Introduction
As the main complication of diabetes-related kidney damage, diabetic nephropathy (DN) is a complex clinical problem worldwide [1,2]. The high frequency of DN in diabetics has been observed despite considerable advancements in therapeutic approaches to diabetes [3,4]. An increasing number of research works have been performed to investigate the mechanisms of DN progression. However, finding effective molecular targets for DN clinical treatment is challenging.
Long non-coding RNAs (lncRNAs) are a vital group of non-coding RNA molecules that are >200 nucleotides in length [5,6]. The novel biologic roles of lncRNAs in DN have been confirmed due to the rapid development of RNA genomics. LncRNA ENSMUST00000147869 inhibits fibrosis and proliferation in mesangial cells (MCs) induced by DN [7]. MIAT/Nrf2 is a vital pathway for renal tubular epithelial injury induced by high glucose [8]. MALAT1 is upregulated in DN and is involved in podocyte injury induced by high glucose via β-catenin [9]. Long non-coding RNA maternally expressed gene 3 (lncRNA MEG3, ID: 55384) is a novel lncRNA that acts as a tumor suppressor in multiple cancer cells. For example, MEG3 suppresses invasion and proliferation by promoting the ubiquitination of EZH2 in gallbladder cancer [10]. MEG3 also inhibits growth by suppressing β-catenin in liver cancer cells via the PKM2/PTEN pathway [11]. However, the potential mechanism and biological function of MEG3 in DN are still barely elucidated.
MicroRNAs (miRNAs) are small noncoding RNAs that are 20–25 nucleotides in length [12,13]. MiRNAs regulate the expression of downstream targets via post-transcriptional modulation [14]. LncRNAs can function as competing endogenous RNAs (ceRNAs), which compete for miRNA binding and block miRNA-mediated target gene silencing [15]. The regulating network of ceRNAs is suggested to play important roles in DN development. LncRNA LINC01619 regulates podocyte injury via the miR-27a/FOXO1 pathway in DN [16]. NEAT1 enhances EMT and extracellular matrix accumulation by targeting the miR-27b-3p/ZEB1 pathway in DN [17]. LncRNA-NR_033515 enhances EMT, proliferation, and fibrogenesis by targeting miR-743b-5p in DN [18].
In the current research, we found that MEG3 was upregulated in DN in vivo and in vitro. MEG3 could enhance cell fibrosis and inflammatory response in MCs and could interact with miR-181a. Our study further demonstrated that MEG3 promotes fibrosis and inflammatory response in DN through the miR-181a/Egr-1/TLR4 pathway.
Results
MEG3 expression was upregulated in DN in vivo and in vitro
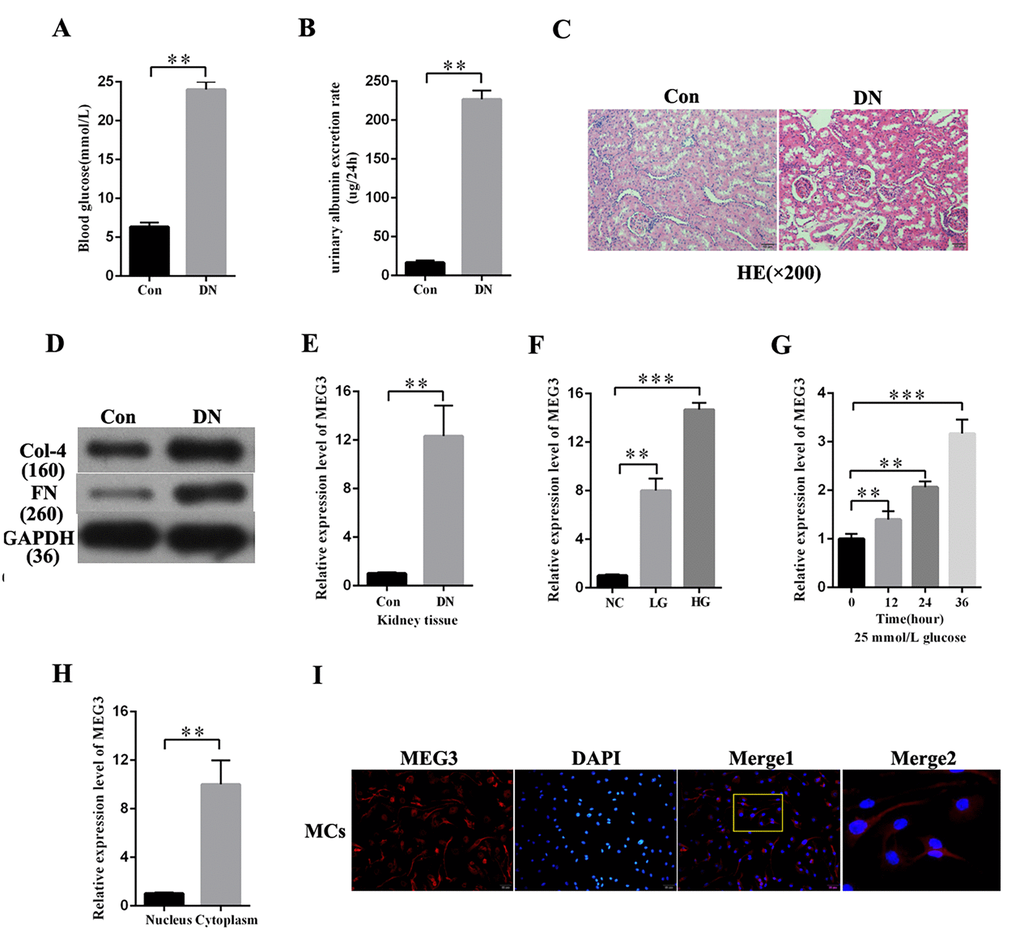
DN rat models were established to investigate the expression profile of MEG3 in DN. As respectively shown in Figures 1A and B, blood glucose level and urinary albumin excretion rate were significantly increased in DN rat models compared with those in the negative group. HE staining showed morphological glomerular changes in diabetic kidney tissue (Figure 1C). The expression level of fibrosis markers (Col-4 and FN) was increased in DN compared with that in the negative group (Figure 1D). MEG3 expression in DN was detected. QRT-PCR results indicated that MEG3 was upregulated in DN tissues (Figure 1E). MEG3 expression was measured in MCs stimulated by D-glucose at 5.5 (NC), 19.5 (LG), and 25 mmol/L (HG) and was found to increase with glucose concentration (Figure 1F). MEG3 expression in MCs cultured in D-glucose at 25 mmol/L was measured at 0, 12, 24, and 36 h. The results showed that MEG3 expression was upregulated with time (Figure 1G). The lncRNAs must be mainly expressed in the cytoplasm to act as an effective miRNA sponge. The qPCR and fluorescence in situ hybridization (FISH) results revealed that MEG3 was mainly localized in the cytoplasm of MCs (Figures 1H–1F). Thus, these results showed that MEG3 expression was upregulated in DN and mainly localized in the cytoplasm of MCs.
MEG3 enhanced the expression of inflammatory cytokines in vitro and in vivo
Increased inflammation is a vital mechanism in the progression of DN. MEG3 overexpression or knockdown was performed to study the effects of MEG3 on the inflammatory response in DN.
