Introduction
Oocyte quality, an indicator of female reproductive health, is essential for embryo development and pregnancy outcome. Following the completion of nuclear and cytoplasmic maturation, fully-grown oocytes acquire the ability to fertilize and support embryo development [1]. Maternal age-related decline in fecundity is largely dependent on oocyte quality [2, 3]. Studies of women of advanced reproductive age revealed the mitochondrial dysfunction, spindle disorganization, and chromosome mis-segregation in oocytes [4]. These defects during meiosis sharply increase the incidence of infertility, miscarriage, and congenital malformation.
Sirtuins (SIRT1–7) have been implicated in influencing diverse biological events, such as aging, energy control, transcription, and apoptosis [5, 6]. For example, SIRT1 was initially discovered to deacetylate p53 tumor suppressor protein and function in chromatin regulation and genome stabilization [7]. SIRT2 was found to be associated with mitotic structures and to ensure normal cell division [8]. SIRT3 has been shown to regulate mitochondrial fatty acid oxidation by controlling the acetylation level of long-chain acyl-CoA dehydrogenase (LCAD) in mouse liver [9]. SIRT4 was identified as a lipoamidase that inhibits pyruvate dehydrogenase complex [10]. SIRT6 was initially characterized as a nuclear ADP-ribosyltransferase [11]. In mice, deficiency for SIRT6 results in genomic instability and metabolic defects, highlighting its importance in aging, energy supply and cancer [12]. Recently, we found that SIRT6 depletion leads to the aberrant spindles and chromosome congression failure in mouse oocytes [13]. It has also been reported that SIRT6 functions in the maintenance of telomeric chromatin structure and genomic integrity via the deacetylation of histone H3 lysine9 (H3K9) [14].
Telomeres are composed of repetitive G-rich sequences and associated proteins. It has been well recognized that telomeres serve multiple functions in protecting the ends of chromosomes from degradation and preventing illegitimate chromosomal recombination [15, 16]. Telomerase, also called terminal transferase, is a DNA polymerase that synthesizes telomeric DNA sequences [17]. In most somatic cell types, telomeres progressively shorten because of the absence of telomerase and the incomplete replication of the linear DNA molecules [18]. Maternal aging adversely impacts telomerase activity and telomere length in ovaries, likely through the long-term oxidative stress and activation of DNA damage checkpoint [19].
Although multiple pathways have been proposed to contribute to the decreased quality of aged oocytes, the underlying mechanisms are still lacking. Herein, our study identified a reduction of SIRT6 expression in aged oocytes. We further revealed that specific depletion of SIRT6 in mouse oocyte could induce the telomere dysfunction and compromises the developmental potential.
Results
Reduced SIRT6 expression in oocytes and embryos from aged mice
To dissect the potential relationship between SIRT6 and quality control of oocyte aging, we first examined the expression of SIRT6 in oocytes from old mice (42-45 weeks old). Analysis of quantitative real-time PCR showed that the relative mRNA levels of Sirt6 were significantly downregulated in aged oocytes in comparison to the oocytes from young mice (Figure 1A). In line with this, about a 60% of reduction in SIRT6 protein expression was detected in old oocytes (Figure 1B). Moreover, we found that the expression of SIRT6 protein was also dramatically lowered in 2-cell embryos isolated from old mice relative to controls (Figure 1C). These findings indicate that SIRT6 reduction might contribute to the compromised quality of aged mice.
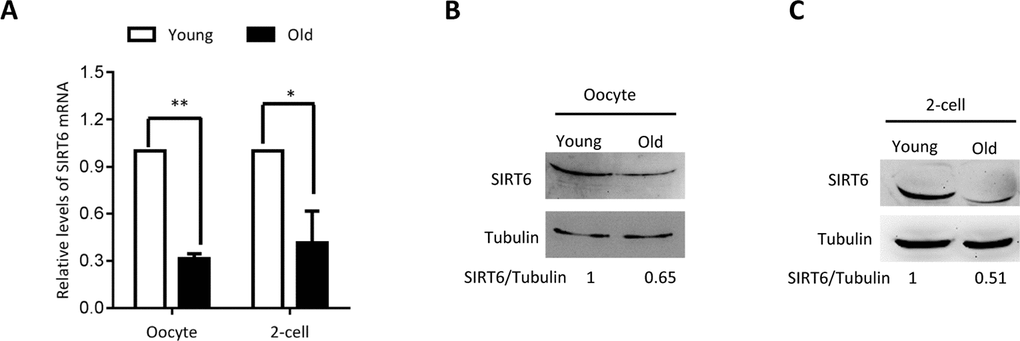
Figure 1. SIRT6 expression in aged mouse oocytes and embryos. Fully-grown oocytes and two-cell embryos from young and aged mouse were collected to evaluate SIRT6 expression. (A) Quantitative RT-PCR showing the lowered SIRT6 mRNA levels in aged oocytes and two-cell embryos. (B–C) SIRT6 protein expression in aged oocytes and two-cell embryos was verified by immunoblotting. Tubulin served as a loading control. Band intensity was calculated using ImageJ software, and the ratio of SIRT6/tubulin expression was normalized and values are indicated. Data are expressed as mean percentage ± SD of three independent experiments. *P <0.05, **P <0.01.
Telomere dysfunction and DNA lesion in aged oocytes and embryos
It has been widely reported that aging is associated with shorter telomere function in diverse cell types and tissues. Therefore, we decided to evaluate the status of telomere in oocytes/embryos from old mice. Telomeres protect the ends of the chromosome from fusion to other chromosomes during cell division. Chromosomal rearrangements due to telomere shortening in cancer cells are thought to result from the extensive chromosome fusion [20].
As shown in Figure 2A, we found that the relative telomere lengths were markedly decreased in both oocytes and 2-cell embryos from old mice. Telomeric repeat-binding factor 1 (TRF1) is a homodimeric protein, which interacts with the telomeric DNA at its C-terminus [21]. TRF1 blocks telomerase access to the telomeres, and overexpression of TRF1 induces a gradual decline in telomere length [22, 23]. Replication stress induces telomere dysfunction, which can activate the DNA damage response factors, forming the so-called telomere dysfunction-induced foci (TIFs) [24]. Next cells were double-stained for phosphorylated H2AX (γH2AX, a surrogate marker for DNA damage) and TRF1. As shown in Figure 2B–2D, the numbers of TIFs are significantly increased in old oocytes and embryos when compared to young cells (arrows). This indicates that the elevated DNA lesions and telomere dysfunction in aged oocytes and embryos. Accumulation of unrepaired DNA damage generates signals that affect genome integrity and cell cycle progression.
