Introduction
Gastric cancer (GC) is a common cancer, causing significant human mortalities each year [1]. In the past decades, due to the developments of sanitation, control of Helicobacter pylori infection and GC screenings, its incidence is declined [2, 3]. However, the prognosis for the metastatic, recurrent, and other advanced GCs is not optimistic [1]. Identification of novel therapeutic targets and biomarkers for GC is extremely important [4, 5]. It is the research focus of our group [6, 7].
LMX1A (LIM homeobox transcription factor 1α) is the primary member of LIM-homeodomain conserved family transcription factors [8]. LMX1A regulates a number of physiological and cancerous behaviors [8]. LMX1A is downregulated in GC [7] and many other cancers [9, 10]. Our group has shown that LMX1A exerted tumor-suppressive functions in GC cells [6, 7, 11]. We found that LMX1A inhibited c-Myc expression to exert tumor suppressive function in GC cells [6]. Also, LMX1A inhibited GC cell metastasis by inhibiting beta-catenin-dependent genes [11].
microRNAs (miRNAs) are small, noncoding RNAs, regulating target gene expression by suppressing mRNAs translation and/or inducing their degradation. Our previous study has identified a LMX1A-targeting miRNA: microRNA-9 (miR-9). We show that miR-9 selectively targets and negatively regulates LMX1A to promote GC cell progression [7]. In human GC tissues, miR-9 upregulation is correlated with LMX1A downregulation [7]. The mechanism of miR-9 upregulation in human GC is still elusive.
Long non-coding RNAs (LncRNAs) are a large family of conserved non-coding RNAs (ncRNAs) with over 200-nucleotide long [12–14]. LncRNA can decrease target miRNA expression by acting as their sponges [15]. LncRNAs regulate key cellular activities, including genomic imprinting, cell proliferation and survival, cell cycle control and differentiation as well as apoptosis and transformation [12–14]. Accumulative studies have confirmed that dysregulation of LncRNA plays a pivotal role in the progression of GC [15, 16] and other human cancers [17, 18]. One aim of this study is to identify possible miR-9-targeting LncRNA, regulating LMX1A expression and GC cell functions.
LncRNAKCNQ1OT1, or potassium voltage-gated channel subfamily Q member 1 (Kcnq1) overlapping transcript 1, is an imprinted antisense LncRNA [19, 20]. It is an overlapping transcript of Kcnq1, locating at Kcnq1 loci on chromosome 11p15.5, being exclusively transcribed from the paternal chromosome [19, 20]. Recent studies have shown that KCNQ1OT1 dysregulation participates in carcinogenesis and progression of human cancers [21, 22]. Its expression and potential functions in GC are largely unknown. In the present study, we will show that LncRNAKCNQ1OT1 inhibits GC cell progression possibly via regulating miR-9 and LMX1A expression.
Results
LncRNA KCNQ1OT1 is downregulated in human GC tissues
It is possible that miR-9 upregulation in GC (see our previous study [7]) could be due to downregulation of certain LncRNAs. As they could function as the ceRNAs of miR-9. To explore this possibility, the human LncRNA database, LncBase (Predicted v.2), was consulted to identify possible miR-9-targeting LncRNAs. These potential LncRNAs were further verified by other LncRNAs/miR databases, including StarBase and miRbase. The bioinformatic analyses identified that one LncRNA, KCNQ1OT1, putatively targeted miR-9. Its expression in GC tissues was tested next. Total RNA was extracted from fresh GC tissues and the adjacent normal epithelial tissues from twelve (12) primary GC patients [7]. KCNQ1OT1 was determined by qPCR assay, using the previously-described primers [23].
The results show that KCNQ1OT1 levels are significantly downregulated in cancer tissues (“Can”) (Figure 1A), as compared to its levels in the surrounding epithelial (“Epi”) tissues (Figure 1A). Therefore, in GC cancer tissues KCNQ1OT1 downregulation correlated with miR-9 upregulation (see the previous results from same set of tissue samples [7]).
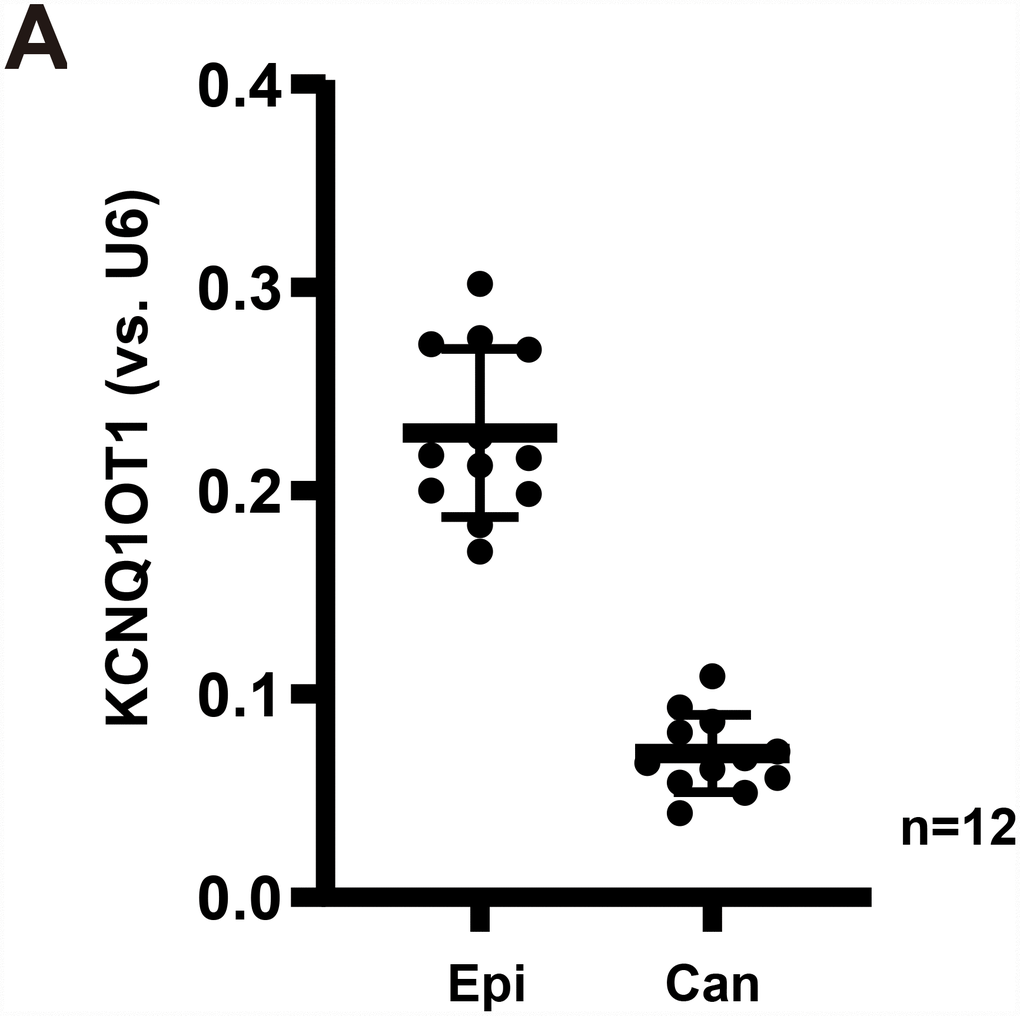
Figure 1. LncRNAKCNQ1OT1 is downregulated in human GC tissues. Expression of LncRNAKCNQ1OT1 (A) in twelve (12) different human gastric cancer (GC) tissues (“Can”) and matched surrounding normal epithelial tissues (“Epi”) was shown, results were normalized to U6 RNA. All values were expressed as the mean ± standard deviation (Same for all Figures). *P <0.05 vs. “Epi” tissues.