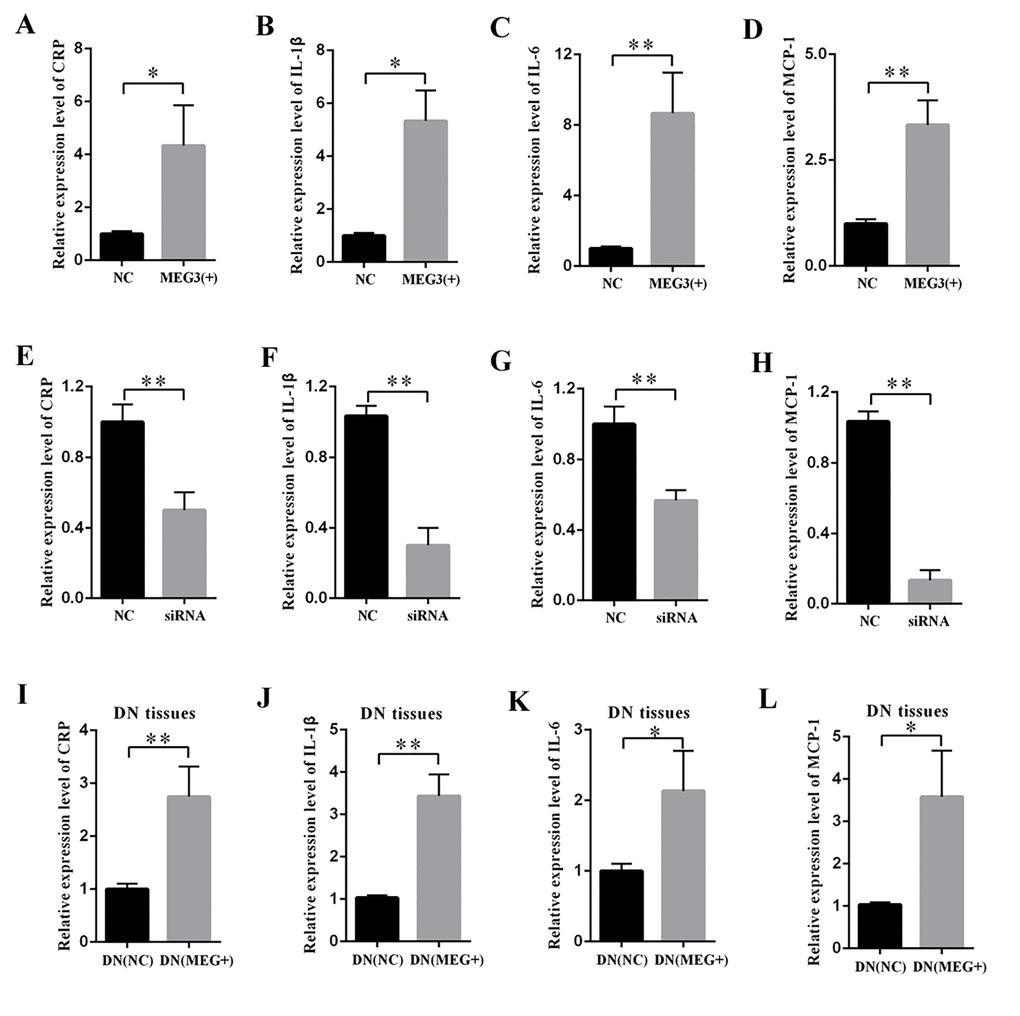
The supernatant of MCs was collected, and DN-related inflammatory cytokines were measured by ELISA. The results revealed that MEG3 overexpression raised the concentrations of CRP (Figure 3A), IL-1β (Figure 3B), IL-6 (Figure 3C), and MCP-1 (Figure 3D). By contrast, MEG3 silencing decreased the concentrations of CRP (Figure 3E), IL-1β (Figure 3F), IL-6 (Figure 3G), and MCP-1 (Figure 3H). DN rats were injected with LV-MEG3 to detect the effect of MEG3 in vivo. ELISA results indicated that MEG3 overexpression increased CRP (Figure 3I), IL-1β (Figure 3J), IL-6 (Figure 3K), and MCP-1 (Figure 3L) expression in the kidney cortex of DN rats. Therefore, these data indicate that MEG3 overexpression promoted inflammation in DN.
MEG3 interacted with miR-181a via a direct targeting and Ago2-dependent manner
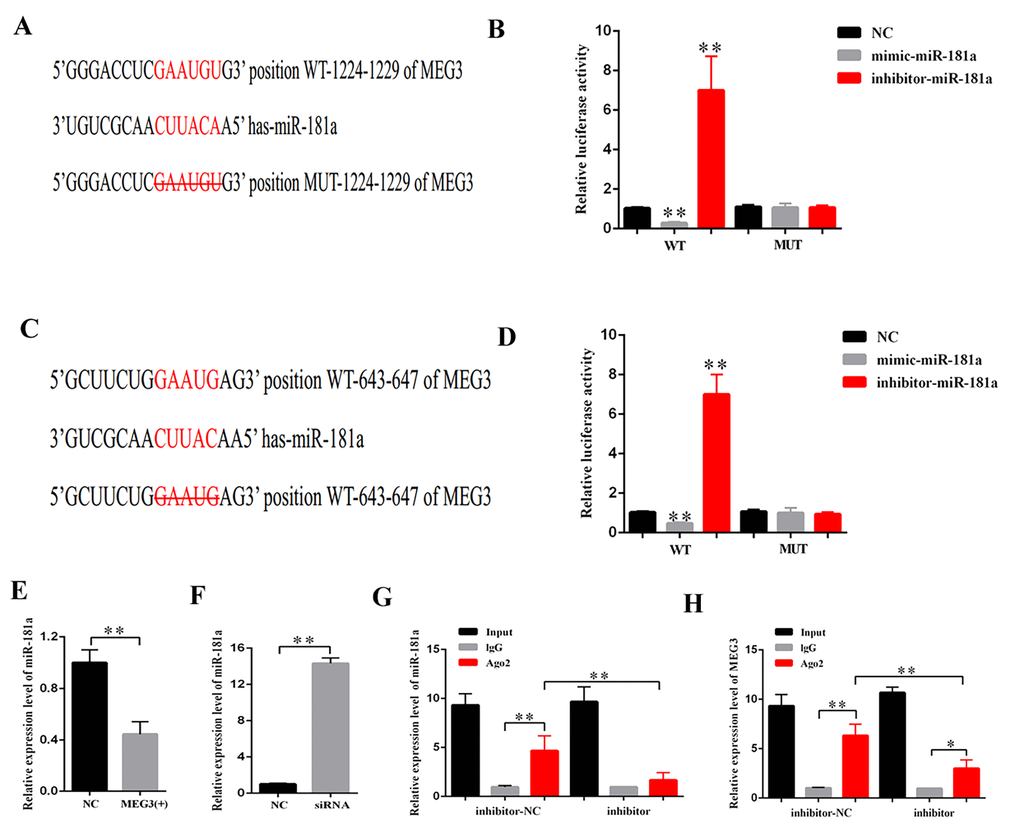
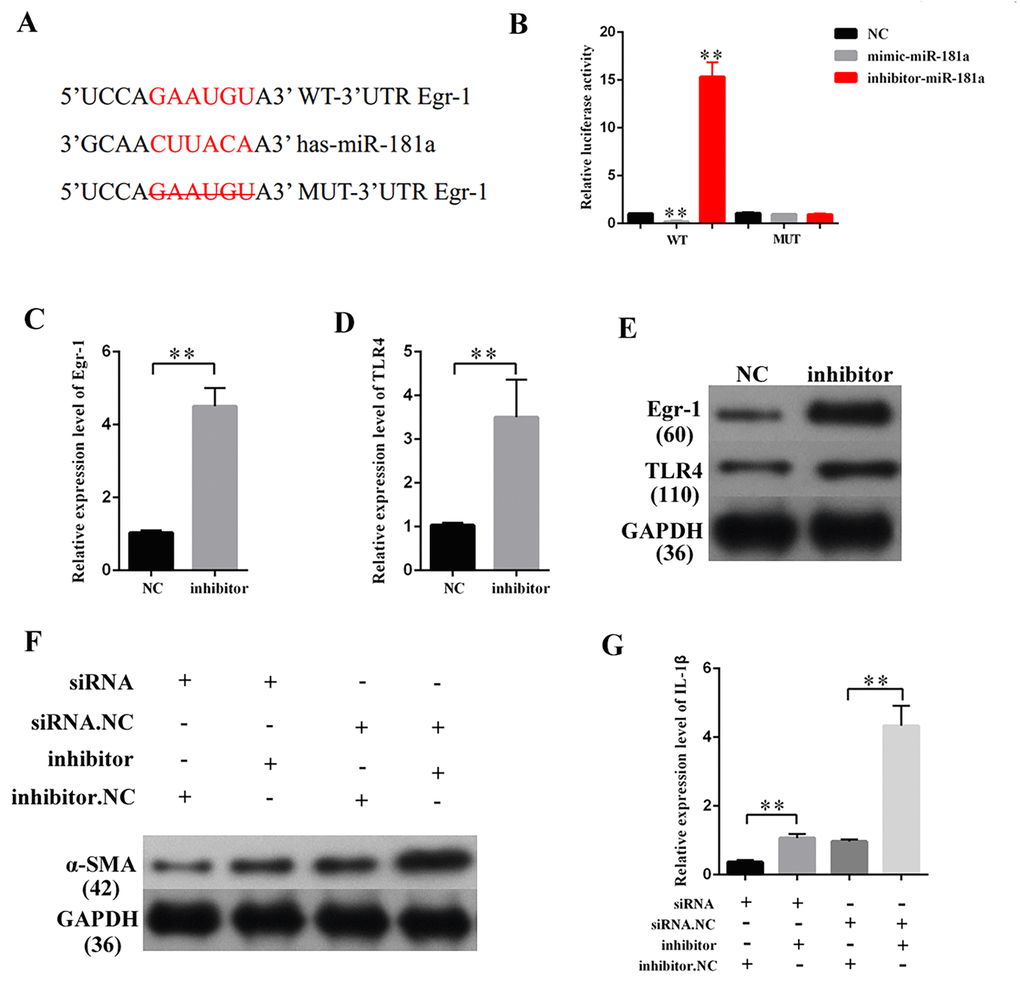
CeRNA functions as a vital regulatory mechanism in DN [19,20]. StarBase v2.0 (http://starbase.sysu.edu.cn/index.php) was used to predict the potential target miRNA of MEG3. Some of the results are presented in Supplementary Table 1. MiR-181a plays a vital role in fibrosis and inflammation [21,22]. Thus, miR-181a, which contains two binding sites