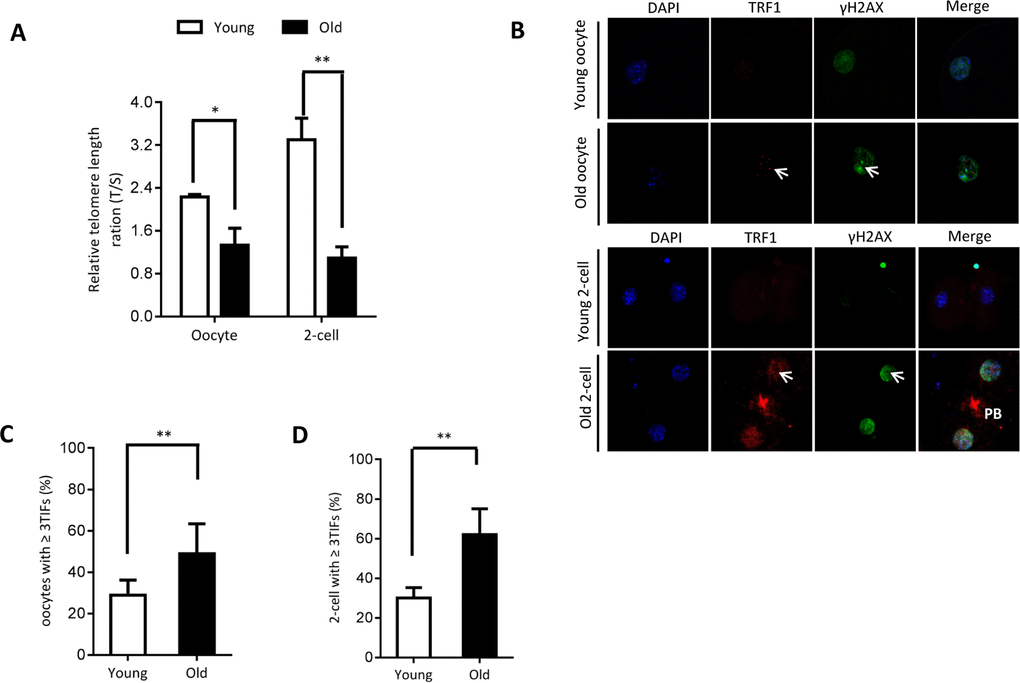
Figure 2. Telomere dysfunction and DNA damage in aged oocytes and embryos. (A) Relative telomere length in oocytes and two-cell embryos is expressed as a T/S ratio determined by quantitative RT-PCR analysis. Data are expressed as mean percentage ± SD of three independent experiments (oocytes: n=114 young, n=106 old; embryos: n=85 young, n=86 old). (B) Representative images of oocytes/two-cell embryos stained with antibodies against TRF1 (red) and γH2AX (green), and co-stained with Hoechst 33342 for chromosomes (blue). PB, polar body. Scale bars, 25 µm. (C–D) Quantification of DNA damage-induced foci (TIFs) from (B). TIFs were detected by co-localization of TRF1 and γ-H2AX, and cells with at least 3 TIFs were scored. Data represent averages of 3–10 fields. Error bars indicate SD (oocytes: n=24 young, n=26 old; embryos: n=35 young, n=38 old). Statistical analyses were performed with Student’s t-test. *P <0.05, **P <0.01.
Effects of SIRT6 depletion in oocyte on embryo development
The phenotypes in old oocytes/embryos mentioned above prompted us to ask whether they are associated with the loss of SIRT6 in oocytes. To address this question, normal MII oocytes were microinjected with specifically designed siRNAs to knock down the SIRT6 level (SIRT6-KD, Figure 3A). Immunoblotting confirmed that siRNA injection led to a dramatic reduction in SIRT6 protein expression in oocytes (Figure 3B). Next, we carried out in vitro fertilization (IVF) of control and SIRT6-KD oocytes, and then zygotes were cultured to check the subsequent embryonic development. As shown in Figure 3C–3D, early embryos derived from SIRT6-KD oocytes displayed a significant decrease in blastocyst formation (red arrows) and increase in cytoplasmic fragmentation in comparison to controls. These observations indicate the compromised developmental potential of early embryos when SIRT6 was abated in oocytes. SIRT6 loss is associated with the dysfunction of telomeres via affecting telomeric chromatin structure. In line with notion, telomeres shortened markedly in 2-cell embryos derived from SIRT6-KD oocytes compared to control embryos (Figure 4A). Moreover, we found that >3 TIFs were frequently detected in 2-cell embryos originated from SIRT6-KD oocytes (Figure 4B–4C). Telomeres are composed of repetitive G-rich sequence and specific binding proteins that together stabilize the ends of chromosomes. These structures are closely related to cellular apoptosis [25, 26]. To evaluate the apoptotic status, Terminal dUTP Nick End Labeling (TUNEL) assay was conducted on control and SIRT6-KD blastocysts. As shown in Figure 4D–4E, TUNEL positive nuclei were hardly detected in normal blastocyst. In contrast, the proportion of apoptotic blastomeres was significantly increased in embryos derived from SIRT6-KD oocytes. Altogether, these results indicate that oocyte SIRT6 is important for maintaining the telomere function and genome integrity in early embryos, thereupon determining the developmental competence.