Forced overexpression of LncRNA KCNQ1OT1 induces miR-9 depletion and LMX1A upregulation in AGS cells
In order to test the potential effect of LncRNAKCNQ1OT1 on miR-9-LMX1A expression, a lentiviral GV248 construct encoding KCNQ1OT1 promoter (“LV-KCNQ1OT1”) was transfected to AGS cells. Following selection by puromycin, two stable cell lines (namely “sL1/sL2”) were established. Testing KCNQ1OT1 expression, by qPCR assays, demonstrated that mature KCNQ1OT1 levels increased over 8 to 10 folds in the stable cells (Figure 2A). Significantly, with KCNQ1OT1 overexpression miR-9 levels were dramatically downregulated (over 60–70%, Figure 2B), resulting in increased LMX1A 3′-UTR luciferase activity (Figure 2C). Moreover, LMX1A mRNA (Figure 2D) and protein (Figure 2E and 2F) levels were significantly increased in AGS cells with LV-KCNQ1OT1. Therefore, KCNQ1OT1 overexpression downregulated miR-9, leading to upregulation of its target LMX1A in AGS cells. The lentiviral control vector (“Vec”) had no significant effect on KCNQ1OT1-miR-9-LMX1A expression in AGS cells (Figure 2A–2F).
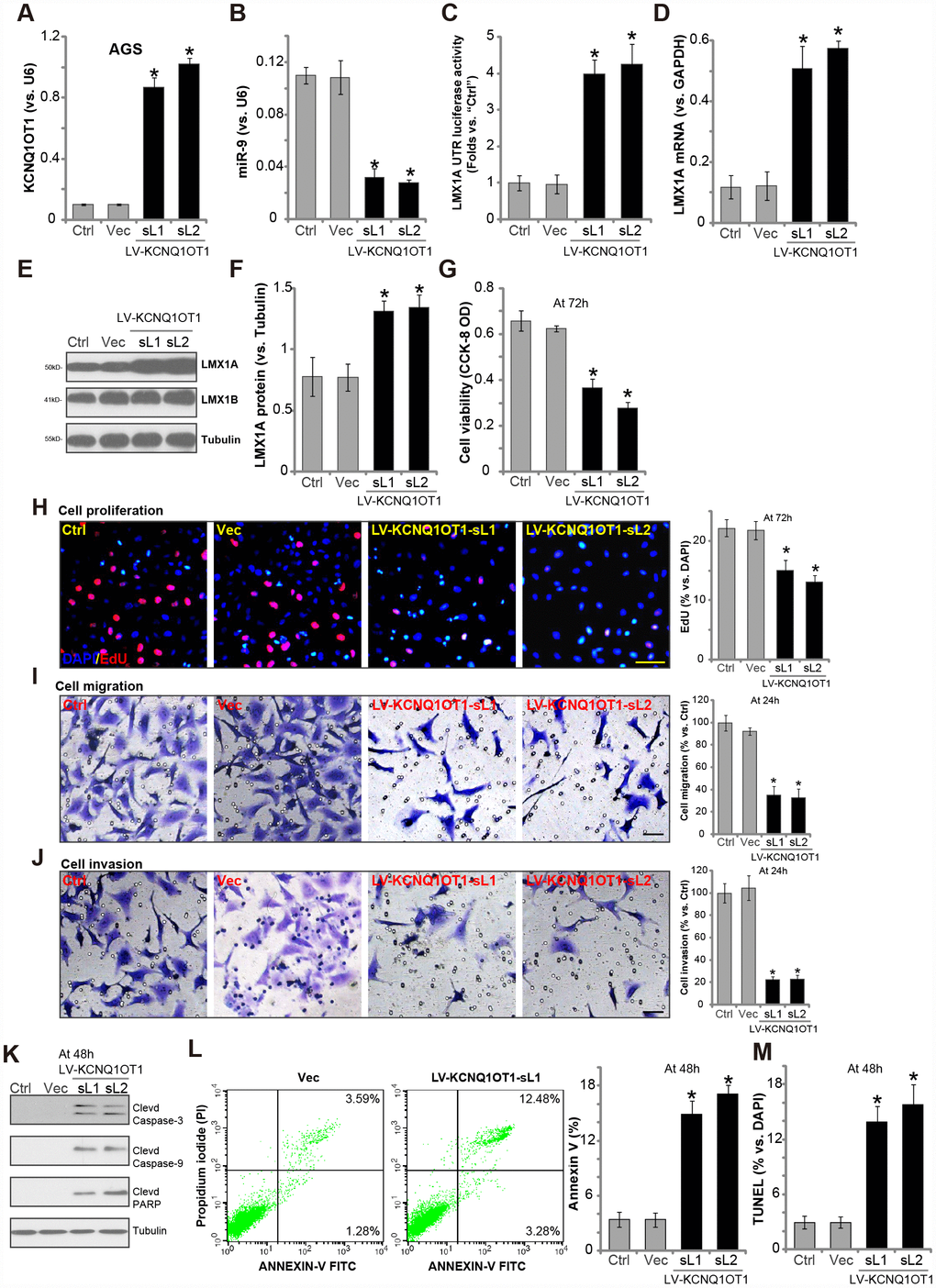
Figure 2. Forced overexpression of LncRNAKCNQ1OT1 induces miR-9 depletion, LMX1A upregulation and AGS cell inhibition. AGS cells were infected with KCNQ1OT1-expressing lentivirus (“LV-KCNQ1OT1”) or scramble control vector lentivirus (“Vec”) for 24h, following puromycin selection two stable lines (“sL1/sL-2”) with LV-KCNQ1OT1 established, expression of KCNQ1OT1 (A), miR-9 (B) and LMX1A mRNA (D) were tested by qPCR assays; The LMX1A 3’-UTR luciferase activity was shown (C); Expression of the listed proteins in total cell lysates were tested by Western blotting assay (E, results quantified in F); Cells were further cultured for the indicated time periods, cell survival and proliferation were tested by CCK-8 assay (G) and EdU staining (H), respectively; Cell migration and invasion were tested by “Transwell” (I) and “Matrigel Transwell” assay (J), respectively; Cell apoptosis was tested by Western blotting (testing apoptosis-associated proteins, K), Annexin V FACS (L) and TUNEL staining assay (M). The exact same number of viable cells of different genetic treatment were plated initially (at 0h) for the functional assays (Same for all following Figures). “Ctrl” stands for the parental control cells (Same for all Figures). For each assay, n=5. *P <0.05 vs. “Vec” cells. Experiments in this figure were repeated five times, and similar results were obtained. Bar=100 μm (H–J).