in MEG3 transcript, was selected as the prediction candidate. Luciferase reporter assay was performed to confirm the direct binding relationship between MEG3 and miR-181a. This binding relationship followed the basic complementary pairing principle. Thus, an MEG3 mutant-type reporter, in which binding sites between MEG3 and miR-181a were knocked out, was built. The results showed that the luciferase activity of MEG3 wild- and mutant-type reporter genes were respectively suppressed and not inhibited by mimic-miR-181a. The luciferase activity of MEG3 wild- and mutant-type reporter genes were respectively increased and decreased by miR-181a inhibitor (Figures 4A–4D). In addition, qPCR results revealed that MEG3 upregulation and downregulation respectively decreased and increased miR-181a expression (Figures 4E–4F). RNA-induced silencing complexes (RISCs) play a central role in posttranscriptional gene regulation. LncRNAs are a vital part of RISCs. MiRNAs bind their targets to repress and/or degrade RNA through RISC in an Ago2-dependent manner [23,24]. In the current study, RNA immunoprecipitation (RIP) assay results showed that MEG3 and miR-181a expression were increased in the anti-Ago2 group compared with that in the control group. The expression levels of MEG3 and miR-181a immunoprecipitated with Ago2 in anti-miR-181a were considerably decreased compared with those in the anti-normal IgG group (Figures 4G–4H). Thus, MEG3 interacted with miR-181a by a direct targeting and Ago2-dependent manner.
MiR-181a inhibition promoted fibrosis and inflammatory response in MCs
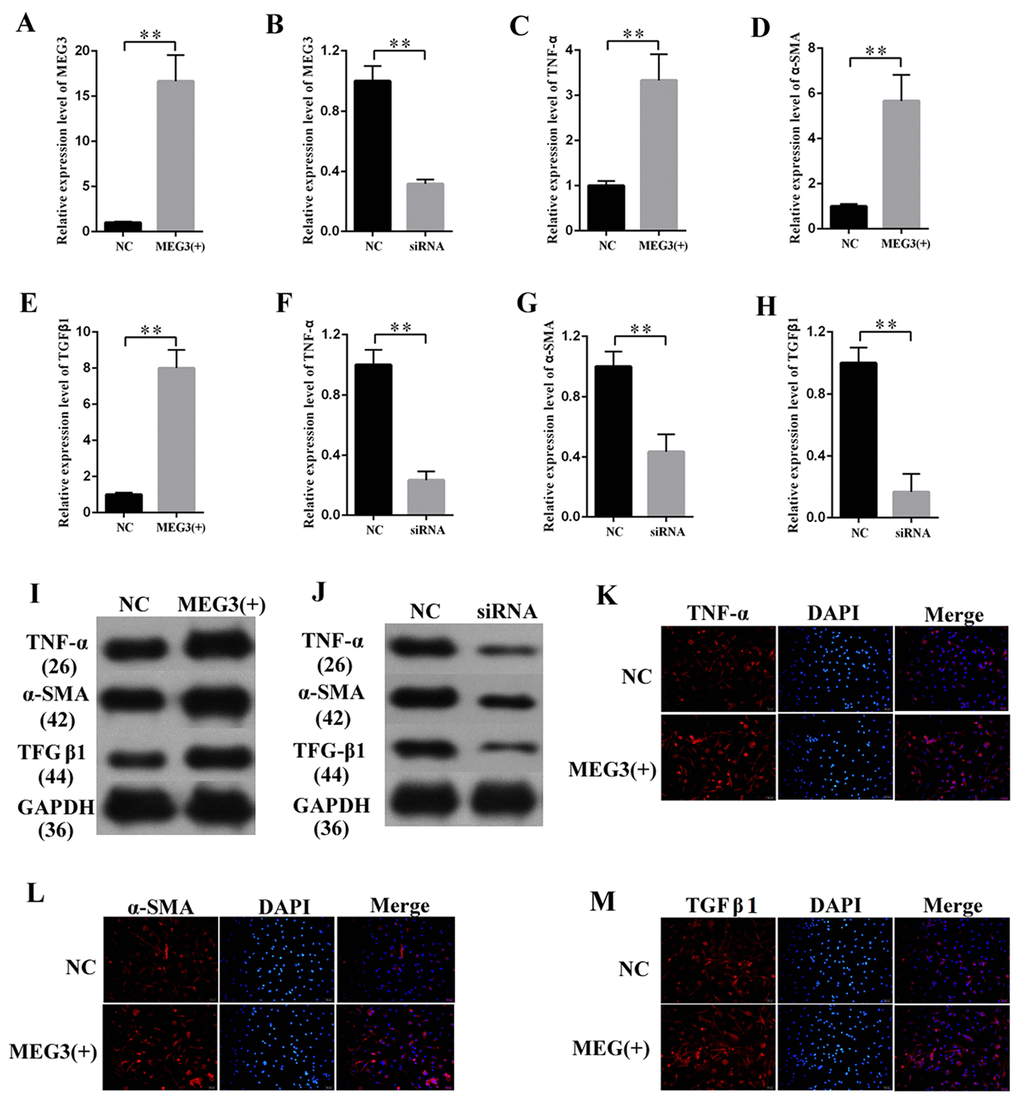
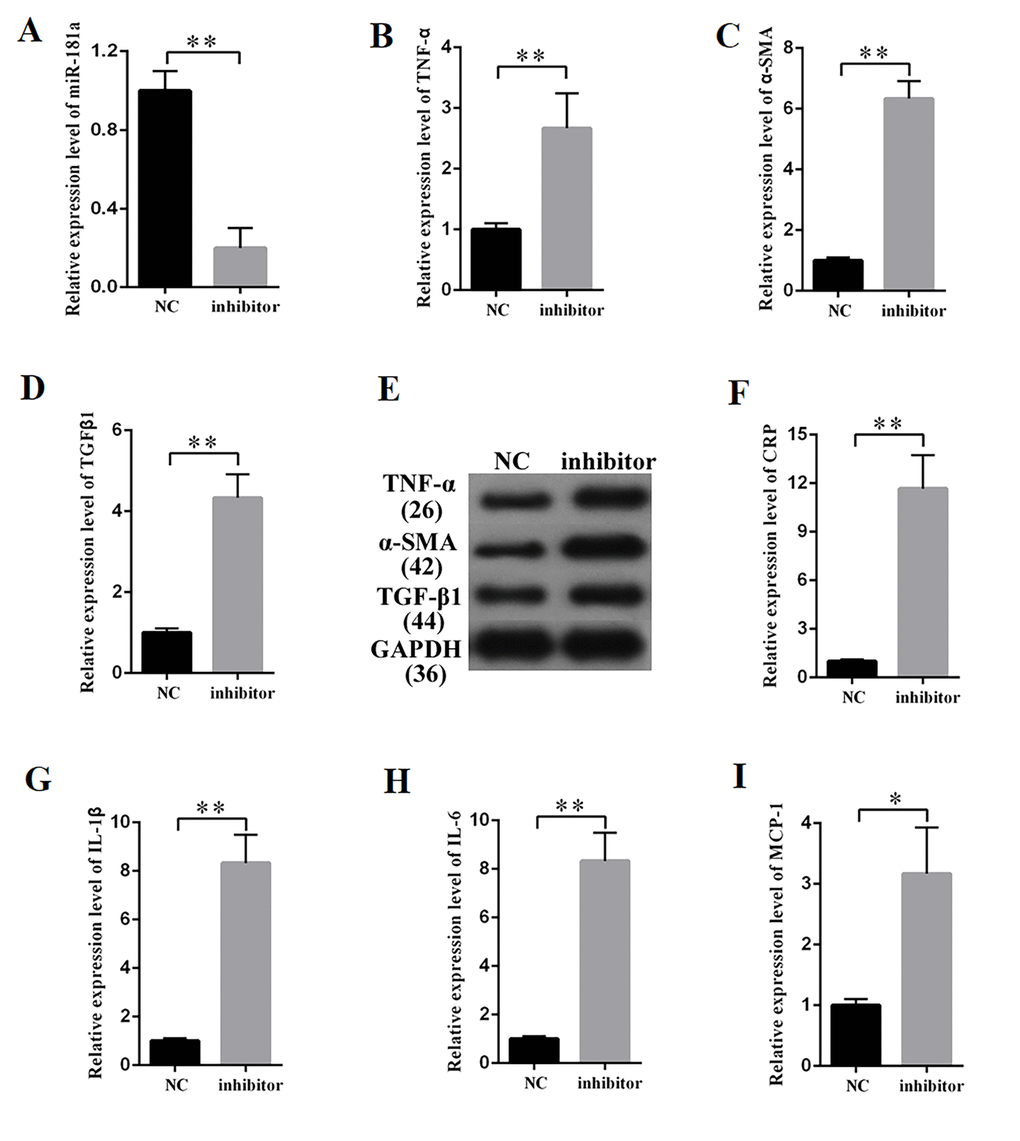
MiR-181a was knocked down to study the effects of miR-181a on the fibrosis and inflammatory response in MCs. The efficiency of a miR-181a inhibitor was determined by qPCR (Figure 5A), which revealed that miR-181a inhibition enhanced the mRNA expression of TNF-α, α-SMA, and TGF-β1 in MCs (Figures 5B, 5C, and 5D). Western blot results revealed that miR-181a inhibition increased TNF-α, α-SMA, and TGF-β1 protein expression in MCs (Figure 5E). Subsequently, the supernatant of MCs transfected with miR-181a was collected and subjected to the measurement of inflammatory cytokines by ELISA. ELISA analysis results revealed that miR-181a inhibition increased the concentrations of CRP, IL-1β, IL-6, and MCP-1 in MCs (Figures 5F, 5G, 5H, and 5I). In summary, miR-181a inhibition enhanced the expression of fibrosis-related proteins and the concentration of inflammatory cytokines in MCs.