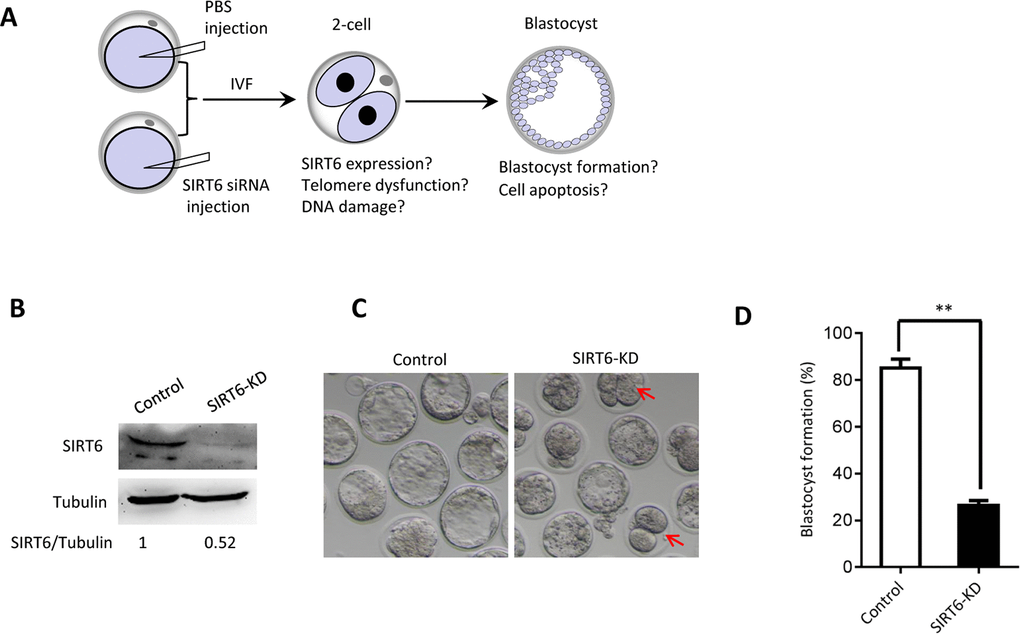
Figure 3. SIRT6 knockdown in oocytes results in the developmental delay of early embryos. (A) Schematic presentation of the SIRT6-knockdown experiments. (B) Efficiency of SIRT6 knockdown (SIRT6-KD) after siRNA injection was verified by immunoblotting. Tubulin served as a loading control. Band intensity was calculated using Image J software. (C) Representative contrast images of blastocyst embryos derived from control and SIRT6-KD oocytes. Red arrows indicate the developmental delay. (D) The percentage of zygotes that successfully progressed to the blastocyst stage during in vitro culture (n= 65 for control; n=55 for SIRT6-KD). Data are expressed as mean percentage ± SD of three independent experiments. Statistical analyses were performed with Student’s t-test. **P <0.01.
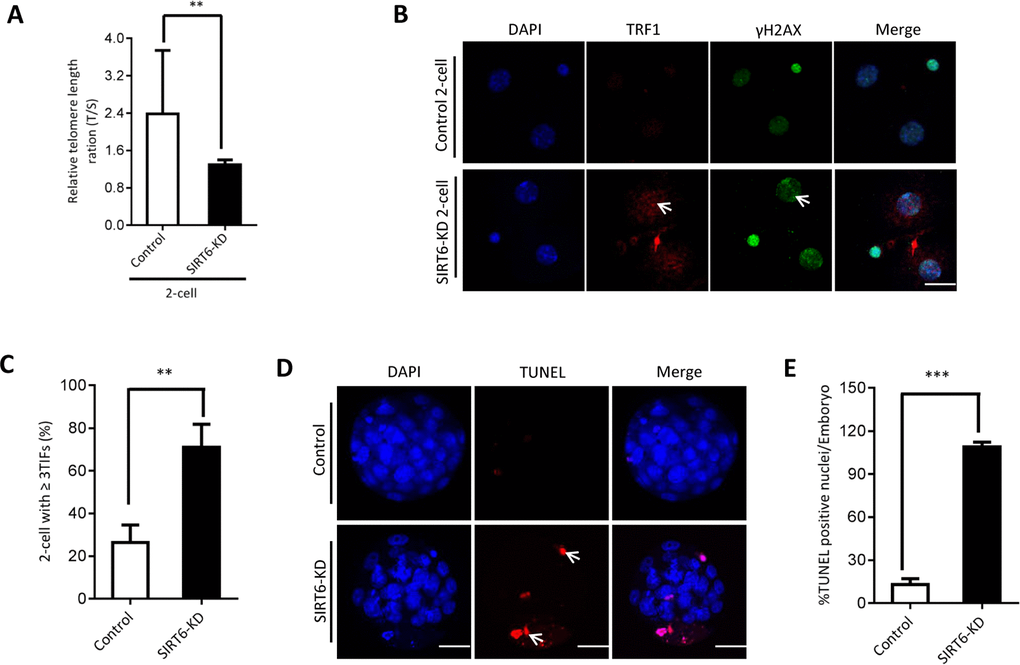
Figure 4. SIRT6 knockdown in oocytes induces DNA damage and apoptosis of early embryos. (A) Relative telomere length in two-cell embryos is expressed as a T/S ratio determined by quantitative RT-PCR analysis. Data are expressed as mean percentage ± SD of three independent experiments (n=125 for control; n=128 for SIRT6-KD). (B) Representative images of young and SIRT6-KD two-cell embryos stained with antibodies against TRF1 (red) and γH2AX (green), and co-stained with Hoechst 33342 for chromosomes (blue). Scale bars, 25 µm. (C) Quantification of DNA damage-induced foci (TIFs) from (B). TIFs were detected by co-localization of TRF1 and γ-H2AX, and cells with at least 3 TIFs were scored (n=35 for control; n=32 for SIRT6-KD). Data represent averages of 3–10 fields. (D) TUNEL analysis of control and SIRT6-KD embryos. Embryos were labeled with Hoechst 33342 (blue) for DNA and by TUNEL for fragmented DNA (red). Arrowheads point to the apoptotic cells in blastocysts. (E) Quantification of control and SIRT6-KD blastocysts with TUNEL positive nuclei (n=82 for control; n=75 for SIRT6-KD). Statistical analyses were performed with Student’s t-test. *P<0.05, ** P<0.01, ***P<0.001.