Forced overexpression of LncRNAKCNQ1OT1 inhibits AGS cell progression in vitro
LMX1A regulates a number of physiological and pathological processes [8]. It is considered as a tumor suppressor [8, 9, 11]. LMX1A downregulation is detected in GC and many other cancers [9, 10]. Our previous study has shown that LMX1A overexpression can inhibit AGS cell progression in vitro. We therefore analyzed the functional consequences of KCNQ1OT1 overexpression.
Results showed that AGS cells with LV-KCNQ1OT1 presented with reduced cell viability (vs. control cells, Figure 2G). Furthermore, KCNQ1OT1 overexpression inhibited EdU incorporation (Figure 2H) in AGS cells, indicating proliferation inhibition. “Transwell” and “Matrigel Transwell” assay results, in Figure 2I and 2J, demonstrated that ectopic overexpression of KCNQ1OT1 potently inhibited AGS cell migration and invasion in vitro. Additionally, apoptosis activation was detected in KCNQ1OT1-overexpressed AGS cells, evidenced by cleavages of caspase-3, caspase-9 and poly adenosine diphosphate (ADP) ribose (PARP) (Figure 2K), as well as increased Annexin V staining (Figure 2L) and nuclear TUNEL ratio (Figure 2M). The lentiviral control vector (“Vec”) had no significant effect on the function of AGS cells (Figure 2G–2M).
Forced overexpression of LncRNAKCNQ1OT1 induces miR-9 reduction and LMX1A upregulation in primary human GC cells
Next, we tested whether LncRNAKCNQ1OT1 exerted similar functions in primary human GC cells. As described previously [7], the primary human GC cells, derived from two independent patients (namely “GC-1/GC-2”), were infected with LV-KCNQ1OT1, which resulted in 5-6 folds increase of KCNQ1OT1 expression (Figure 3A). On the contrary, miR-9 levels were significantly decreased (Figure 3B), with LMX1A mRNA (Figure 3C) and protein (Figure 3D) levels increased in KCNQ1OT1-overexpressed GC cells. LMX1B protein expression was unaffected by KCNQ1OT1 overexpression (Figure 3D).
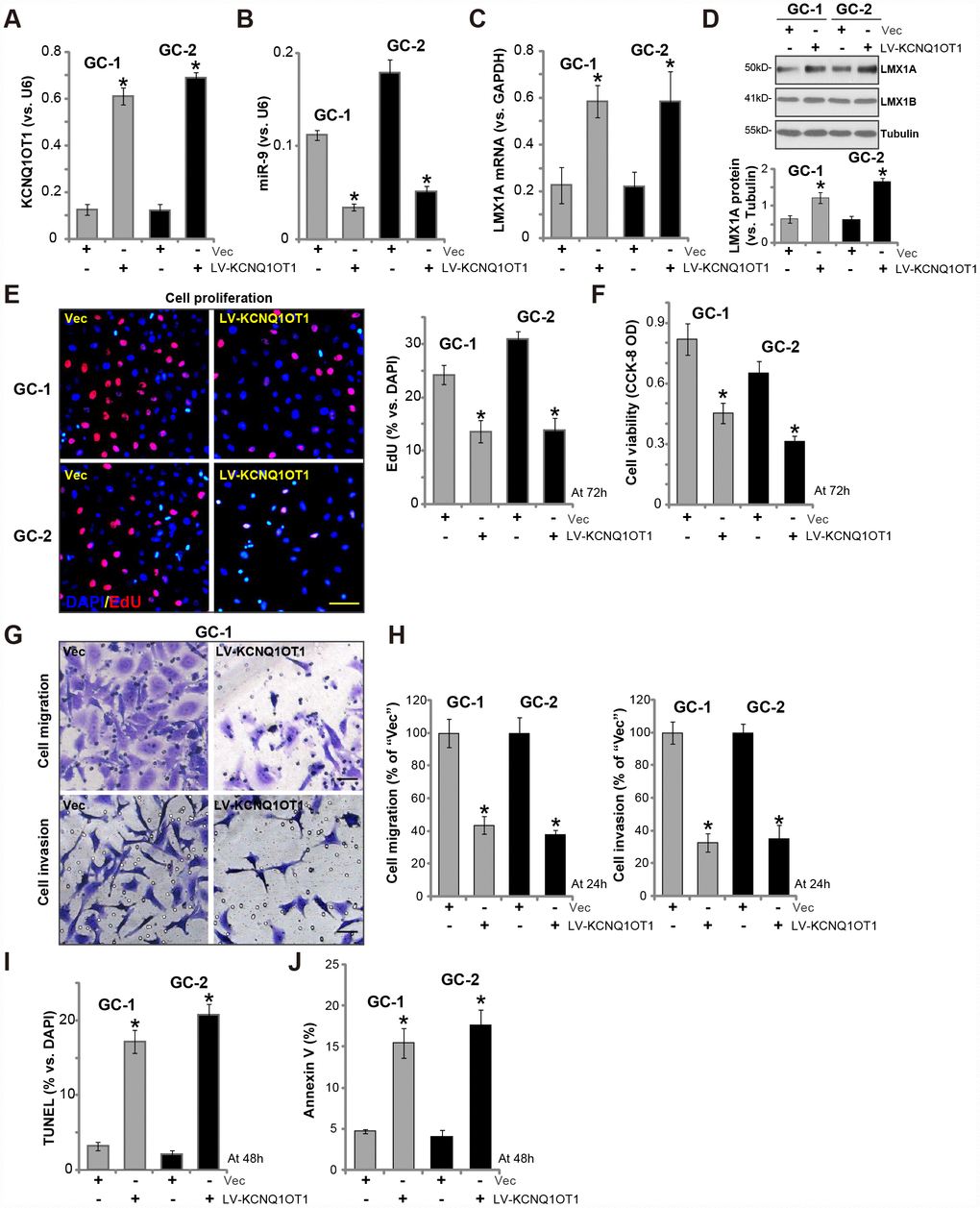
Figure 3. Forced overexpression of LncRNAKCNQ1OT1 induces miR-9 reduction and LMX1A upregulation in primary human GC cells. The primary human GC cells (“GC-1/GC-2”) were infected with LncRNAKCNQ1OT1-expressing lentivirus (“LV-KCNQ1OT1”) or scramble control vector lentivirus (“Vec”) for 72h, expression of KCNQ1OT1 (A), miR-9 (B) and LMX1A mRNA (C) were tested by qPCR assay; LMX1A and LMX1B protein expression in total cell lysates was tested by Western blotting assay (D, LMX1A protein results were quantified); Cells were further cultured for the indicated time periods, cell proliferation and viability were tested by EdU staining assay (E) and CCK-8 (F), respectively, with cell migration and invasion tested by “Transwell” and “Matrigel Transwell” assay (G and H); Cell apoptosis was quantified via the TUNEL staining assay (I) and Annexin V-FACS assay (J). For each assay, n=5. *P <0.05 vs. “Vec” cells. Experiments in this figure were repeated three times, and similar results were obtained. Bar=100 μm (E and G).
