Introduction
A recent study suggested PIK3CA mutations to be a major mediator of therapy resistance in breast cancer [1]. More than 70% of hormone receptor (HR)-positive breast cancers have molecular aberrations in Phosphatidylinositol 3-kinase (PI3K)-AKT-mTOR pathways [2]. PI3K is a heterodimer composed of a regulatory subunit p85 and a catalytic subunit p110 [3]. The PIK3CA gene encodes the PI3K catalytic subunit p110α [4]. According to the circulating tumor DNA (ctDNA) sequencing results, about 50% of HR-positive metastatic breast cancers (MBCs) have PIK3CA missense mutations; 10-30% of metastatic triple-negative breast cancers (TNBC) and HER2-positive breast cancers have PIK3CA missense mutations [5]. About 80% of PIK3CA mutations occur in helix domain (HD) exon 9 and kinase domain (KD) exon 20 [6]. 26% of PIK3CA mutations are in exon 9 (hotspots: E545K and E542K), and 50% of PIK3CA mutations are in exon 20 (hotspot: H1047R) [7]. While PIK3CA-HD mutations are less potent in inducing mammary tumors [8], these mutations are independently associated with poor prognosis (early recurrence and death) [9]. Although PIK3CA-KD mutations are associated with lymph node infiltration and are much more aggressive in carcinogenesis compared to PIK3CA-HD mutations [8, 10–11], PIK3CA-KD mutations are associated with optimal prognosis [9].
PIK3CA-HD mutations are more frequent in older age-onset patients, and they are not associated with lymph node infiltration as PIK3CA-KD mutations [10–11]. Importantly, the therapeutic response of PIK3CA-HD mutant tumors is significantly poorer than tumors with PIK3CA-KD mutations [12]. Mechanism studies suggest that, instead of binding to the p85 subunit, PIK3CA-HD mutation-encoded protein PIK3CA(E545K) interacts with Ras-GTP [13]. Compared to PIK3CA(H1047R), PIK3CA(E545K) is less efficient in activating the downstream Akt signaling [13].
Most MBC patients receive chemotherapy coupled with/without endocrine therapy or anti-HER2 therapy. The activation of PI3K/AKT/mTOR pathway, however, would promote tumor progression and induce drug resistance to endocrine therapy and chemotherapy [14–16]. In this study, we examined the genetic aberrations of PI3K/AKT pathway and assessed the effect of these aberrations on progression-free survival (PFS) in metastatic breast cancer (MBC) patients with late-line treatment. We analyzed the ctDNA mutation profile and investigated the association of PI3K/AKT pathway aberrations with the clinical and genetic features of MBC tumors. We compared distinct types of PI3K/AKT pathway mutations, especially PIK3CA-HD and PIK3CA-KD mutations, to determine the different effects of PI3K/AKT pathway mutations on the prognosis of MBC patients with late-line treatment.
Results
Genetic aberrations of PI3K/AKT pathway molecules in MBCs after early-line salvage therapy
A total of 193 MBC patients after 1st or 2nd salvage treatment received ctDNA testing and then late-line therapy between December 2016 and December 2018. Ninety three patients (48%) had ctDNA mutations of the PI3K/AKT pathway molecules. Among these ninety three patients, thirty six patients (38.7%) had PIK3CA–KD missense mutations, including thirty one p.H1047R, one p.H1047L, two p.G1049R, and one p.M1043V missense mutations (Figure 1A). Twenty six patients (28%) had PIK3CA–HD missense mutations, including sixteen p.H545K, six p.H542K, one p.E545G, two p.Q546R and one p.Q546H missense mutations (Figure 1B). Ten patients (10.8%) had PIK3CA-OD aberrations, including five p.N345K, two p.C420R and one p.Y985S missense mutation, one p.L113del and one p.G106_E109del mutation (Figure 1C). The other twenty one patients (22.6%) had aberrations in other PI3K/AKT pathway molecules except PIK3CA, including six AKT1 p.E17K missense mutations, two other AKT1 mutations, two AKT2 mutations, two PTEN frameshifts, two PIK3CB, three PIK3CG, two PIK3R1 and two PIK3R2 mutations (Figure 1D).