SIRT6 overexpression partially rescues defective phenotypes of embryos from aged mice
Our above findings suggest the potential connection between SIRT6 insufficiency in oocytes and telomere dysfunction in early embryos from old mice. Therefore, we next sought to determine whether elevating expression of SIRT6 in aged oocytes could alleviate the deficient telomere in early embryos. To do this, we conducted SIRT6 overexpression experiments (SIRT6- OE) followed by IVF (Figure 5A). Immunoblotting assay verified the ectopic expression of exogenous SIRT6 protein in oocytes (Figure 5B). The telomere status and apoptosis in the resulting two-cell embryos and blastocysts were then evaluated, respectively. As shown in Figure 5C, ectopic expression of SIRT6 in aged oocytes promoted the telomere elongation in the subsequent 2-cell embryos. In line with this, the frequency of TIFs occurred in old embryos overexpressing SIRT6 was remarkably lowered (Figure 6A–6B). Moreover, we noticed that forced overexpression of SIRT6 reduces the percentage of apoptotic blastomeres in old blastocyst (Figure 6C–6D). Collectively, these data indicates that loss of SIRT6 in aged oocytes is an important factor inducing the dysfunctional telomeres and developmental abnormalities in early embryos.
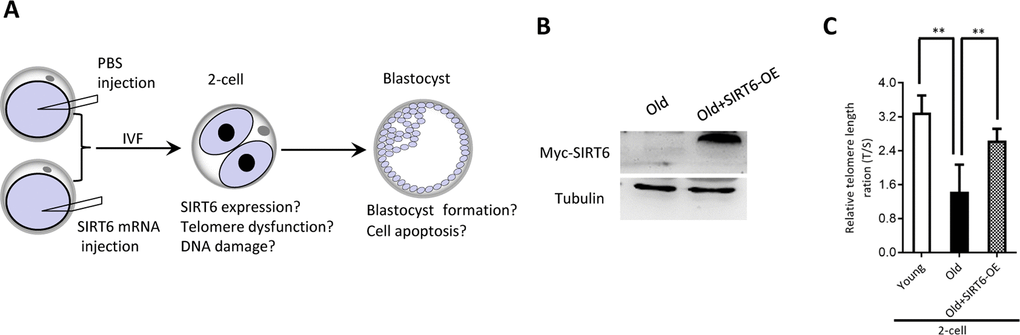
Figure 5. SIRT6 overexpression in aged oocytes increases the telomere length in early embryos. (A) Schematic presentation of the SIRT6 overexpression experiments. (B) Efficiency of SIRT6 overexpression (SIRT6-OE) after mRNA injection was verified by immunoblotting. (C) Relative telomere length expressed in two-cell embryos is expressed as a T/S ratio determined by quantitative RT-PCR analysis. Data are expressed as mean percentage ± SD of three independent experiments (n=86 for young; n=85 for old; n=80 for old+SIRT6-OE). **P <0.01.
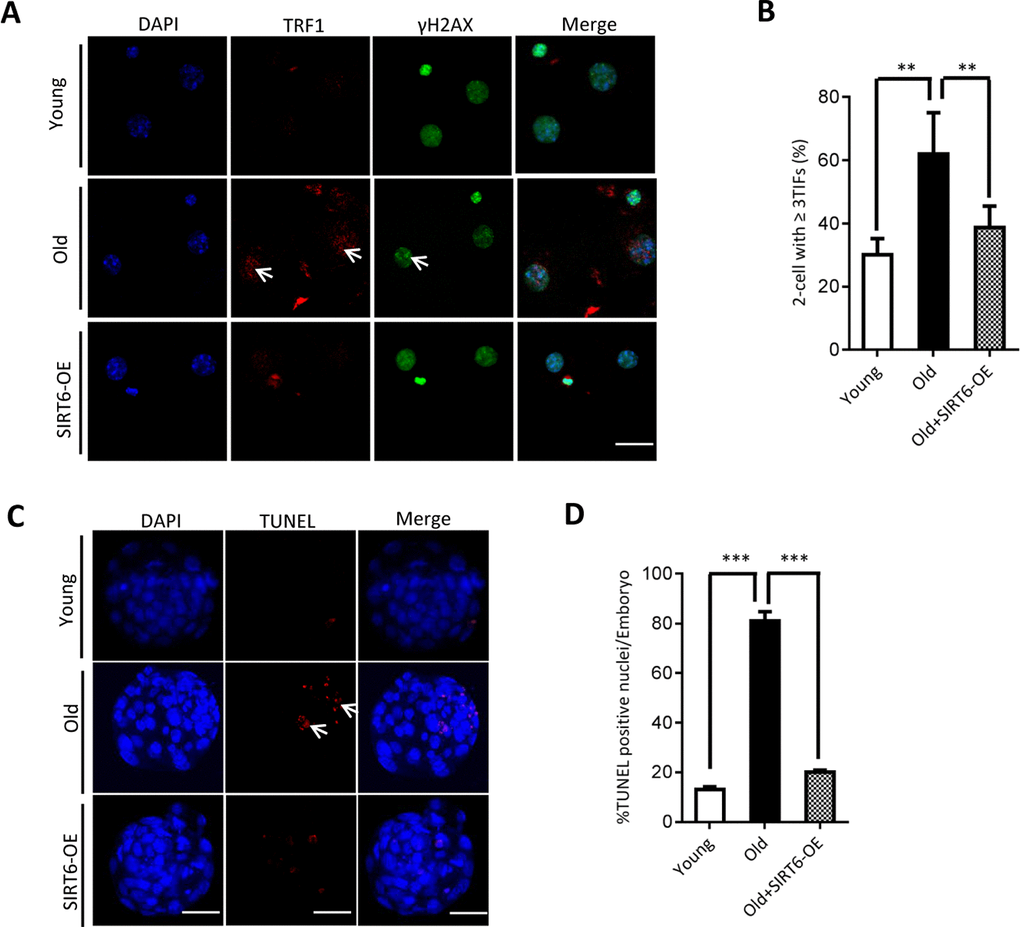
Figure 6. SIRT6 overexpression in aged oocytes partly prevented the defective phenotype of early embryos. (A) Representative images of young, old, and SIRT6-OE two-cell embryos stained with antibodies against TRF1 (red) and γH2AX (green), and co-stained with Hoechst 33342 for chromosomes (blue). Scale bars, 25 µm. (B) Quantification of DNA damage-induced foci (TIFs) from (A). TIFs were detected by co-localization of TRF1 and γ-H2AX, and cells with at least 3 TIFs were scored (n=26 for young; n=28 for old; n=25 for old+SIRT6-OE). (C) TUNEL analysis of young, old, and SIRT6-OE embryos. Embryos were labeled with Hoechst 33342 (blue) for DNA and by TUNEL for fragmented DNA (red). Arrowheads point to the apoptotic cells in blastocysts. Scale bars, 25 μm. (D) Quantification of young (n=90), old (n=82), and SIRT6-OE (n=78) blastocysts with TUNEL positive nuclei. Data are expressed as mean percentage ± SD from three independent experiments. ** P<0.01, ***P<0.001.