Functional studies show that KCNQ1OT1 overexpression by LV-KCNQ1OT1 inhibited EdU incorporation (Figure 3E) and cell viability (Figure 3F) in primary human GC cells. Furthermore, LV-KCNQ1OT1-expressing primary GC cells presented with decreased cell migration and invasion (vs. Vector control cells, Figure 3G, 3H). Contrarily, the increased TUNEL staining (Figure 3I, vs control cells) and Annexin V ratio (Figure 3J) were detected in LV-KCNQ1OT1-expressing primary GC cells, indicating apoptosis activation. Therefore, ectopic KCNQ1OT1 overexpression inhibited viability, proliferation, migration and invasion, but inducing apoptosis activation in primary GC cells.
LncRNA KCNQ1OT1 siRNA induces miR-9 upregulation and LMX1A downregulation, promoting GC cell progression in vitro
Next, the siRNA strategy was employed to knockdown KCNQ1OT1. Two siRNAs, targeting non-overlapping sequence of KCNQ1OT1 (namely “seq1/seq2” [24]), were transfected to AGS cells. Analyzing KCNQ1OT1 expression by qPCR confirmed that the applied siRNAs resulted in over 60-70% reduction of KCNQ1OT1 expression in AGS cells (Figure 4A). Significantly, KCNQ1OT1 siRNA induced miR-9 upregulation (Figure 4B) and LMX1A mRNA/protein downregulation (Figure 4C). Importantly, KCNQ1OT1-silenced cells presented with increased cell viability (Figure 4D) and EdU staining (Figure 4E), as compared to cells with scramble control siRNA (“si-C”) (Figure 4D, 4E). AGS cell migration and invasion, tested by “Transwell” assay (Figure 4F) and “Matrigel Transwell” assay (Figure 4G), were augmented by KCNQ1OT1 siRNAs as well.
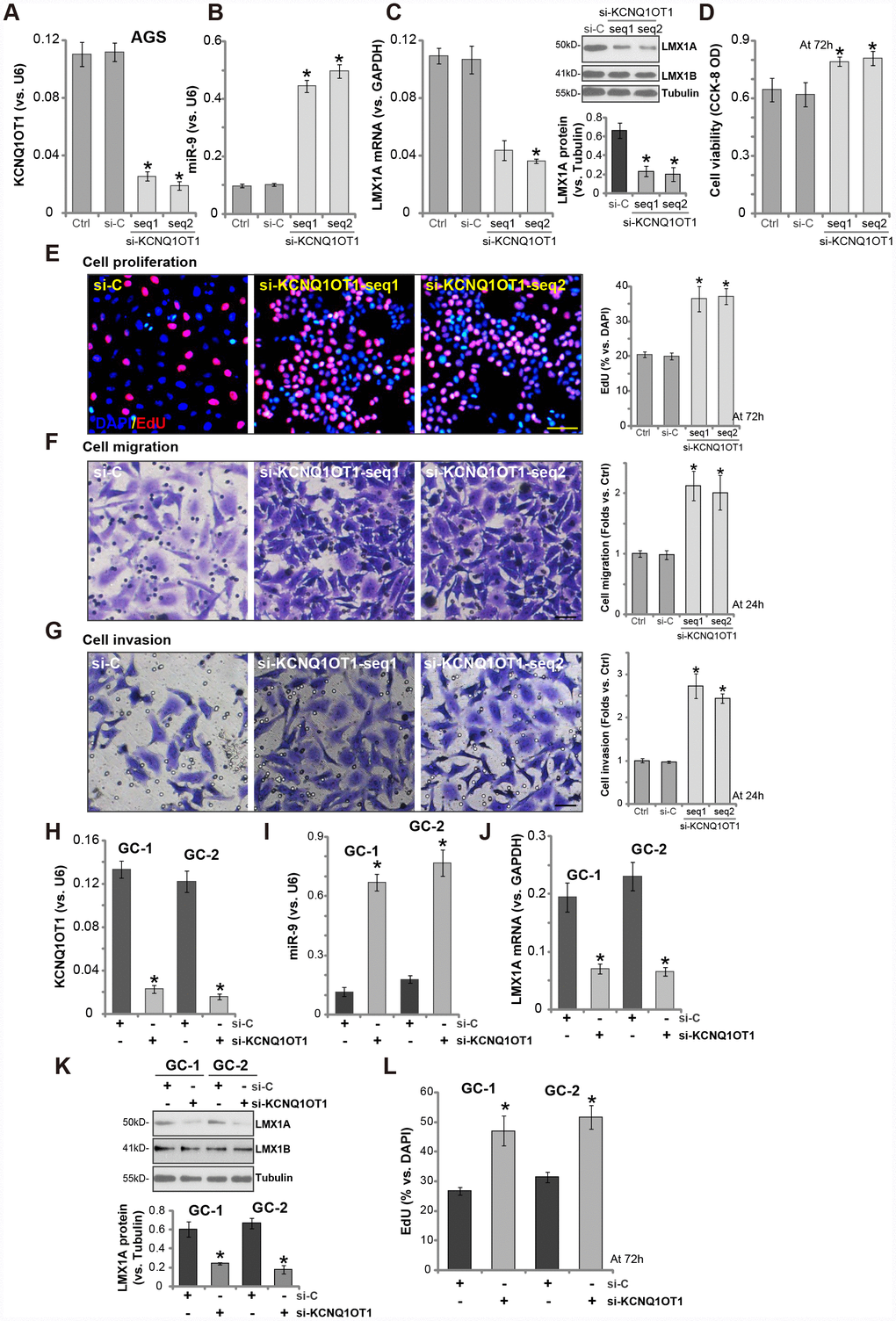
Figure 4. LncRNA KCNQ1OT1 siRNA induces miR-9 upregulation and LMX1A downregulation, promoting GC cell progression in vitro. AGS cells (A–G) or primary human GC cells (“GC-1/GC-2”) (H–L) were transfected with 500 nM of KCNQ1OT1 siRNA (“seq1/seq2”, two rounds, total 48h) or the scramble control siRNA (“si-C”, two rounds, total 48h), expression of KCNQ1OT1 (A and H), miR-9 (B and I) and LMX1A mRNA (C and J) were tested by qPCR assays; LMX1A and LMX1B protein expression was tested by Western blotting assay (C and K, LMX1A protein results were quantified); Cells were further cultured for the indicated time periods, cell viability (D) and proliferation (E and L) were tested by the listed assays; Cell migration and invasion were tested by “Transwell” (F) and “Matrigel Transwell” (G) assays respectively. For each assay, n=5. *P <0.05 vs. “si-C” cells. Experiments in this figure were repeated three times, and similar results were obtained. Bar=100 μm (E–G).