MEG3 promoted Egr-1 expression, fibrosis, and inflammatory response by acting as a miR-181a sponge
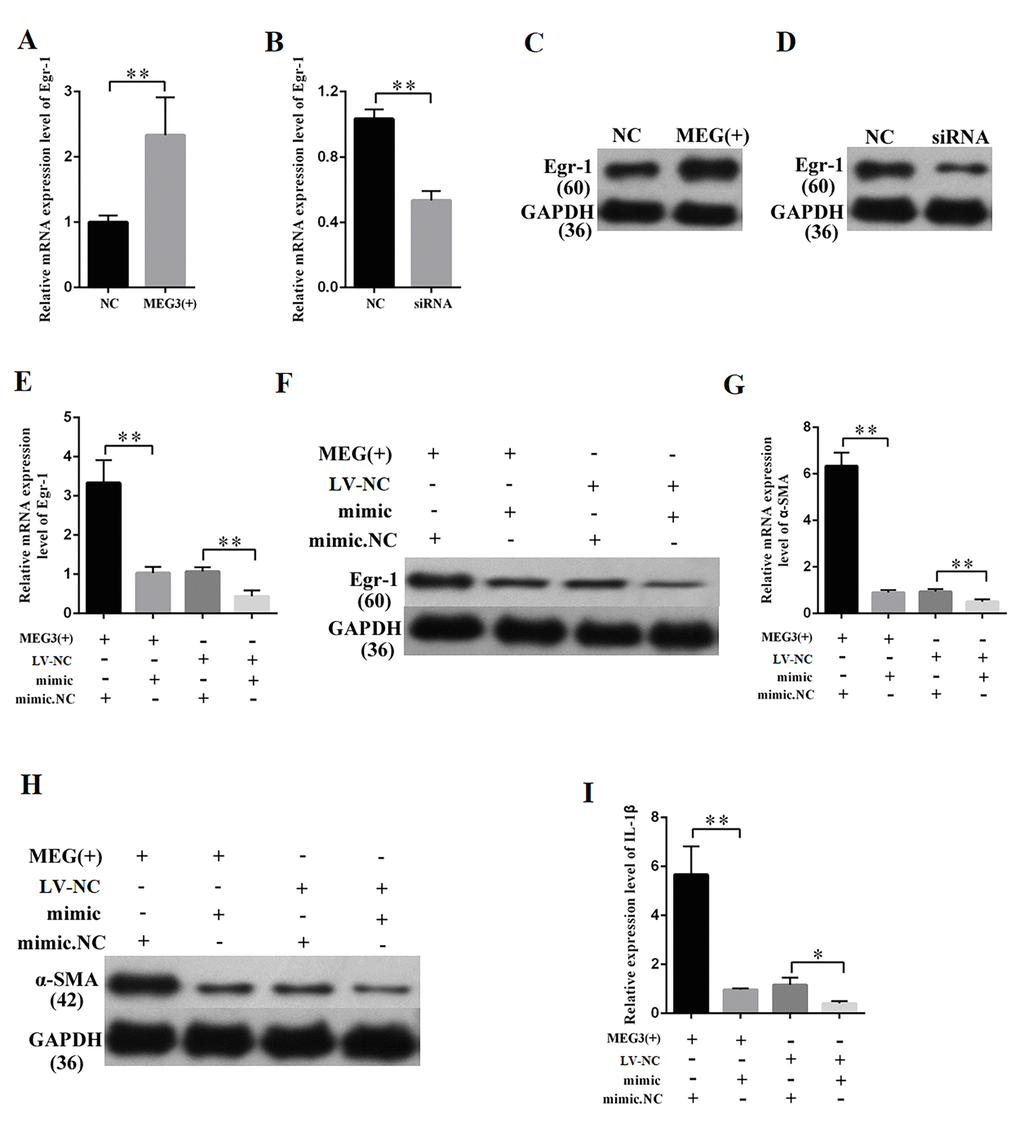
MEG3 overexpression and knockdown were performed to further study the relationship between MEG3 and Egr-1. QRT-PCR and Western blot results revealed that the overexpression and silencing of MEG3 respectively increased and decreased the Egr-1 expression (Figures 7A–7D). QRT-PCR and Western blot were then performed by respectively co-transfecting LV-MEG3 and mimic-miR-181a into MCs to investigate whether miR-181a mediates the function of MEG3. The results showed that mimic-miR-181a restored Egr-1 upregulation by MEG3 overexpression (Figures 7E–7F). QRT-PCR and Western blot indicated that miR-181a overexpression attenuated the promotive roles of MEG3 on α-SMA (Figures 7G–7H). ELISA indicated that miR-181a overexpression attenuated the promotive roles of MEG3 on IL-1β (Figure 7I). In summary, MEG3 upregulated Egr-1 expression and enhanced fibrosis and inflammatory response by acting as a miR-181a sponge.
MEG3 overexpression enhanced DN development in vivo
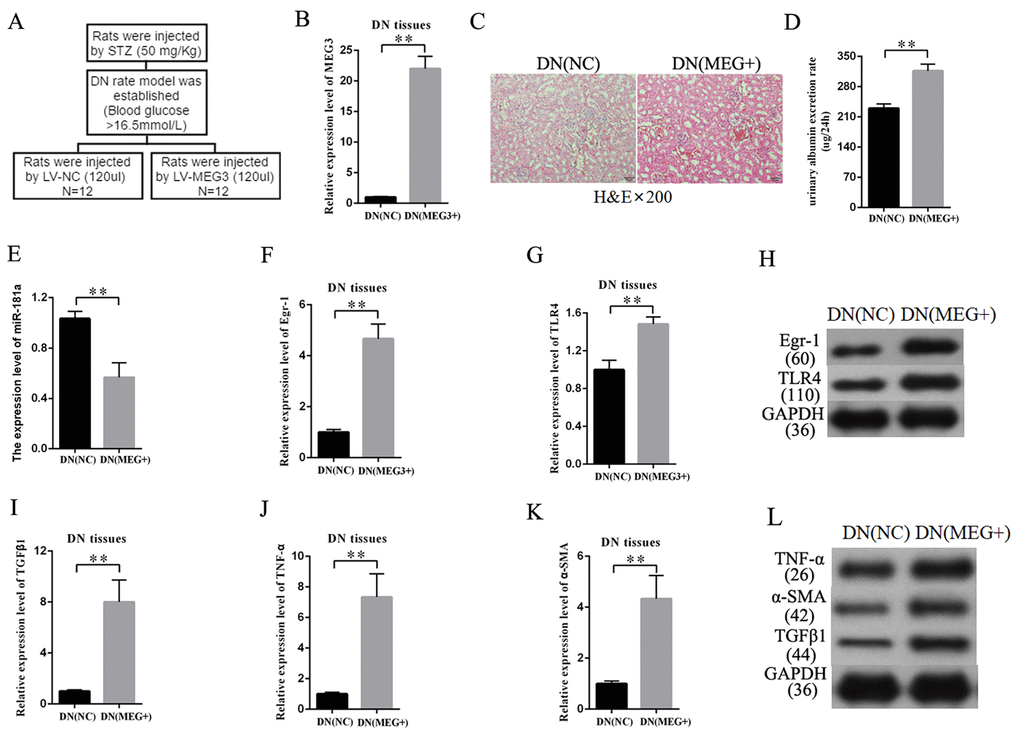
An overexpression-MEG3 DN rat model was built to further study the effects of MEG3 on DN in vivo. The establishment procedure is shown in Figure 8A. The efficiency of LV-MEG3 was detected in kidney tissues by qRT-PCR. The results indicated that MEG3 was successfully overexpressed in DN tissues after LV-MEG3 injection (Figure 8B). H&E staining results showed that MEG3 overexpression promoted DN development (Figure 8C) and increased urinary albumin excretion rate in DN rat models compared with that in the NC group (Figure 8D). The expression of miR-181a, Egr-1, and TLR4 was detected in kidney tissues to validate the participation of the miR-181a/Egr-1/TLR4 axis in the roles of MEG3 in DN. MEG3 overexpression decreased miR-181a expression (Figure 8E). QRT-PCR results indicated that Egr-1 and TLR4 expression was enhanced in DN tissues by LV-MEG3 injection (Figures 8F–8G). Western blot results indicated that MEG3 overexpression upregulated Egr-1 and TLR4 protein expression in DN tissues (Figure 8H). In addition, fibrosis-related genes were detected in the DN rat model. QPCR and Western blot results showed that TNF-α, α-SMA, and TGF-β1 were upregulated in the overexpression-MEG3 DN rat model (Figures 8I–8L). These results demonstrated that MEG3 overexpression enhanced DN development via the miR-181a/Egr-1/TLR4 axis.
Discussion
DN is a common cause of renal failure [27]. Medical treatment for this disease is currently unavailable [28]. Numerous patients with DN require kidney dialysis throughout life. Thus, understanding the potential molecular mechanism of DN is important for the discovery of novel therapeutic targets and the development of new specific drugs.