Discussion
Telomere is a region of repetitive nucleotide sequences at each end of a eukaryotic chromosome. Telomeres can not only prevent genetic information loss, but also protect the chromosomes from end-to-end fusions to keep genome stability [27, 28]. SIRT6 interacts with multiple chromatin-related proteins and possesses ADP-ribosylase and deacetylase activity [14, 29]. Depletion of SIRT6 has been reported to result in cellular senescence because of the telomere dysfunction [14]. Silencing of a telomere-proximal gene also requires SIRT6 [30]. Moreover, to activate the downstream DDR factors and DNA repair, SIRT6 is recruited to DNA double-strand breaks (DSBs) following DNA damage [31, 32]. In this study, we found that SIRT6 knockdown in oocytes causes the reduction in telomere length in early embryos. Telomere-binding proteins TRF1 and TRF2 bind specifically to the double-stranded TTAGGG repeats, and are important for the control of telomeric length and structure [33]. TRF1 has been reported to play a key role in both telomerase-dependent telomere maintenance and alternative lengthening [34, 35]. In support of this conception, our data showed that the frequency of TIFs was dramatically increased in 2-cell embryos when SIRT6 was knocked down in oocytes (Figure 4).
Consequently, the SIRT6-depleted embryos fragmentation, with the apoptotic blastomeres (Figure 3). Altogether, these findings strongly suggest that SIRT6 is an essential factor determining the telomere structure and genomic integrity during mouse early embryogenesis. It has been reported that SIRT6 can modulate the acetylation level of H3K56 and H3K9 in mitotic cells [14, 36]. Therefore, telomere dysfunction in SIRT6-KD oocytes is likely correlated with the altered telomere chromatin structure induced by elevated acetylation. Our ongoing research is to address this question.
Presence of telomerase activity in female germ cells ensures the proper meiotic process and the generation of euploid eggs [37]. It has been shown that DNA damage and telomere shortening resulted from the overproduction of free radicals promote ovarian aging [38]. Decreased Tert expression and telomerase activity have been detected in oocytes from reproductively aged females [39]. Similarly, here we notice the reduced telomere length and increased DNA lesions in oocytes and 2-cell embryos from aged mice (Figure 2). Telomere shortening compromises the chromosome synapsis and meiotic apparatus in germ cells, acting to produce aneuploid embryos after fertilization [40, 41]. Most aneuploidies in eggs arise in frequency with advancing maternal age [3, 42]. It is worthy of noting that SIRT6 expression was also remarkably decreased in these old oocytes (Figure 1). We have previously revealed that SIRT6 is critical for the proper assembly of meiotic structure during mouse oocyte meiosis [13]. Importantly, overexpression of SIRT6 in aged oocytes could partly prevent the telomere dysfunction and apoptosis in the subsequent early embryos (Figure 6). Consistent with this observation, telomere shortening was reported to be involved in the cytoplasmic fragmentation of preimplantation embryos [43].
In conclusion, our results suggest that SIRT6-controlled telomere structure is a crucial factor controlling oocyte quality from aged mice. Meanwhile, we propose that maternal aging results in the SIRT6 insufficiency in oocytes, which in turn disrupts the telomere function and genomic integrity, and consequently, contributing to the compromised developmental potential of the resultant early embryos.
Materials and Methods
All chemicals and culture media were purchased from Sigma (St. Louis, MO, USA) unless otherwise stated.
Mice
All animal experiments followed the rules of the Animal Care and Use Committee of Nanjing Medical University. ICR young female mice (3-4 week old) were used in our experiments. 42-45 weeks old female mice (near the end of their reproductive lifespan) were used as a natural aging mouse model.
Antibodies
The following antibodies were purchased: rabbit polyclonal anti-SIRT6 antibody (Sigma, Cat#: S4197), rabbit polyclonal anti-TRF1 antibody (Abcam, Cat#: ab1423), mouse monoclonal anti-γH2AX antibody (Abcam, Cat#: ab22551), mouse monoclonal anti-Myc antibody (Abcam, Cat#: ab18185).
Oocyte collection
To obtain MII oocytes, female mice were superovulated by injecting 5U of pregnant mare’s serum gonadotropin (PMSG) followed by 5U of human chorionic gonadotropin (hCG) 48h after PMSG priming. Mice were euthanized by cervical dislocation 14h after the hCG injection. Cumulus–oocyte complexes (COCs) were isolated from oviduct ampullae, and denuded MII oocytes were then obtained by removing the cumulus mass in medium containing 0.5mg/ml hyaluronidase at 37°C
In vitro fertilization, embryo collection and culture
IVF assays were performed following our published protocols [44]. Sperm were collected from the dissected epididymis of ICR mice aged 10–20 weeks and left to capacitate for 1 hour in HTF fertilization medium (Millipore, Merck) supplemented with 10 mg/ml BSA. Then, dispersed spermatozoa were added to HTF drops containing COCs for fertilization. Zygotes were cultured into KSOM medium (Millipore, Merck) at 37 °C in a humidified atmosphere of 5% CO2, 5% O2 and 90% N2.
Plasmid construction and mRNA synthesis
Total RNA was extracted from mouse oocytes/embryos using Arcturus Pico Pure RNA Isolation Kit (Applied Biosystems, CA, USA), and the cDNA was produced with QIA quick PCR Purification Kit (Qiagen, Germany). PCR products were cloned into the pCS2+ vector with Myc tags. mRNA synthesis was conducted by in vitro transcription with SP6 mMESSAGE mMACHINE (Ambion, CA) as we reported previously [45]. The related primer sequences can be found in Supplementary Table 1.