In primary human GC cells (“GC-1/GC-2”), transfection with the KCNQ1OT1 siRNA (“seq2”) similarly resulted in KCNQ1OT1 downregulation (Figure 4H), miR-9 upregulation (Figure 4I) and LMX1A downregulation (Figure 4J, 4K). Cell proliferation, tested by an EdU staining assay, was enhanced as well (Figure 4L). These results further confirmed a tumor-suppressive function by KCNQ1OT1, and KCNQ1OT1 siRNA promoted GC cell progression in vitro.
In LMX1A-knockout AGS cells altering KCNQ1OT1 expression fails to change cell viability and proliferation
To verify that LMX1A is one target of KCNQ1OT1, we utilized the CRISPR/Cas9 gene-editing method to knockout LMX1A (see our previous study [7]). Two stable LMX1A knockout (“Cas9-LMX1A-ko”) AGS cell lines were established (namely “S1/S2”) [7]. Results in Figure 5A confirmed LMX1A protein depletion in the stable cells, with LMX1B unaffected (Figure 5A). LV-KCNQ1OT1 (see Figure 2) or KCNQ1OT1 siRNA (“seq2”, see Figure 3) were tranduced to the Cas9-LMX1A-ko cells, both significantly altered KCNQ1OT1 expression (Figure 5B). Significantly, neither LV-KCNQ1OT1 nor KCNQ1OT1 siRNA altered viability (Figure 5C) and proliferation (EdU ratio, Figure 5D) in Cas9-LMX1A-ko AGS cells. Thus, in LMX1A-knockout AGS cells KCNQ1OT1 did not change cell viability and proliferation, indicating that LMX1A is one target of KCNQ1OT1.
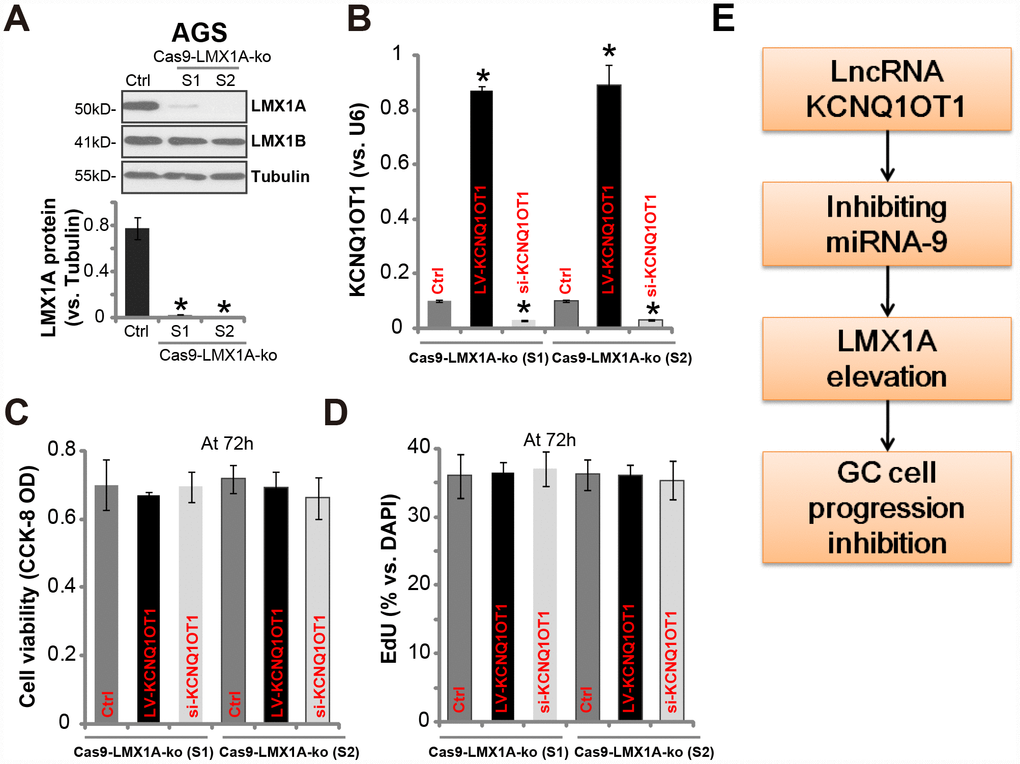
Figure 5. In LMX1A-knockout AGS cells altering KCNQ1OT1 expression fails to change cell viability and proliferation. AGS cells were transfected with the lentiCRISPR/Cas9 LMX1A knockout constructs containing non-overlapping sgRNA sequences (“S1/S2”), following FACS sorting and puromycin selection two stable lines were obtained (“Cas9-LMX1A-ko”). LMX1A and LMX1B protein expression was tested (A, LMX1A protein results were quantified). LV-KCNQ1OT1 or KCNQ1OT1 siRNA (500 nM) were transduced to the Cas9-LMX1A-ko AGS cells (“S1/S2”) for 72h, KCNQ1OT1 expression (B), cell viability (C) and proliferation (D) were shown. (E) The proposed signaling cascade of this study. For each assay, n=5. *P <0.05 vs. “Ctrl” cells. Experiments in this figure were repeated three times, and similar results were obtained.