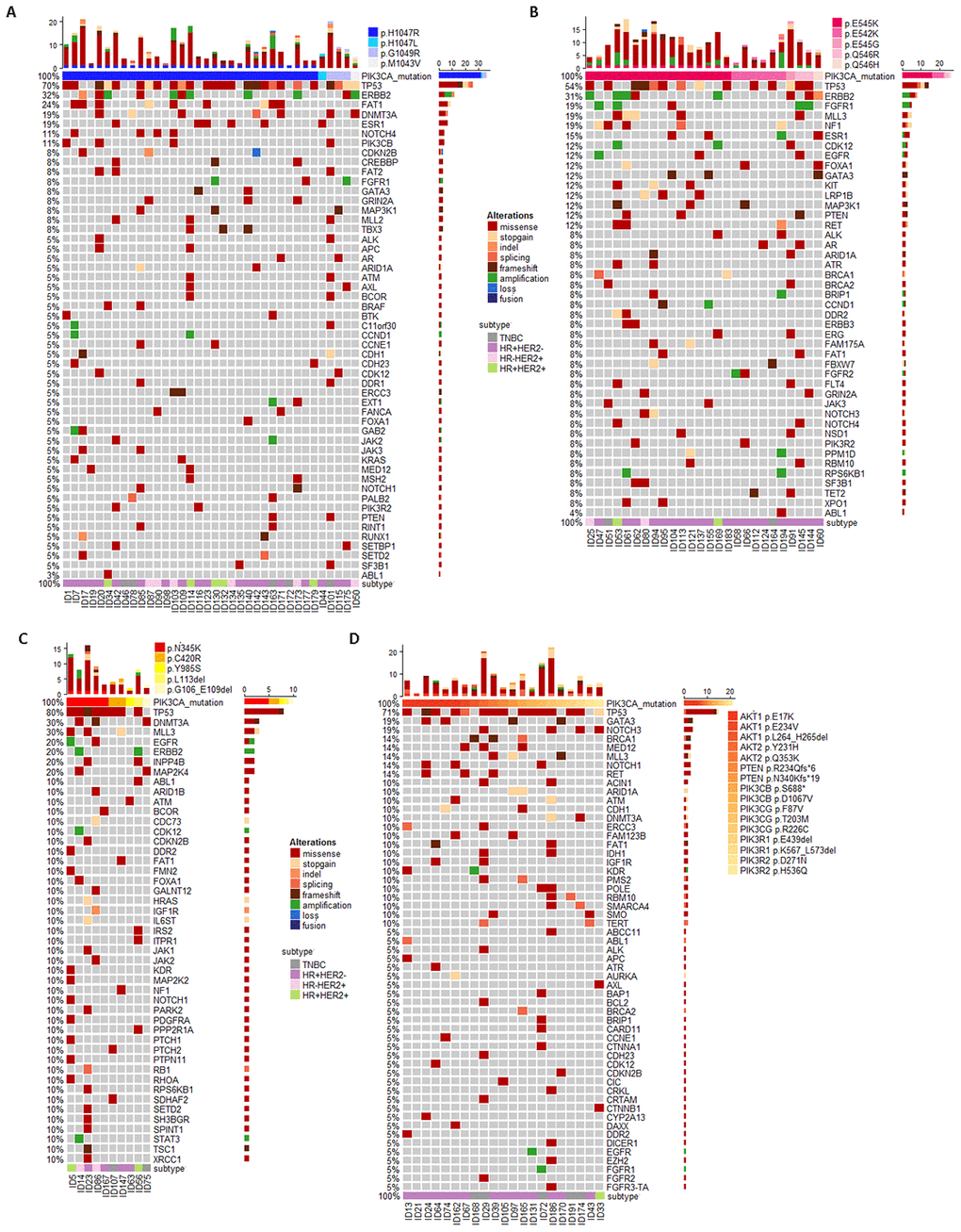
Figure 1. Circulating tumor DNA (ctDNA) gene mutation profiles for MBC patients who progressed after early-line therapy and had PIK3CA mutations in kinase domain (PIK3CA–KD, A) helix domain (PIK3CA–HD, B), other region (PIK3CA–OD, C) and other PI3K/AKT pathway aberrations (P/A, D).
Clinical features were distinct with respect to different PI3K/AKT aberrations (Table 1). Compared to the wild-type (WT) patients, patients with PI3K/AKT pathway aberrations had longer time from BC diagnosis to metastasis (TTM) (p = 0.001). More PI3K/AKT pathway aberrant patients had visceral metastases, and more PI3K/AKT pathway aberrant patients were ER-positive or PR-positive. In addition, compared to PIK3CA-KD mutant patients, PIK3CA-HD carriers had an even higher ER/PR-positive rate.