Knockdown and overexpression analysis
Microinjection was performed employing an inverted microscope (Eclipse Ti-S, Nikon) equipped with a micromanipulator (Narishige) as we reported previously [46]. In knockdown experiments, SIRT6-specific siRNA was diluted to give a stock concentration of 1mM, and 5pl solution was injected into oocytes. In overexpression experiment, 10pl SIRT6 cRNA solution (10 ng/µl) was injected into the cytoplasm of MII-arrested oocytes. Oocytes were cultured for 3 hours before the subsequent manipulation. The siRNA sequences can be found in Supplementary Table 1.
Western blotting
A total of 100 oocytes were lysed in Laemmli sample buffer with protease inhibitor, incubated at 95°C for 5 min, then stored at -20°C. Protein extracts were separated using SDS–PAGE and transferred to PVDF membranes. After blocking, the membranes were incubated at 4 °C overnight with anti-SIRT6 antibody (1:1,000), or anti-Myc antibody (1:1,000). After three washes, protein samples were incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (1:5,000; Thermo Fisher Scientific) at room temperature for 1 hour. The bands were visualized using an ECL Plus Western Blotting Detection System (GE Healthcare, Little Chalfont, UK).
Quantitative real-time PCR
Quantitative RT–PCR was performed as we described previously [47]. Total cellular RNA was isolated from 50 oocytes/embryos using an Arcturus PicoPure RNA Isolation kit (Applied Biosystems) RNA queous-Micro Kit (Ambion, TX). GAPDH was used as an internal control. Average telomere length was measured from total genomic DNA using a RT-PCR assay according to the published protocol [48, 49]. Primer sequences are listed in Supplementary Table 1.
Immunofluorescence
Oocytes/embryos were permeabilized with 0.5% Triton for 4-5 min, washed in PBS containing 0.1% polyvinyl pyrrolidone (PBS–PVP) for three times. Following a fixation step with 3.7% paraformaldehyde, oocytes or two-cell embryos were washed in PBS–PVP for 3 times and treated with blocking buffer (1% BSA-supplemented PBS) for 1 hour. Samples were incubated overnight at 4°C with rabbit polyclonal anti-TRF1 antibody (1:300) and mouse monoclonal anti-γH2AX antibody (1:200). After multiple washes, cells were incubated with Alexa Fluor 555–conjugated goat anti-rabbit and Alexa Fluor 488–conjugated goat anti-mouse secondary antibodies (1:150; Molecular Probes) for 1 hour at room temperature. Chromosomes were counterstained with Hoechst 33342 (1:250) for 10 min. Samples were mounted on slides with a drop of antifade medium (Vectashield, Vector Laboratories, CA, USA) and then examined under a laser scanning confocal microscope (LSM 710; Carl Zeiss, Oberkochen, Germany). For each antibody used, immunofluorescence was performed on oocytes/embryos from young and old mice in parallel and identical conditions. Images were always acquired using the same confocal microscope settings. The amount of γH2AX foci was analyzed using ImageJ (NIH, USA). To evaluate apoptosis, fixed embryos were processed for TUNEL analysis (In Situ Cell Death Detection Kit, Fluorescein, Roche).
Statistical analysis
All experiments were repeated at least three times. Data are presented as mean ± SD, unless otherwise indicated. Statistical comparisons were made with Student’s test and ANOVA when appropriate using Prism 5 software (GraphPad, San Diego, CA, USA). P<0.05 was considered to be significant.
Supplementary Materials
Author Contributions
JG and QW designed research; CL, CL, ZH, JZ and LH performed research; JG and QW analyzed data; JG and QW wrote paper.
Conflicts of Interest
The authors have nothing to disclose.
Funding
This work was supported by National Natural Science Foundation (NO. 31601202 to JG and NO. 31571543 to QW), National Key R&D Program (NO. 2018YFC1004000) of China, and the Science Foundation for Distinguished Young Scholars of Jiangsu Province (BK20180035 to QW).
References
- 1. Eppig JJ. Coordination of nuclear and cytoplasmic oocyte maturation in eutherian mammals. Reprod Fertil Dev. 1996; 8:485–89. https://doi.org/10.1071/RD9960485 [PubMed]
- 2. MacLennan M, Crichton JH, Playfoot CJ, Adams IR. Oocyte development, meiosis and aneuploidy. Semin Cell Dev Biol. 2015; 45:68–76. https://doi.org/10.1016/j.semcdb.2015.10.005 [PubMed]
- 3. Nagaoka SI, Hassold TJ, Hunt PA. Human aneuploidy: mechanisms and new insights into an age-old problem. Nat Rev Genet. 2012; 13:493–504. https://doi.org/10.1038/nrg3245 [PubMed]
- 4. Eichenlaub-Ritter U, Vogt E, Yin H, Gosden R. Spindles, mitochondria and redox potential in ageing oocytes. Reprod Biomed Online. 2004; 8:45–58. https://doi.org/10.1016/S1472-6483(10)60497-X [PubMed]
- 5. Baur JA, Ungvari Z, Minor RK, Le Couteur DG, de Cabo R. Are sirtuins viable targets for improving healthspan and lifespan? Nat Rev Drug Discov. 2012; 11:443–61. https://doi.org/10.1038/nrd3738 [PubMed]