Discussion
Studies have proposed a possible role of KCNQ1OT1 in carcinogenesis and progression of multiple cancers [21–23]. Recently, a high level of KCNQ1OT1 was detected in lung cancer, promoting proliferation and invasion as well as chemoresistance of lung cancer cells [23, 25]. Similarly, KCNQ1OT1 upregulation and mutation are associated with progression of hepatocellular carcinoma (HCC) and glioma [26, 27]. Feng et al., reported that KCNQ1OT1 upregulation in breast cancer is important for cancer cell growth in vitro and in vivo [21]. Li et al., however demonstrated that KCNQ1OT1 downregulation attenuated myocardial ischemia/reperfusion injury from acute myocardial infarction, possibly by regulating adiponectin receptor 1 (AdipoR1)-p38 /NF-κB signaling cascade [28]. Here, we show that KCNQ1OT1 expression is downregulated in human GC tissues, which correlated with miR-9 upregulation.
Very recent studies have proposed that KCNQ1OT1 can function as a ceRNA. In HCC cells, KCNQ1OT1 sponges miR-504 to regulate the expression of miR-504 target cyclin-dependent kinase 16 (CDK16) [29]. Furthermore, KCNQ1OT1 directly sponges miR-370 in glioma cells [26]. KCNQ1OT1 induces IL-10 expression and macrophage polarization by sponging miR-21a [30]. It promoted caspase-1 activation via inhibiting miR-214 [24]. We here propose that KCNQ1OT1 could act as a ceRNA to sponge miR-9 in GC cells, regulating LMX1A expression and GC cell functions.
In AGS cells and primary human GC cells, forced overexpression of KCNQ1OT1 induced potent miR-9 downregulation but LMX1A upregulation. Contrarily, KCNQ1OT1 silencing, by two non-overlapping siRNAs, induced miR-9 accumulation and LMX1A downregulation. Significantly, GC cell progression, including survival, proliferation, migration and invasion, were inhibited by KCNQ1OT1 overexpression, but enhanced with KCNQ1OT1 silencing. More importantly, in LMX1A knockout AGC cells, KCNQ1OT1 overexpression or silencing was completely ineffective. Thus, we propose that KCNQ1OT1 inhibits GC cell progression via regulating miR-9-LMX1A expression (Figure 5E). Taken together, we conclude that LncRNAKCNQ1OT1 inhibits GC cell progression via regulating microRNA-9-LMX1A expression.
Materials and Methods
Chemicals and reagents
Puromycin, neomycin and polybrene were obtained from Sigma-Aldrich (St. Louis, Mo). All antibodies were purchased from Abcam (Cambridge, MA). RNA reagents, Lipofectamine 2000 and other transcription reagents were obtained from Invitrogen (Shanghai, China). All sequences and plasmids were provided by Genechem (Shanghai, China).
Cell culture
AGS cells were cultured as previously described [7]. The primary human GC cells, derived from two independent written informed-consent GC patients (“GC-1” and “GC-2”), were cultured in the described medium [31]. The protocols were approved by the Ethics Board of Minhang Hospital, Fudan University (Ethics number: 2015-052, Principle Investigator, Dr. Feng Li), in according to Declaration of Helsinki.
Human tissues
As described previously [7], fresh GC cancer tissues along with matched surrounding normal gastric epithelial tissues from twelve (12) primary informed-consent GC patients (all male, 41–67 years old, with no prior chemotherapy and radiotherapy) were acquired. Tissues were washed, minced, and homogenized before further analysis.
qPCR assay
For each treatment, aliquots of 100 ng total RNAs were reverse transcribed into cDNA with Reverse Transcriptase M-MLV (Promega, Madison, WI). The quantitative reverse transcriptase PCR (qPCR) assay was performed by the previously-described protocol [7], using ΔΔCt method for target mRNA quantification and GAPDH as the internal control. KCNQ1OT1 and miR-9 expression was normalized to U6 RNA. Primers for miR-9, LMX1A, GAPDH and U6 were reported previously [7]. The primers for KCNQ1OT1, forward, 5′-CTTTGCAGCAACCTCCTTGT; reverse, 5′-TGGGGTGAGGGATCTGAA, were also reported early [32].
Forced KCNQ1OT1 overexpression
The PCR primer sets, 5′-GGGGTACCCCAGGTGACAAGGTGCAGGCGC and reverse primer: 5′-ACAGAGTTCCTCGTTGGGAGCTTGAAGATCTTC [32], were applied to amplify the upstream KCNQ1OT1 promoter region from GC cells, inserted into the hU6-MCS-Ubiquitin-EGFP-IRES-puromycin vector (GV248, Genechem). The construct, along with the lentiviral packaging constructs [7], were transfected to HEK-293 cells to generate KCNQ1OT1 expression lentivirus (“LV-KCNQ1OT1”). The lentivirus was filtered, enriched and added to GC cells. When necessary, stable cells were selected by puromycin (5.0 μg/mL) for a total of 4-5 passages. Control cells were infected with control lentivirus with empty vector.
KCNQ1OT1 siRNA
GC cells were seeded at 50% confluence. Two different siRNAs (designed and verified by RIBOBIO, Guangzhou, China [24]) against non-overlapping sequence of KCNQ1OT1 were individually transfected (siRNA concentration at 500 nM) by Lipofectamine 2000 reagent for 24h, repeated for a second round for another 24h (total 48h). KCNQ1OT1 knockdown was confirmed by qPCR assay. Control cells were transfected with the scramble non-sense control siRNA (“si-C”).
Luciferase reporter assay
As reported [7], the human LMX1A 3’-UTR with the putative binding sites of miR-9 (see the sites at [7]) was amplified by PCR and then inserted into the firefly luciferase reporter vector, pGL4.13 (luc2/SV40) (Promega), at the XbaI site and downstream from the stop codon of the luciferase gene. AGS cells were plated at 1 × 105 cells per well into six-well plates (60% confluence), transfected with pGL4.13 construct along with the Renilla luciferase reporter vector and pRL-SV40 (Promega). Following the indicated genetic treatments, LMX1A 3’-UTR luciferase activity value was tested, and results were normalized to that in the control cells.
Cell viability assay
Cells were seeded onto the 96-well plates (3 × 103 cells/ well) [7]. Cell viability was determined via the Cell Counting kit-8 (CCK-8, Dojindo Laboratories, Kumamoto, Japan), with optical density (OD) values measured at 570 nm.