CeRNA has become an important mechanism of lncRNAs to participate in the regulation of coding gene expression in DN. LncRNA NR_033515 enhances EMT and fibrogenesis by acting as a sponge for miR-743b-5p in DN [18]. LncRNA 1700020I14Rik inhibits cell fibrosis and proliferation in DN by targeting miR-34a-5p [29]. LncRNA Erbb4-IR enhances DN-related injury by targeting miR-29b [30]. MEG3 plays a vital role in diabetes [31]. Genetic variants of MEG3 confer risk of type 2 diabetes in populations [32]. MEG3 enhances hepatic insulin resistance via miR-214/ATF4 axis [33]. However, the biological role and specific underlying mechanisms of MEG3 in DN are completely unknown.
In this research, we found that MEG3 expression level was remarkably enhanced in DN in vivo and vitro. QPCR and FISH results showed that MEG3 was mainly localized in the cytoplasm of MCs. MEG3 enhanced fibrosis and inflammatory response in MCs. MEG3 overexpression improved the expression of fibrosis-related proteins and inflammatory cytokines. Conversely, MEG3 silencing suppressed fibrosis and inflammatory response. MEG3 is mainly located in the cell cytoplasm, thereby suggesting that MEG3 also plays a role in MCs by ceRNA. Bioinformatics analysis predicted MEG3 putative binding sites with miR-181a. Luciferase assay and RIP assay showed that MEG3 interacts with miR-181a through direct targeting by and in an Ago2-dependent manner, respectively. MiR-181a inhibition suppressed fibrosis and inflammatory response in MCs. Egr-1 was demonstrated to be a direct target gene of miR-181a. Egr-1 is involved in the development of DN. Egr-1 silencing inhibits renal tubulointerstitial fibrosis in DN [25]. Egr-1 also increases pro-fibrotic gene expression in a high-sugar environment in HK-2 cells [34]. In the current study, a rescuing experiment revealed that MEG3 promoted Egr-1 expression, fibrosis, and inflammatory response in a ceRNA-dependent manner. Finally, MEG3 could enhance the development of DN in vivo. In DN rat models, MEG3 overexpression increased the urinary albumin excretion rate and upregulated the expression of fibrosis proteins. MEG3 upregulation decreased miR-181 expression and increased Egr-1 and TLR4 expression
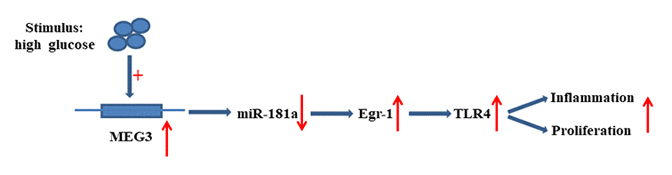
The present research revealed that MEG3 functions as ceRNA to upregulate Egr-1 expression by acting as a sponge for miR-181a. MEG3 promoted fibrosis and inflammatory response in DN. The relationship between MEG3, miR-181a, Egr1, and TLR4 is illustrated in Figure 9. Therefore, MEG3 may act as a potential therapeutic target and biomarker for further DN clinical application.
Materials and Methods
Cell culture
Cell culture was performed in accordance with a previous study [35]. RAW264.7 cells were cultured in RPMI1640 (Gibco, USA) supplemented with 20% FBS (Gibco, USA) in a 37 °C incubator with 5% CO2. MCs were cultured in 5.5 mmol/L glucose, low glucose (LG, 5.5 mmol/L glucose plus 19.5 mmol/L mannitol), or high glucose (HG, 25 mmol/L glucose).
Cell transfection, lentivirus production, and transduction
A lentiviral vector for MEG3 was constructed by Bio-Link Gene (China). MiR-181a mimic, miR-181a inhibitor, and the corresponding controls were purchased from RiboBio (Guangzhou, China). The sequence of Egr-1 was cloned into a pcDNA3.1 vector. An empty vector functioned as negative control. Lipofectamine 3000 (Invitrogen, USA) was used for transfection in accordance with the manufacturer’s instructions.
ELISA
ELISA was performed in accordance with a previous study [36]. The supernatants of MCs and DN tissues in lysis buffer were collected and measured by CRP, IL-1β, IL-6, and MCP-1 ELISA Kits (Boster Bioengineering Co, China) in accordance with the manufacturer’s instructions.
Isolation of nuclear and cytoplasmic RNA
Nuclear and cytoplasmic RNA were isolated in accordance with a previous work [16]. Cytoplasmic and nuclear RNA were extracted and separated using a Nuclear and Cytoplasmic RNA Purification Kit (Norgen, USA). The isolated RNA was detected by qRT-PCR.
Fluorescence in situ hybridization
MCs were permeabilized with pre-cooled Triton X-100 (0.1%), and the slides were treated with 4% paraformaldehyde. Hybridization was then performed overnight with the lncRNA MEG3 probe. The cells were washed with SCC buffer. DAPI was used to stain the coverslip, and laser scanning confocal microscopy was utilized to test fluorescence.
QRT-PCR
All RNAs were isolated using the TRIzol reagent (Life Technologies, US). MRNA expression was measured via SYBR Green qPCR on an ABI 7300 real-time PCR machine. TNF-α, α-SMA, and TGF-β1 mRNA expression levels were respectively examined using the following specific primers: F-5′-TTGTCCTCAGTTTCGGGAGAT-3′ R-5′-CAAGTCTGTGTCCTGAACGC-3′; F-5′-CCTGTGTTGTGGTTTACACTGG-3′ R-5′- GGGGGAATTATCTTTCCTGGT; F-5′-AGGAGCAAAATCTGTCCGATCT-3′ R-5′- GTGGGGGAATTATCTTTCCT.
Bioinformatics analysis
The candidate target miRNA of MEG3 was analyzed using StarBase v2.0 (http://starbase.sysu.edu.cn/index.php). The candidate target genes of miR-181a were analyzed via the miRNA target prediction program miRNA.org (http://www.microrna.org/microrna/home.do).
Western blot
Protein (16–20 mg) extracted from the samples was utilized for Western blot. Antibodies utilized in the research included anti-TNF-α (1:1000; HK), anti-α-SMA (1:1000; HK), anti-TGF-β1 (1:1000; HK), anti-Egr-1 (1:1000; HK), and anti-TLR4 (1:1000; HK), and GAPDH (1:1000; USA) was utilized as loading control.
Luciferase reporter assay
Luciferase reporter assay was performed according to a previous study [37]. Specifically, pmirGLO-MEG-WT or pmirGLO-Egr-1-WT reporter plasmids (120 ng) and corresponding mutated vectors were transfected into 293T cells with miR-181a inhibitor or miR181a mimic (100 nM) with Lipofectamine 3000. In accordance with standard protocols, 293T cells were lysed 48 h after transfection, and luciferase activity was measured using the PicaGene Dual SeaPansy luminescence kit (Toyo Ink, Japan).
RIP assays
RIP assay was performed according to the literature [35] to determine the association between 1700020I14Rik and the RISC. The cell lysate was incubated with RIP immunoprecipitation buffer, which contained magnetic beads conjugated with Ago2 antibody (Abcam, USA) and NC IgG (Abcam, Cambridge, USA). RIP experiments were performed using Magna RIP kit (Millipore, USA). Co-precipitated RNAs were detected via qRT-PCR.