- 6. Preyat N, Rossi M, Kers J, Chen L, Bertin J, Gough PJ, Le Moine A, Rongvaux A, Van Gool F, Leo O. Intracellular nicotinamide adenine dinucleotide promotes TNF-induced necroptosis in a sirtuin-dependent manner. Cell Death Differ. 2016; 23:29–40. https://doi.org/10.1038/cdd.2015.60 [PubMed]
- 7. Hisahara S, Chiba S, Matsumoto H, Tanno M, Yagi H, Shimohama S, Sato M, Horio Y. Histone deacetylase SIRT1 modulates neuronal differentiation by its nuclear translocation. Proc Natl Acad Sci USA. 2008; 105:15599–604. https://doi.org/10.1073/pnas.0800612105 [PubMed]
- 8. North BJ, Verdin E. Interphase nucleo-cytoplasmic shuttling and localization of SIRT2 during mitosis. PLoS One. 2007; 2:e784. https://doi.org/10.1371/journal.pone.0000784 [PubMed]
- 9. Hirschey MD, Shimazu T, Goetzman E, Jing E, Schwer B, Lombard DB, Grueter CA, Harris C, Biddinger S, Ilkayeva OR, Stevens RD, Li Y, Saha AK, et al. SIRT3 regulates mitochondrial fatty-acid oxidation by reversible enzyme deacetylation. Nature. 2010; 464:121–25. https://doi.org/10.1038/nature08778 [PubMed]
- 10. Mathias RA, Greco TM, Oberstein A, Budayeva HG, Chakrabarti R, Rowland EA, Kang Y, Shenk T, Cristea IM. Sirtuin 4 is a lipoamidase regulating pyruvate dehydrogenase complex activity. Cell. 2014; 159:1615–25. https://doi.org/10.1016/j.cell.2014.11.046 [PubMed]
- 11. Liszt G, Ford E, Kurtev M, Guarente L. Mouse Sir2 homolog SIRT6 is a nuclear ADP-ribosyltransferase. J Biol Chem. 2005; 280:21313–20. https://doi.org/10.1074/jbc.M413296200 [PubMed]
- 12. Mostoslavsky R, Chua KF, Lombard DB, Pang WW, Fischer MR, Gellon L, Liu P, Mostoslavsky G, Franco S, Murphy MM, Mills KD, Patel P, Hsu JT, et al. Genomic instability and aging-like phenotype in the absence of mammalian SIRT6. Cell. 2006; 124:315–29. https://doi.org/10.1016/j.cell.2005.11.044 [PubMed]
- 13. Han L, Ge J, Zhang L, Ma R, Hou X, Li B, Moley K, Wang Q. Sirt6 depletion causes spindle defects and chromosome misalignment during meiosis of mouse oocyte. Sci Rep. 2015; 5:15366. https://doi.org/10.1038/srep15366 [PubMed]
- 14. Michishita E, McCord RA, Berber E, Kioi M, Padilla-Nash H, Damian M, Cheung P, Kusumoto R, Kawahara TL, Barrett JC, Chang HY, Bohr VA, Ried T, et al. SIRT6 is a histone H3 lysine 9 deacetylase that modulates telomeric chromatin. Nature. 2008; 452:492–96. https://doi.org/10.1038/nature06736 [PubMed]
- 15. Hande MP, Samper E, Lansdorp P, Blasco MA. Telomere length dynamics and chromosomal instability in cells derived from telomerase null mice. J Cell Biol. 1999; 144:589–601. https://doi.org/10.1083/jcb.144.4.589 [PubMed]
- 16. Zhao Y, Hoshiyama H, Shay JW, Wright WE. Quantitative telomeric overhang determination using a double-strand specific nuclease. Nucleic Acids Res. 2008; 36:e14. https://doi.org/10.1093/nar/gkm1063 [PubMed]
- 17. Cawthon RM. Telomere measurement by quantitative PCR. Nucleic Acids Res. 2002; 30:e47. https://doi.org/10.1093/nar/30.10.e47 [PubMed]
- 18. Allsopp RC, Vaziri H, Patterson C, Goldstein S, Younglai EV, Futcher AB, Greider CW, Harley CB. Telomere length predicts replicative capacity of human fibroblasts. Proc Natl Acad Sci USA. 1992; 89:10114–18. https://doi.org/10.1073/pnas.89.21.10114 [PubMed]
- 19. Hanna CW, Bretherick KL, Gair JL, Fluker MR, Stephenson MD, Robinson WP. Telomere length and reproductive aging. Hum Reprod. 2009; 24:1206–11. https://doi.org/10.1093/humrep/dep007 [PubMed]
- 20. Bailey SM, Murnane JP. Telomeres, chromosome instability and cancer. Nucleic Acids Res. 2006; 34:2408–17. https://doi.org/10.1093/nar/gkl303 [PubMed]
- 21. Kim MK, Kang MR, Nam HW, Bae YS, Kim YS, Chung IK. Regulation of telomeric repeat binding factor 1 binding to telomeres by casein kinase 2-mediated phosphorylation. J Biol Chem. 2008; 283:14144–52. https://doi.org/10.1074/jbc.M710065200 [PubMed]
- 22. Smogorzewska A, van Steensel B, Bianchi A, Oelmann S, Schaefer MR, Schnapp G, de Lange T. Control of human telomere length by TRF1 and TRF2. Mol Cell Biol. 2000; 20:1659–68. https://doi.org/10.1128/MCB.20.5.1659-1668.2000 [PubMed]
- 23. van Steensel B, de Lange T. Control of telomere length by the human telomeric protein TRF1. Nature. 1997; 385:740–43. https://doi.org/10.1038/385740a0 [PubMed]
- 24. Takai H, Smogorzewska A, de Lange T. DNA damage foci at dysfunctional telomeres. Curr Biol. 2003; 13:1549–56. https://doi.org/10.1016/S0960-9822(03)00542-6 [PubMed]
- 25. Counter CM, Avilion AA, LeFeuvre CE, Stewart NG, Greider CW, Harley CB, Bacchetti S. Telomere shortening associated with chromosome instability is arrested in immortal cells which express telomerase activity. EMBO J. 1992; 11:1921–29. https://doi.org/10.1002/j.1460-2075.1992.tb05245.x [PubMed]
- 26. Webb CJ, Wu Y, Zakian VA. DNA repair at telomeres: keeping the ends intact. Cold Spring Harb Perspect Biol. 2013; 5:a012666. https://doi.org/10.1101/cshperspect.a012666 [PubMed]
- 27. Shampay J, Szostak JW, Blackburn EH. DNA sequences of telomeres maintained in yeast. Nature. 1984; 310:154–57. https://doi.org/10.1038/310154a0 [PubMed]