In vitro cell migration and invasion assays
As described [33], GC cells were seeded on “Transwell” upper chamber (at 3 × 4000 cells per chamber, BD Biosciences). The complete medium (with 10% FBS) was added to the lower compartments. After 24h the migrated cells on the lower surface were stained. Matrigel (Sigma) was added in the chamber surface when analyzing cell invasion. Five repeated views in each condition were included to calculate the average number of migrated/invasive cells.
EdU assay
EdU (5-ethynyl-20-deoxyuridine) Apollo-567 Kit (RIBOBIO, Shanghai, China) was employed to test cell proliferation. Briefly, cells were seeded onto the twelve-well plates. Following treatment, EdU and DAPI dyes were added, and cells were incubated for additional 8h. Cell nuclei were then visualized under a fluorescent microscope. For each condition at least 500 cells in five random views were included to calculate EdU ratio (EdU/DAPI×100%).
Apoptosis assays
Testing cell apoptosis, by the Annexin V FACS and TUNEL staining assay, was described in detail in our previous study [7].
LMX1A knockout
The two different lentiCRISPR-GFP-puro LMX1A knockout constructs (see our previous study [7]) were independently transfected to AGS cells by Lipofectamine 2000. GFP-positive cells were FACS-sorted, cultured for another three weeks. Stable cells were achieved by puromycin selection, named as “Cas9-LMX1A-ko” cells.
Western blotting
Total cellular lysates were resolved by SDS-PAGE (10%) gels, transferred to a PVDF membrane. The latter was incubated with blocking buffer and following primary and secondary antibodies. ECL method was applied to detect the immuno-reactive bands.
Statistical analyses
The GraphPad Prism software was employed for statistical analyses. All values were expressed as the mean ± standard deviation (SD). Repeated-measures analysis of variance (RMANOVA) followed by Dunnett’s post hoc test for multiple comparisons were applied to evaluate statistical significance. All differences were considered significant at P < 0.05. To determine significance between two treatment groups, the two-tailed t tests were carried out.
Author Contributions
All listed authors designed the study, performed the experiments and the statistical analysis, and wrote the manuscript. All authors have read the manuscript and approved the final version.
Conflicts of Interest
The authors declare that they have no conflicts of interests.
Funding
This project was supported by Minhang District University Building Project (2017MWDXK03). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018; 68:7–30. https://doi.org/10.3322/caac.21442 [PubMed]
- 2. Choi KS, Suh M. Screening for gastric cancer: the usefulness of endoscopy. Clin Endosc. 2014; 47:490–96. https://doi.org/10.5946/ce.2014.47.6.490 [PubMed]
- 3. Hashim D, Boffetta P, La Vecchia C, Rota M, Bertuccio P, Malvezzi M, Negri E. The global decrease in cancer mortality: trends and disparities. Ann Oncol. 2016; 27:926–33. https://doi.org/10.1093/annonc/mdw027 [PubMed]
- 4. Shah MA. Gastrointestinal cancer: targeted therapies in gastric cancer-the dawn of a new era. Nat Rev Clin Oncol. 2014; 11:10–11. https://doi.org/10.1038/nrclinonc.2013.231 [PubMed]
- 5. Wadhwa R, Song S, Lee JS, Yao Y, Wei Q, Ajani JA. Gastric cancer-molecular and clinical dimensions. Nat Rev Clin Oncol. 2013; 10:643–55. https://doi.org/10.1038/nrclinonc.2013.170 [PubMed]
- 6. Qian P, Li J, Zhang X, Li F, Bei S, Li H, Sun Q, Feng L. LMX1A inhibits C-Myc expression through ANGPTL4 to exert tumor suppressive role in gastric cancer. PLoS One. 2019; 14:e0221640. https://doi.org/10.1371/journal.pone.0221640 [PubMed]
- 7. Zhang X, Qian Y, Li F, Bei S, Li M, Feng L. microRNA-9 selectively targets LMX1A to promote gastric cancer cell progression. Biochem Biophys Res Commun. 2018; 505:405–12. https://doi.org/10.1016/j.bbrc.2018.09.101 [PubMed]
- 8. Doucet-Beaupré H, Ang SL, Lévesque M. Cell fate determination, neuronal maintenance and disease state: the emerging role of transcription factors Lmx1a and Lmx1b. FEBS Lett. 2015; 589:3727–38. https://doi.org/10.1016/j.febslet.2015.10.020 [PubMed]
- 9. Dong W, Feng L, Xie Y, Zhang H, Wu Y. Hypermethylation-mediated reduction of LMX1A expression in gastric cancer. Cancer Sci. 2011; 102:361–66. https://doi.org/10.1111/j.1349-7006.2010.01804.x [PubMed]
- 10. Chang CC, Huang RL, Wang HC, Liao YP, Yu MH, Lai HC. High methylation rate of LMX1A, NKX6-1, PAX1, PTPRR, SOX1, and ZNF582 genes in cervical adenocarcinoma. Int J Gynecol Cancer. 2014; 24:201–09. https://doi.org/10.1097/IGC.0000000000000054 [PubMed]
- 11. Feng L, Xie Y, Zhao Z, Lian W. LMX1A inhibits metastasis of gastric cancer cells through negative regulation of β-catenin. Cell Biol Toxicol. 2016; 32:133–39. https://doi.org/10.1007/s10565-016-9326-0 [PubMed]
- 12. Niu ZS, Niu XJ, Wang WH. Long non-coding RNAs in hepatocellular carcinoma: potential roles and clinical implications. World J Gastroenterol. 2017; 23:5860–74. https://doi.org/10.3748/wjg.v23.i32.5860 [PubMed]
- 13. Huo X, Han S, Wu G, Latchoumanin O, Zhou G, Hebbard L, George J, Qiao L. Dysregulated long noncoding RNAs (lncRNAs) in hepatocellular carcinoma: implications for tumorigenesis, disease progression, and liver cancer stem cells. Mol Cancer. 2017; 16:165. https://doi.org/10.1186/s12943-017-0734-4 [PubMed]