Immunofluorescence assay
For immunofluorescence analysis, the fixed cells were permeabilized with 0.1% Triton X-100 and blocked in 5% goat serum solution. The coverslips were added with primary antibodies against TNF-α (HK, 1:50), α-SMA (HK, 1:50), or TGF-β1 (HK, 1:50) at 4 °C overnight and then with Cy3-labeled secondary antibody (Proteintech, USA, 1:50) for 1 h at room temperature. After treatment with DAPI, the coverslips were sealed with glycerine. Images were observed with QImaging Micro-publisherTM 5.0 RTV (Olympus Corporation, Japan) and analyzed with Image-Pro Plus (Medium Cybernetics, Bethesda, MD, USA).
In vivo study
Eight-week-old male rats (SD rats) were purchased from Shanghai Laboratory Animal Center (Shanghai, China) and kept in an SPF lab. Two groups were set up (N = 12 for each group): DN + LV‐NC group and DN + LV‐MEG3 group. Streptozotocin (STZ) was used to establish the DN rat model. STZ was dissolved in 0.1 M citrate acid buffer, which was used to build DN rat models (50 mg/kg). The rats were injected once daily for six days. Blood glucose levels of >16.5 mmol/L were identified as successful DN models. Lentiviral –MEG3 (LV-MEG3) was used to establish the DN rat model. LV‐MEG3 or LV‐NC (120 µl) was injected into DN models. All DN rat models were sacrificed at the end of the third week after injection. The study was approved by the Ethics Committee of Qingpu Branch of Zhongshan Hospital Affiliated to Fudan University, and the procedures were performed in accordance with the regulations of the Ethics Committee.
Statistical analysis
Student’s t-test and one-way ANOVA were performed for statistical analysis by using the GraphPad Prism 5.0. Statistical data are expressed as mean ± standard deviation (SD). Differences were considered significant at P < 0.05.
Supplementary Materials
Conflicts of Interest
The authors declare that there is no conflict of interest.
Funding
This work was supported by the Fund of Shanghai Municipal Commission of Health and Family Planning (N0. 201840254).
References
- 1. Long J, Badal SS, Ye Z, Wang Y, Ayanga BA, Galvan DL, Green NH, Chang BH, Overbeek PA, Danesh FR. Long noncoding RNA Tug1 regulates mitochondrial bioenergetics in diabetic nephropathy. J Clin Invest. 2016; 126:4205–18. https://doi.org/10.1172/JCI87927 [PubMed]
- 2. Madhusudhan T, Wang H, Dong W, Ghosh S, Bock F, Thangapandi VR, Ranjan S, Wolter J, Kohli S, Shahzad K, Heidel F, Krueger M, Schwenger V, et al. Defective podocyte insulin signalling through p85-XBP1 promotes ATF6-dependent maladaptive ER-stress response in diabetic nephropathy. Nat Commun. 2015; 6:6496. https://doi.org/10.1038/ncomms7496 [PubMed]
- 3. Bailey CJ. Potential new treatments for type 2 diabetes. Trends Pharmacol Sci. 2000; 21:259–65. https://doi.org/10.1016/S0165-6147(00)01506-6 [PubMed]
- 4. Cernea S, Pozzilli P. New potential treatments for protection of pancreatic B-cell function in Type 1 diabetes. Diabet Med. 2008; 25:1259–67. https://doi.org/10.1111/j.1464-5491.2008.02556.x [PubMed]
- 5. Kopp F, Mendell JT. Functional Classification and Experimental Dissection of Long Noncoding RNAs. Cell. 2018; 172:393–407. https://doi.org/10.1016/j.cell.2018.01.011 [PubMed]
- 6. Schmitt AM, Chang HY. Long Noncoding RNAs in Cancer Pathways. Cancer Cell. 2016; 29:452–63. https://doi.org/10.1016/j.ccell.2016.03.010 [PubMed]
- 7. Wang M, Yao D, Wang S, Yan Q, Lu W. Long non-coding RNA ENSMUST00000147869 protects mesangial cells from proliferation and fibrosis induced by diabetic nephropathy. Endocrine. 2016; 54:81–92. https://doi.org/10.1007/s12020-016-0950-5 [PubMed]
- 8. Zhou L, Xu DY, Sha WG, Shen L, Lu GY, Yin X. Long non-coding MIAT mediates high glucose-induced renal tubular epithelial injury. Biochem Biophys Res Commun. 2015; 468:726–32. https://doi.org/10.1016/j.bbrc.2015.11.023 [PubMed]
- 9. Jaulin-Bastard F, Arsanto JP, Le Bivic A, Navarro C, Vély F, Saito H, Marchetto S, Hatzfeld M, Santoni MJ, Birnbaum D, Borg JP. Interaction between Erbin and a Catenin-related protein in epithelial cells. J Biol Chem. 2002; 277:2869–75. https://doi.org/10.1074/jbc.M109652200 [PubMed]
- 10. Jin L, Cai Q, Wang S, Wang S, Mondal T, Wang J, Quan Z. Long noncoding RNA MEG3 regulates LATS2 by promoting the ubiquitination of EZH2 and inhibits proliferation and invasion in gallbladder cancer. Cell Death Dis. 2018; 9:1017. https://doi.org/10.1038/s41419-018-1064-1 [PubMed]
- 11. Zheng Q, Lin Z, Xu J, Lu Y, Meng Q, Wang C, Yang Y, Xin X, Li X, Pu H, Gui X, Li T, Xiong W, Lu D. Long noncoding RNA MEG3 suppresses liver cancer cells growth through inhibiting β-catenin by activating PKM2 and inactivating PTEN. Cell Death Dis. 2018; 9:253. https://doi.org/10.1038/s41419-018-0305-7 [PubMed]
- 12. Bartel DP. Metazoan MicroRNAs. Cell. 2018; 173:20–51. https://doi.org/10.1016/j.cell.2018.03.006 [PubMed]
- 13. Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016; 17:272–83. https://doi.org/10.1038/nrg.2016.20 [PubMed]
- 14. Mohajeri M, Banach M, Atkin SL, Butler AE, Ruscica M, Watts GF, Sahebkar A. MicroRNAs: Novel Molecular Targets and Response Modulators of Statin Therapy. Trends Pharmacol Sci. 2018; 39:967–81. https://doi.org/10.1016/j.tips.2018.09.005 [PubMed]
- 15. Song YX, Sun JX, Zhao JH, Yang YC, Shi JX, Wu ZH, Chen XW, Gao P, Miao ZF, Wang ZN. Non-coding RNAs participate in the regulatory network of CLDN4 via ceRNA mediated miRNA evasion. Nat Commun. 2017; 8:289. https://doi.org/10.1038/s41467-017-00304-1 [PubMed]
- 16. Bai X, Geng J, Li X, Wan J, Liu J, Zhou Z, Liu X. Long Noncoding RNA LINC01619 Regulates MicroRNA-27a/Forkhead Box Protein O1 and Endoplasmic Reticulum Stress-Mediated Podocyte Injury in Diabetic Nephropathy. Antioxid Redox Signal. 2018; 29:355–76. https://doi.org/10.1089/ars.2017.7278 [PubMed]
