencing. Clin Cancer Res. 2018; 24:2092–99. https://doi.org/10.1158/1078-0432.CCR-17-1960 [PubMed]
- 100. Yamada A, Horimatsu T, Okugawa Y, Nishida N, Honjo H, Ida H, Kou T, Kusaka T, Sasaki Y, Yagi M, Higurashi T, Yukawa N, Amanuma Y, et al. Serum miR-21, miR-29a, and miR-125b Are Promising Biomarkers for the Early Detection of Colorectal Neoplasia. Clin Cancer Res. 2015; 21:4234–42. https://doi.org/10.1158/1078-0432.CCR-14-2793 [PubMed]
- 101. Taslim C, Weng DY, Brasky TM, Dumitrescu RG, Huang K, Kallakury BV, Krishnan S, Llanos AA, Marian C, McElroy J, Schneider SS, Spear SL, Troester MA, et al. Discovery and replication of microRNAs for breast cancer risk using genome-wide profiling. Oncotarget. 2016; 7:86457–68. https://doi.org/10.18632/oncotarget.13241 [PubMed]
- 102. Zhang L, Xu Y, Jin X, Wang Z, Wu Y, Zhao D, Chen G, Li D, Wang X, Cao H, Xie Y, Liang Z. A circulating miRNA signature as a diagnostic biomarker for non-invasive early detection of breast cancer. Breast Cancer Res Treat. 2015; 154:423–34. https://doi.org/10.1007/s10549-015-3591-0 [PubMed]
- 103. Chang CW, Wu HC, Terry MB, Santella RM. microRNA Expression in Prospectively Collected Blood as a Potential Biomarker of Breast Cancer Risk in the BCFR. Anticancer Res. 2015; 35:3969–77. [PubMed]
- 104. Farina NH, Ramsey JE, Cuke ME, Ahern TP, Shirley DJ, Stein JL, Stein GS, Lian JB, Wood ME. Development of a predictive miRNA signature for breast cancer risk among high-risk women. Oncotarget. 2017; 8:112170–83. https://doi.org/10.18632/oncotarget.22750 [PubMed]
- 105. Anderson S, Ahnn S, Duff K. NSABP Breast Cancer Prevention Trial risk assessment program, version 2. NSABP Biostatistical Center Technical Report. Pittsburgh (PA): Department of Biostatistics, University of Pittsburgh; Aug 14, 1992.
- 106. Gail MH, Costantino JP, Pee D, Bondy M, Newman L, Selvan M, Anderson GL, Malone KE, Marchbanks PA, McCaskill-Stevens W, Norman SA, Simon MS, Spirtas R, et al. Projecting individualized absolute invasive breast cancer risk in African American women. J Natl Cancer Inst. 2007; 99:1782–92. https://doi.org/10.1093/jnci/djm223 [PubMed]
- 107. Banegas MP, John EM, Slattery ML, Gomez SL, Yu M, LaCroix AZ, Pee D, Chlebowski RT, Hines LM, Thompson CA, Gail MH. Projecting Individualized Absolute Invasive Breast Cancer Risk in US Hispanic Women. J Natl Cancer Inst. 2016; 109:215. https://doi.org/10.1093/jnci/djw215 [PubMed]
- 108. Matsuno RK, Costantino JP, Ziegler RG, Anderson GL, Li H, Pee D, Gail MH. Projecting individualized absolute invasive breast cancer risk in Asian and Pacific Islander American women. J Natl Cancer Inst. 2011; 103:951–61. https://doi.org/10.1093/jnci/djr154 [PubMed]
- 109. Claus EB, Risch N, Thompson WD. Genetic analysis of breast cancer in the cancer and steroid hormone study. Am J Hum Genet. 1991; 48:232–42. [PubMed]
- 110. Tice JA, Cummings SR, Ziv E, Kerlikowske K. Mammographic breast density and the Gail model for breast cancer risk prediction in a screening population. Breast Cancer Res Treat. 2005; 94:115–22. https://doi.org/10.1007/s10549-005-5152-4 [PubMed]
- 111. Tice JA, Cummings SR, Smith-Bindman R, Ichikawa L, Barlow WE, Kerlikowske K. Using clinical factors and mammographic breast density to estimate breast cancer risk: development and validation of a new predictive model. Ann Intern Med. 2008; 148:337–47. https://doi.org/10.7326/0003-4819-148-5-200803040-00004 [PubMed]
- 112. Decarli A, Calza S, Masala G, Specchia C, Palli D, Gail MH. Gail model for prediction of absolute risk of invasive breast cancer: independent evaluation in the Florence-European Prospective Investigation Into Cancer and Nutrition cohort. J Natl Cancer Inst. 2006; 98:1686–93. https://doi.org/10.1093/jnci/djj463 [PubMed]
- 113. Rockhill B, Spiegelman D, Byrne C, Hunter DJ, Colditz GA. Validation of the Gail et al. model of breast cancer risk prediction and implications for chemoprevention. J Natl Cancer Inst. 2001; 93:358–66. https://doi.org/10.1093/jnci/93.5.358 [PubMed]
- 114. Schonberg MA, Li VW, Eliassen AH, Davis RB, LaCroix AZ, McCarthy EP, Rosner BA, Chlebowski RT, Rohan TE, Hankinson SE, Marcantonio ER, Ngo LH. Performance of the Breast Cancer Risk Assessment Tool Among Women Age 75 Years and Older. J Natl Cancer Inst. 2015; 108. https://doi.org/10.1093/jnci/djv348 [PubMed]
- 115. Tice JA, Miike R, Adduci K, Petrakis NL, King E, Wrensch MR. Nipple aspirate fluid cytology and the Gail model for breast cancer risk assessment in a screening population. Cancer Epidemiol Biomarkers Prev. 2005; 14:324–28. https://doi.org/10.1158/1055-9965.EPI-04-0289 [PubMed]
- 116. Barlow WE, White E, Ballard-Barbash R, Vacek PM, Titus-Ernstoff L, Carney PA, Tice JA, Buist DS, Geller BM, Rosenberg R, Yankaskas BC, Kerlikowske K. Prospective breast cancer risk prediction model for women undergoing screening mammography. J Natl Cancer Inst. 2006; 98:1204–14. https://doi.org/10.1093/jnci/djj331 [PubMed]