Table 1. Clinical characteristics of MBC patients with wild-type PIK3CA and PI3K/AKT pathways aberrations.
| Variables | Levels | WT (n=100) | PI3K/AKT pathway aberrations | p-value*† | p-value**† | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PIK3CA-KD (n=36) | PIK3CA-HD (n=26) | PIK3CA-OD (n=10) | P/A (n=21) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age at diagnosis (years) | 43.7 ± 8.9, 42.9 (36.3, 49.8) | 45.0 ± 9.5, 42.9 (37.9, 51.6) | 45.9 ± 9.9, 44.6 (38.1, 51.0) | 45.0 ± 11.4, 47.0 (37.6, 55.3) | 41.7 ± 11.4, 40.0 (35.1, 52.0) | 0.55 | 0.70 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age at metastasis (years) # | 46.4 ± 9.2, 46.2 (38.8, 52.3) | 49.3 ± 9.3, 50.1 (41.9, 56.4) | 51.2 ± 9.8, 52.5 (40.3, 56.4) | 48.7 ± 12.1, 54.0 (38.0, 57.3) | 44.4 ± 10.8, 42.2 (37.2, 52.3) | 0.09 | 0.45 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TTM ## | 2.6 ± 2.8, 1.6 (0.3, 4.3) | 4.4 ± 4.3, 2.8 (1.5, 5.4) | 5.1 ± 3.6, 5.0 (3.4, 6.4) | 3.7 ± 4.7, 2.3 (0.4, 3.0) | 3.3 ± 2.9, 2.2 (0.6, 5.9) | 0.001 | 0.53 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ER | Positive | 40 (40.0%) | 22 (61.1%) | 21 (80.8%) | 5 (50.0%) | 16 (76.2%) | 0.0003 | 0.10 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Negative | 60 (60.0%) | 14 (38.9%) | 5 (19.2%) | 5 (50.0%) | 5 (23.8%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PR | Positive | 33 (33.0%) | 19 (52.8%) | 19 (73.1%) | 6 (60.0%) | 13 (61.9%) | <0.0001 | 0.10 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Negative | 67 (67.0%) | 17 (47.2%) | 7 (26.9%) | 4 (40.0%) | 8 (38.1%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HER2 | Positive | 30 (30.0%) | 11 (30.6%) | 4 (15.4%) | 4 (40.0%) | 1 (4.8%) | 0.21 | 0.23 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Negative | 70 (70.0%) | 25 (69.4%) | 22 (84.6%) | 6 (60.0%) | 20 (95.2%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HR/HER2 subtype | TNBC | 36 (36.0%) | 5 (13.9%) | 2 (7.7%) | 2 (20.0%) | 5 (23.8%) | 0.27 | 0.46 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HR+/HER2- | 34 (34.0%) | 20 (55.6%) | 20 (76.9%) | 4 (40.0%) | 15 (71.4%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HR-/HER2+ | 21 (21.0%) | 6 (16.7%) | 2 (7.7%) | 2 (20.0%) | 0 (0%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HR+/HER2+ | 9 (9.0%) | 5 (13.9%) | 2 (7.7%) | 2 (20.0%) | 1 (4.8%) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metastasis sites ### | Bone-only | 16 (16.0%) | 2 (5.6%) | 4 (15.4%) | 2 (20.0%) | 4 (19.1%) | 0.54 | 0.22 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Visceral | 47 (47.0%) | 28 (77.8%) | 18 (69.2%) | 5 (50.0%) | 14 (66.7%) | 0.001 | 0.45 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Soft tissue | 53 (53.0%) | 26 (72.2%) | 16 (61.5%) | 3 (30.0%) | 12 (57.1%) | 0.25 | 0.37 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Note: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Patients with PI3K/AKT pathway aberrations were divided into four groups: WT (wild-type group), PIK3CA-KD (PIK3CA kinase-domain mutation) group, PIK3CA-HD (PIK3CA helix-domain mutation) group, PIK3CA-OD (other PIK3CA mutation) group, and P/A (other PI3K/AKT pathway mutations) group. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| p-values† were calculated using Student’s t-tests for continuous variables and using Chi-square tests (Mentel-Haenszel for >2 levels comparison), or Fisher’s exact tests (n<5) for categorical variables | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| p-values* compared variables between wild-type patients and PI3K/AKT aberrant patients; p-value** compared variables between PIK3CA-KD mutant patients and PIK3CA-HD mutant patients. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age at metastasis # represented the age (in year) of patients when the metastasis occurred. TTM ## represented the time (in year) from diagnosis to metastasis. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metastatic sites ### compared the patients with bone-only metastases, visceral metastases (such as liver, lung, brain, ovary, etc), and soft tissue metastases (lymphnode, mediastinum, plura and contra-lateral breast). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviation: ER (Estrogen Receptor), PR (Progesterone Receptor), HER2 (Human Epidermal Growth Factor Receptor-2). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Poor prognosis with PIK3CA mutations in helical domain
All included patients failed and progressed in the early–line (≤2) salvage therapy. Then, the later line regimen was given by TPC (treatment physician choice). The median PFS for patients with WT, PIK3CA-KD, PIK3CA-HD, PIK3CA-OD, other PI3K/AKT (P/A) mutations were 7.7 months (95% CI: 5.4-9.6), 5.1 months (95% CI: 3.5-13.0), 3.2 months (95% CI: 2.1-4.2), 11.2 months (95% CI: 3.0-NE), and 4.6 months (95% CI: 3.3-14.8), respectively (Figure 2). Apart from PIK3CA-HD mutations, patients with PI3K/AKT pathway aberrations had similar PFS to WT patients (Logrank p=0.63). PIK3CA-HD mutant patients had a significantly lower PFS than WT patients (Logrank p<0.0001), PIK3CA-KD mutant patients (Logrank p=0.02), PIK3CA-OD mutant patients (Logrank p=0.0006) and P/A aberrant patients (Logrank p=0.01).
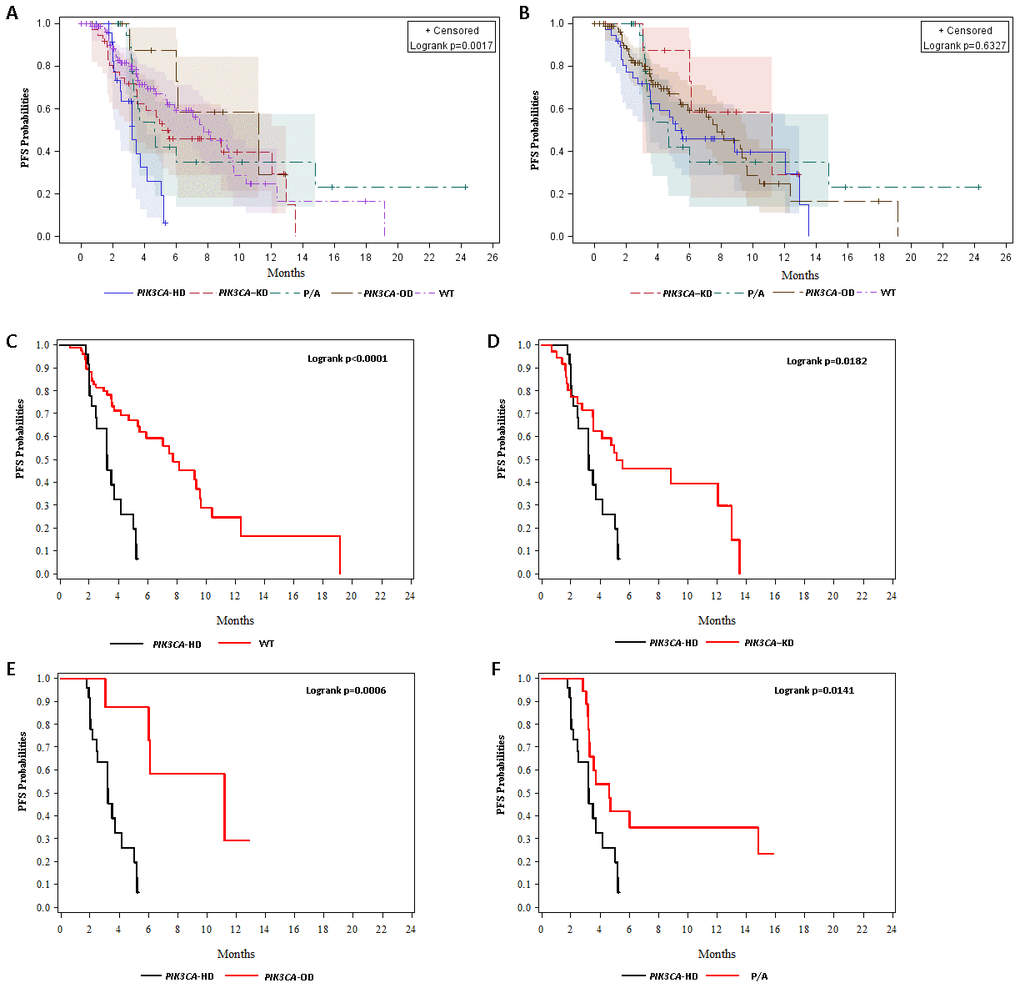