- 28. Blackburn EH. Telomere states and cell fates. Nature. 2000; 408:53–56. https://doi.org/10.1038/35040500 [PubMed]
- 29. Yang B, Zwaans BM, Eckersdorff M, Lombard DB. The sirtuin SIRT6 deacetylates H3 K56Ac in vivo to promote genomic stability. Cell Cycle. 2009; 8:2662–63. https://doi.org/10.4161/cc.8.16.9329 [PubMed]
- 30. Tennen RI, Bua DJ, Wright WE, Chua KF. SIRT6 is required for maintenance of telomere position effect in human cells. Nat Commun. 2011; 2:433. https://doi.org/10.1038/ncomms1443 [PubMed]
- 31. McCord RA, Michishita E, Hong T, Berber E, Boxer LD, Kusumoto R, Guan S, Shi X, Gozani O, Burlingame AL, Bohr VA, Chua KF. SIRT6 stabilizes DNA-dependent protein kinase at chromatin for DNA double-strand break repair. Aging (Albany NY). 2009; 1:109–21. https://doi.org/10.18632/aging.100011 [PubMed]
- 32. Jia G, Su L, Singhal S, Liu X. Emerging roles of SIRT6 on telomere maintenance, DNA repair, metabolism and mammalian aging. Mol Cell Biochem. 2012; 364:345–50. https://doi.org/10.1007/s11010-012-1236-8 [PubMed]
- 33. Li Y, Meng X, Wang W, Liu F, Hao Z, Yang Y, Zhao J, Yin W, Xu L, Zhao R, Hu J. Cardioprotective Effects of SIRT6 in a Mouse Model of Transverse Aortic Constriction-Induced Heart Failure. Front Physiol. 2017; 8:394. https://doi.org/10.3389/fphys.2017.00394 [PubMed]
- 34. Chong L, van Steensel B, Broccoli D, Erdjument-Bromage H, Hanish J, Tempst P, de Lange T. A human telomeric protein. Science. 1995; 270:1663–67. https://doi.org/10.1126/science.270.5242.1663 [PubMed]
- 35. Kimura M, Cherkas LF, Kato BS, Demissie S, Hjelmborg JB, Brimacombe M, Cupples A, Hunkin JL, Gardner JP, Lu X, Cao X, Sastrasinh M, Province MA, et al. Offspring’s leukocyte telomere length, paternal age, and telomere elongation in sperm. PLoS Genet. 2008; 4:e37. https://doi.org/10.1371/journal.pgen.0040037 [PubMed]
- 36. Kouzarides T. Chromatin modifications and their function. Cell. 2007; 128:693–705. https://doi.org/10.1016/j.cell.2007.02.005 [PubMed]
- 37. Rocca MS, Foresta C, Ferlin A. Telomere length: lights and shadows on their role in human reproduction. Biol Reprod. 2019; 100:305–317. https://doi.org/10.1093/biolre/ioy208 [PubMed]
- 38. Liu M, Yin Y, Ye X, Zeng M, Zhao Q, Keefe DL, Liu L. Resveratrol protects against age-associated infertility in mice. Hum Reprod. 2013; 28:707–17. https://doi.org/10.1093/humrep/des437 [PubMed]
- 39. Yamada-Fukunaga T, Yamada M, Hamatani T, Chikazawa N, Ogawa S, Akutsu H, Miura T, Miyado K, Tarín JJ, Kuji N, Umezawa A, Yoshimura Y. Age-associated telomere shortening in mouse oocytes. Reprod Biol Endocrinol. 2013; 11:108. https://doi.org/10.1186/1477-7827-11-108 [PubMed]
- 40. Mathieu N, Pirzio L, Freulet-Marrière MA, Desmaze C, Sabatier L. Telomeres and chromosomal instability. Cell Mol Life Sci. 2004; 61:641–56. https://doi.org/10.1007/s00018-003-3296-0 [PubMed]
- 41. Ozturk S, Sozen B, Demir N. Telomere length and telomerase activity during oocyte maturation and early embryo development in mammalian species. Mol Hum Reprod. 2014; 20:15–30. https://doi.org/10.1093/molehr/gat055 [PubMed]
- 42. Qiao J, Wang ZB, Feng HL, Miao YL, Wang Q, Xu Y, Wei YC, Yan J, Wang WH, Shen W, Sun SC, Schatten H, Sun QY. The root of reduced fertility in aged women and possible therapentic options: current status and future perspects. Mol Aspects Med. 2014; 38:54–85. https://doi.org/10.1016/j.mam.2013.06.001 [PubMed]
- 43. Keefe DL, Franco S, Liu L, Trimarchi J, Cao B, Weitzen S, Agarwal S, Blasco MA. Telomere length predicts embryo fragmentation after in vitro fertilization in women—toward a telomere theory of reproductive aging in women. Am J Obstet Gynecol. 2005; 192:1256–60. https://doi.org/10.1016/j.ajog.2005.01.036 [PubMed]
- 44. Han L, Ren C, Li L, Li X, Ge J, Wang H, Miao YL, Guo X, Moley KH, Shu W, Wang Q. Publisher Correction: embryonic defects induced by maternal obesity in mice derive from Stella insufficiency in oocytes. Nat Genet. 2018; 50:768. https://doi.org/10.1038/s41588-018-0077-0 [PubMed]
- 45. Qiu D, Hou X, Han L, Li X, Ge J, Wang Q. Sirt2-BubR1 acetylation pathway mediates the effects of advanced maternal age on oocyte quality. Aging Cell. 2018; 17:e12698. https://doi.org/10.1111/acel.12698 [PubMed]
- 46. Zeng J, Jiang M, Wu X, Diao F, Qiu D, Hou X, Wang H, Li L, Li C, Ge J, Liu J, Ou X, Wang Q. SIRT4 is essential for metabolic control and meiotic structure during mouse oocyte maturation. Aging Cell. 2018; 17:e12789. https://doi.org/10.1111/acel.12789 [PubMed]
- 47. Hou X, Zhang L, Han L, Ge J, Ma R, Zhang X, Moley K, Schedl T, Wang Q. Differing roles of pyruvate dehydrogenase kinases during mouse oocyte maturation. J Cell Sci. 2015; 128:2319–29. https://doi.org/10.1242/jcs.167049 [PubMed]
- 48. Callicott RJ, Womack JE. Real-time PCR assay for measurement of mouse telomeres. Comp Med. 2006; 56:17–22. [PubMed]
- 49. Liu L, Bailey SM, Okuka M, Muñoz P, Li C, Zhou L, Wu C, Czerwiec E, Sandler L, Seyfang A, Blasco MA, Keefe DL. Telomere lengthening early in development. Nat Cell Biol. 2007; 9:1436–41. [PubMed]