- 14. Yang X, Xie X, Xiao YF, Xie R, Hu CJ, Tang B, Li BS, Yang SM. The emergence of long non-coding RNAs in the tumorigenesis of hepatocellular carcinoma. Cancer Lett. 2015; 360:119–24. https://doi.org/10.1016/j.canlet.2015.02.035 [PubMed]
- 15. Hao NB, He YF, Li XQ, Wang K, Wang RL. The role of miRNA and lncRNA in gastric cancer. Oncotarget. 2017; 8:81572–82. https://doi.org/10.18632/oncotarget.19197 [PubMed]
- 16. Sun M, Nie FQ, Wang ZX, De W. Involvement of lncRNA dysregulation in gastric cancer. Histol Histopathol. 2016; 31:33–39. https://doi.org/10.14670/HH-11-655 [PubMed]
- 17. Chen L, Dzakah EE, Shan G. Targetable long non-coding RNAs in cancer treatments. Cancer Lett. 2018; 418:119–24. https://doi.org/10.1016/j.canlet.2018.01.042 [PubMed]
- 18. Arun G, Diermeier SD, Spector DL. Therapeutic Targeting of Long Non-Coding RNAs in Cancer. Trends Mol Med. 2018; 24:257–77. https://doi.org/10.1016/j.molmed.2018.01.001 [PubMed]
- 19. Mitsuya K, Meguro M, Lee MP, Katoh M, Schulz TC, Kugoh H, Yoshida MA, Niikawa N, Feinberg AP, Oshimura M. LIT1, an imprinted antisense RNA in the human KvLQT1 locus identified by screening for differentially expressed transcripts using monochromosomal hybrids. Hum Mol Genet. 1999; 8:1209–17. https://doi.org/10.1093/hmg/8.7.1209 [PubMed]
- 20. Lee MP, DeBaun MR, Mitsuya K, Galonek HL, Brandenburg S, Oshimura M, Feinberg AP. Loss of imprinting of a paternally expressed transcript, with antisense orientation to KVLQT1, occurs frequently in Beckwith-Wiedemann syndrome and is independent of insulin-like growth factor II imprinting. Proc Natl Acad Sci USA. 1999; 96:5203–08. https://doi.org/10.1073/pnas.96.9.5203 [PubMed]
- 21. Feng W, Wang C, Liang C, Yang H, Chen D, Yu X, Zhao W, Geng D, Li S, Chen Z, Sun M. The Dysregulated Expression of KCNQ1OT1 and Its Interaction with Downstream Factors miR-145/CCNE2 in Breast Cancer Cells. Cell Physiol Biochem. 2018; 49:432–46. https://doi.org/10.1159/000492978 [PubMed]
- 22. Zhang S, Ma H, Zhang D, Xie S, Wang W, Li Q, Lin Z, Wang Y. LncRNA KCNQ1OT1 regulates proliferation and cisplatin resistance in tongue cancer via miR-211-5p mediated Ezrin/Fak/Src signaling. Cell Death Dis. 2018; 9:742. https://doi.org/10.1038/s41419-018-0793-5 [PubMed]
- 23. Dong Z, Yang P, Qiu X, Liang S, Guan B, Yang H, Li F, Sun L, Liu H, Zou G, Zhao K. KCNQ1OT1 facilitates progression of non-small-cell lung carcinoma via modulating miRNA-27b-3p/HSP90AA1 axis. J Cell Physiol. 2019; 234:11304–11314. https://doi.org/10.1002/jcp.27788 [PubMed]
- 24. Jin X, Jin H, Shi Y, Guo Y, Zhang H. Long Non-Coding RNA KCNQ1OT1 Promotes Cataractogenesis via miR-214 and Activation of the Caspase-1 Pathway. Cell Physiol Biochem. 2017; 42:295–305. https://doi.org/10.1159/000477330 [PubMed]
- 25. Ren K, Xu R, Huang J, Zhao J, Shi W. Knockdown of long non-coding RNA KCNQ1OT1 depressed chemoresistance to paclitaxel in lung adenocarcinoma. Cancer Chemother Pharmacol. 2017; 80:243–50. https://doi.org/10.1007/s00280-017-3356-z [PubMed]
- 26. Gong W, Zheng J, Liu X, Liu Y, Guo J, Gao Y, Tao W, Chen J, Li Z, Ma J, Xue Y. Knockdown of Long Non-Coding RNA KCNQ1OT1 Restrained Glioma Cells’ Malignancy by Activating miR-370/CCNE2 Axis. Front Cell Neurosci. 2017; 11:84. https://doi.org/10.3389/fncel.2017.00084 [PubMed]
- 27. Wan J, Huang M, Zhao H, Wang C, Zhao X, Jiang X, Bian S, He Y, Gao Y. A novel tetranucleotide repeat polymorphism within KCNQ1OT1 confers risk for hepatocellular carcinoma. DNA Cell Biol. 2013; 32:628–34. https://doi.org/10.1089/dna.2013.2118 [PubMed]
- 28. Li X, Dai Y, Yan S, Shi Y, Han B, Li J, Cha L, Mu J. Down-regulation of lncRNA KCNQ1OT1 protects against myocardial ischemia/reperfusion injury following acute myocardial infarction. Biochem Biophys Res Commun. 2017; 491:1026–33. https://doi.org/10.1016/j.bbrc.2017.08.005 [PubMed]
- 29. Li C, Miao R, Zhang J, Qu K, Liu C. Long non-coding RNA KCNQ1OT1 mediates the growth of hepatocellular carcinoma by functioning as a competing endogenous RNA of miR-504. Int J Oncol. 2018. [Epub ahead of print]. https://doi.org/10.3892/ijo.2018.4313 [PubMed]
- 30. Gao X, Ge J, Li W, Zhou
W , Xu L. LncRNA KCNQ1OT1 ameliorates particle-induced osteolysis through inducing macrophage polarization by inhibiting miR-21a-5p. Biol Chem. 2018; 399:375–86. https://doi.org/10.1515/hsz-2017-0215 [PubMed] - 31. Yang L, Zheng LY, Tian Y, Zhang ZQ, Dong WL, Wang XF, Zhang XY, Cao C. C6 ceramide dramatically enhances docetaxel-induced growth inhibition and apoptosis in cultured breast cancer cells: a mechanism study. Exp Cell Res. 2015; 332:47–59. https://doi.org/10.1016/j.yexcr.2014.12.017 [PubMed]
- 32. Sunamura N, Ohira T, Kataoka M, Inaoka D, Tanabe H, Nakayama Y, Oshimura M, Kugoh H. Regulation of functional KCNQ1OT1 lncRNA by β-catenin. Sci Rep. 2016; 6:20690. https://doi.org/10.1038/srep20690 [PubMed]
- 33. Wang SS, Lv Y, Xu XC, Zuo Y, Song Y, Wu GP, Lu PH, Zhang ZQ, Chen MB. Triptonide inhibits human nasopharyngeal carcinoma cell growth via disrupting Lnc-RNA THOR-IGF2BP1 signaling. Cancer Lett. 2019; 443:13–24. https://doi.org/10.1016/j.canlet.2018.11.028 [PubMed]