Figure 2. Kaplan-Meier (KM) curves for progression-free survival (PFS) probabilities. (A) KM curves for PFS probabilities stratified by wild-type (WT) and PIK3CA–KD mutations, PIK3CA–HD mutations, PIK3CA–OD mutations, and other PI3K/AKT pathway aberrations (P/A). (B) KM curves for PFS probabilities stratified by wild-type (WT) and PIK3CA–KD mutations, PIK3CA–OD mutations, and other PI3K/AKT pathway aberrations (P/A). (C) KM curves for PFS probabilities stratified stratified by wild-type (WT) and PIK3CA–HD mutations. (D) KM curves for PFS probabilities stratified stratified by PIK3CA–HD mutations and PIK3CA–KD mutations. (E) KM curves for PFS probabilities stratified by PIK3CA–HD mutations and PIK3CA–OD mutations. (F) KM curves for PFS probabilities stratified by PIK3CA–HD mutations and other PI3K/AKT pathway aberrations (P/A).
ctDNA mutation profile
Due to the metastatic tumor burden, we could detect ctDNA mutations in most MBC patients [5]. Compared to WT patients, patients with PI3K/AKT pathway aberrations had significantly higher rates of TP53, ERBB2, FAT1, and FGFR aberrations (Table 2). In addition, patients with PIK3CA-HD mutations had an obviously distinct ctDNA mutation profile (Figure 1). For example, compared to the PIK3CA-KD mutations, PIK3CA-HD mutant patients had a higher FGFR aberration rate (Fisher’s exact test, p=0.10), a higher NF1 mutation rate (Fisher’s exact test, p=0.07), and a lower DNMT3A mutation rate (Fisher’s exact test, p=0.04) (Table 2).
Table 2. Somatic mutations accompanied with PI3K/AKT pathway gene aberrations.
| Variables | WT (n=100) | PI3K/AKT pathway mutations# | p-value*† | p-value**† | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PIK3CA-KD (n=36) | PIK3CA-HD (n=26) | PIK3CA-OD (n=10) | P/A (n=21) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TP53 | 33 (32.0%) | 26 (72.2%) | 14 (53.9%) | 9 (90.0%) | 14 (66.7%) | <0.0001 | 0.14 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ERBB2 | 8 (8.0%) | 10 (27.8%) | 8 (30.8%) | 2 (20.0%) | 0 (0%) | 0.008 | 0.80 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FAT1 | 3 (3.0%) | 9 (25.0%) | 2 (7.7%) | 1 (10.0%) | 2 (9.5%) | 0.004 | 0.10 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ESR1 | 5 (5.0%) | 6 (16.7%) | 4 (15.4%) | 0 (0%) | 0 (0%) | 0.18 | 0.90 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DNMT3A | 6 (6.0%) | 7 (19.4%) | 0 (0%) | 2 (20.0%) | 2 (9.5%) | 0.15 | 0.04 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FGFR | 7 (7.0%) | 4 (11.1%) | 8 (36.4%) | 0 (0%) | 3 (14.3%) | 0.05 | 0.10 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NF1 | 4 (4.0%) | 1 (2.8%) | 5 (19.2%) | 1 (10.0%) | 0 (0%) | 0.36 | 0.07 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TMB-High | 5 (5.0%) | 8 (22.2%) | 14 (53.9%) | 3 (30.0%) | 4 (19.1%) | <0.0001 | 0.01 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| #Patients with PI3K/AKT pathway aberrations were divided into four groups: WT group, PIK3CA-KD group, PIK3CA-HD group, PIK3CA-OD group, and P/A (other PI3K/AKT pathway mutations) group. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| p-values† were calculated using Student’s t-tests for continuous variables and using Chi-square tests (Mentel-Haenszel for >2 levels comparison), or Fisher’s exact tests (n<5) for categorical variables. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| p-values* compared variables between WT patients and PI3K/AKT pathway aberrant patients; p-value** compared variables between PIK3CA-KD mutant patients and PIK3CA-HD mutant patients. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