- 17. Wang X, Xu Y, Zhu YC, Wang YK, Li J, Li XY, Ji T, Bai SJ. LncRNA NEAT1 promotes extracellular matrix accumulation and epithelial-to-mesenchymal transition by targeting miR-27b-3p and ZEB1 in diabetic nephropathy. J Cell Physiol. 2019; 234:12926–33. https://doi.org/10.1002/jcp.27959 [PubMed]
- 18. Gao J, Wang W, Wang F, Guo C. LncRNA-NR_033515 promotes proliferation, fibrogenesis and epithelial-to-mesenchymal transition by targeting miR-743b-5p in diabetic nephropathy. Biomed Pharmacother. 2018; 106:543–52. https://doi.org/10.1016/j.biopha.2018.06.104 [PubMed]
- 19. Peng W, Si S, Zhang Q, Li C, Zhao F, Wang F, Yu J, Ma R. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate gastric cancer progression. J Exp Clin Cancer Res. 2015; 34:79. https://doi.org/10.1186/s13046-015-0197-7 [PubMed]
- 20. Ghaedi H, Mozaffari MA, Salehi Z, Ghasemi H, Zadian SS, Alipoor S, Hadianpour S, Alipoor B. Co-expression profiling of plasma miRNAs and long noncoding RNAs in gastric cancer patients. Gene. 2019; 687:135–42. https://doi.org/10.1016/j.gene.2018.11.034 [PubMed]
- 21. Chen P, Pan J, Zhang X, Shi Z, Yang X. The Role of MicroRNA-181a in Myocardial Fibrosis Following Myocardial Infarction in a Rat Model. Med Sci Monit. 2018; 24:4121–27. https://doi.org/10.12659/MSM.908056 [PubMed]
- 22. Song J, Yang S, Yin R, Xiao Q, Ma A, Pan X. MicroRNA-181a regulates the activation of the NLRP3 inflammatory pathway by targeting MEK1 in THP-1 macrophages stimulated by ox-LDL. J Cell Biochem. 2019. https://doi.org/10.1002/jcb.28637 [PubMed]
- 23. Yoon JH, Abdelmohsen K, Srikantan S, Yang X, Martindale JL, De S, Huarte M, Zhan M, Becker KG, Gorospe M. LincRNA-p21 suppresses target mRNA translation. Mol Cell. 2012; 47:648–55. https://doi.org/10.1016/j.molcel.2012.06.027 [PubMed]
- 24. Chiyomaru T, Fukuhara S, Saini S, Majid S, Deng G, Shahryari V, Chang I, Tanaka Y, Enokida H, Nakagawa M, Dahiya R, Yamamura S. Long non-coding RNA HOTAIR is targeted and regulated by miR-141 in human cancer cells. J Biol Chem. 2014; 289:12550–65. https://doi.org/10.1074/jbc.M113.488593 [PubMed]
- 25. Liu F, Zhang ZP, Xin GD, Guo LH, Jiang Q, Wang ZX. miR-192 prevents renal tubulointerstitial fibrosis in diabetic nephropathy by targeting Egr1. Eur Rev Med Pharmacol Sci. 2018; 22:4252–60. https://doi.org/10.26355/eurrev_201807_15420 [PubMed]
- 26. Ha YM, Park EJ, Kang YJ, Park SW, Kim HJ, Chang KC. Valsartan independent of AT1 receptor inhibits tissue factor, TLR-2 and -4 expression by regulation of Egr-1 through activation of AMPK in diabetic conditions. J Cell Mol Med. 2014; 18:2031–43. https://doi.org/10.1111/jcmm.12354 [PubMed]
- 27. Rask-Madsen C, King GL. Vascular complications of diabetes: mechanisms of injury and protective factors. Cell Metab. 2013; 17:20–33. https://doi.org/10.1016/j.cmet.2012.11.012 [PubMed]
- 28. Fineberg D, Jandeleit-Dahm KA, Cooper ME. Diabetic nephropathy: diagnosis and treatment. Nat Rev Endocrinol. 2013; 9:713–23. https://doi.org/10.1038/nrendo.2013.184 [PubMed]
- 29. Lee FT, Mountain AJ, Kelly MP, Hall C, Rigopoulos A, Johns TG, Smyth FE, Brechbiel MW, Nice EC, Burgess AW, Scott AM. Enhanced efficacy of radioimmunotherapy with 90Y-CHX-A’'-DTPA-hu3S193 by inhibition of epidermal growth factor receptor (EGFR) signaling with EGFR tyrosine kinase inhibitor AG1478. Clin Cancer Res. 2005; 11:7080s–86s. https://doi.org/10.1158/1078-0432.CCR-1004-0019 [PubMed]
- 30. Sun SF, Tang PM, Feng M, Xiao J, Huang XR, Li P, Ma RC, Lan HY. Novel lncRNA Erbb4-IR Promotes Diabetic Kidney Injury in db/db Mice by Targeting miR-29b. Diabetes. 2018; 67:731–44. https://doi.org/10.2337/db17-0816 [PubMed]
- 31. Sathishkumar C, Prabu P, Mohan V, Balasubramanyam M. Linking a role of lncRNAs (long non-coding RNAs) with insulin resistance, accelerated senescence, and inflammation in patients with type 2 diabetes. Hum Genomics. 2018; 12:41. https://doi.org/10.1186/s40246-018-0173-3 [PubMed]
- 32. Ghaedi H, Zare A, Omrani MD, Doustimotlagh AH, Meshkani R, Alipoor S, Alipoor B. Genetic variants in long noncoding RNA H19 and MEG3 confer risk of type 2 diabetes in an Iranian population. Gene. 2018; 675:265–71. https://doi.org/10.1016/j.gene.2018.07.002 [PubMed]
- 33. Zhu X, Li H, Wu Y, Zhou J, Yang G, Wang W. lncRNA MEG3 promotes hepatic insulin resistance by serving as a competing endogenous RNA of miR-214 to regulate ATF4 expression. Int J Mol Med. 2019; 43:345–57. https://doi.org/10.3892/ijmm.2018.3975 [PubMed]
- 34. Xu P, Guan MP, Bi JG, Wang D, Zheng ZJ, Xue YM. High glucose down-regulates microRNA-181a-5p to increase pro-fibrotic gene expression by targeting early growth response factor 1 in HK-2 cells. Cell Signal. 2017; 31:96–104. https://doi.org/10.1016/j.cellsig.2017.01.012 [PubMed]
- 35. Li A, Peng R, Sun Y, Liu H, Peng H, Zhang Z. LincRNA 1700020I14Rik alleviates cell proliferation and fibrosis in diabetic nephropathy via miR-34a-5p/Sirt1/HIF-1α signaling. Cell Death Dis. 2018; 9:461. https://doi.org/10.1038/s41419-018-0527-8 [PubMed]
- 36. Guo J, Li J, Zhao J, Yang S, Wang L, Cheng G, Liu D, Xiao J, Liu Z, Zhao Z. MiRNA-29c regulates the expression of inflammatory cytokines in diabetic nephropathy by targeting tristetraprolin. Sci Rep. 2017; 7:2314. https://doi.org/10.1038/s41598-017-01027-5 [PubMed]
- 37. Huang T, Yin L, Wu J, Gu JJ, Wu JZ, Chen D, Yu HL, Ding K, Zhang N, Du MY, Qian LX, Lu ZW, He X. MicroRNA-19b-3p regulates nasopharyngeal carcinoma radiosensitivity by targeting TNFAIP3/NF-κB axis. J Exp Clin Cancer Res. 2016; 35:188. https://doi.org/10.1186/s13046-016-0465-1 [PubMed]