High TMB in helical domain mutant patients
Tumor mutation burden (TMB) has been considered as a marker for immunotherapy. The more mutations the tumor has, the higher possibility of production and subsequent presentation of tumor-associated antigens (TAA) on MHC molecules, which leads to a higher tumor cell cytotoxicity after the inhibition of immune checkpoint signals [17, 18]. BCs are “cold” tumors with less TAAs than “hot” tumors (non-small cell lung cancer and malignant melanoma) [19, 20]. In this study, we defined TMB-H if the TMB was larger than the top 25% TMB value of all BC samples in Geneplus database (9 Muts/Mb). This cut-off value of 9 Muts/Mb was reasonable because it was equal to the lung cancer of 9 Muts/Mb (the third third tertile) [21] and close to the gastric cancer of 12 Muts/Mb (top 20%) [22]. Here, we found that PIK3CA-HD mutant MBC patients had a significantly higher TMB level than patients with PIK3CA-KD mutations (p=0.006), PIK3CA-OD mutations (p=0.045), or other P/A pathway mutations (p=0.01) (Figure 3).
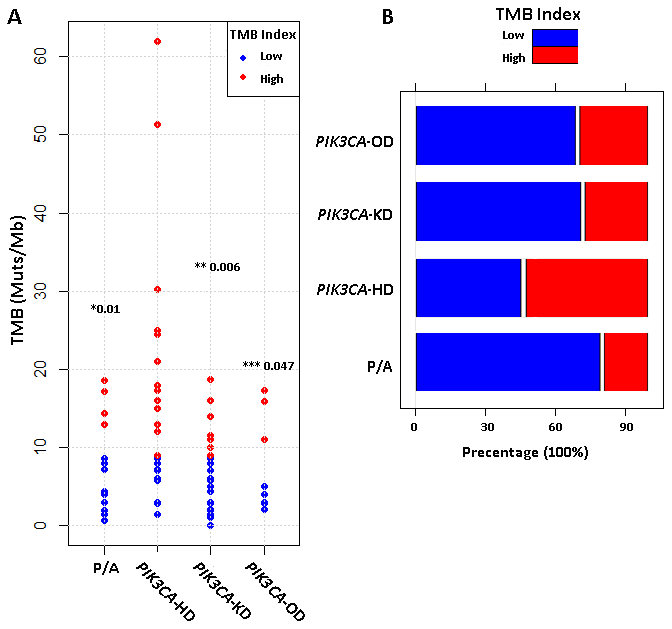
Figure 3. Tumor mutation burden (TMB) for different metastatic breast cancer subtypes. (A) Comparison of TMB levels (mutations per million bases, muts/Mb) among four types of PI3K/AKT pathway aberrations. p-value for PIK3CA-HD vs PIK3CA-KD (*), PIK3CA-HD vs PIK3CA-OD (**) and PIK3CA-HD vs P/A (***) were calculated by using non-parametric wilcoxon rank-sum test. (B) Barplot compared the percentages of TMB-High (red) and TMB-Low (blue) patients among four different types of PI3K/AKT pathway aberrations.
Risk prognostic factors for MBC patients with late-line therapy
Cox regression analyses suggested that none of the clinical indicators had significant impacts on PFS for late-line therapies, including the age of diagnosis, age of BC metastasis, TTM, primary BC laterality, ER, PR, HER2 status and the sites of metastatic lesions (Supplementary Table 1). However, we found that some genetic indicators, including PIK3CA-HD mutations, FGFR aberrations, and high TMB levels, were significant risk factors for poor PFS for late-line therapy in MBCs (Table 3). Multivariate Cox regression analyses showed that compared to PI3K/AKT pathway WT MBCs, the hazard ratios (HR) for patients with PIK3CA-HD mutations, TP53 mutations, FGFR aberrations, and high TMB levels were 2.0 (95% CI = 1.02-3.93, p=0.045), 2.22 (95% CI = 1.35-3.66, p=0.0002), 2.17 (95% CI = 1.17-4.02, p=0.01), and 1.62 (95% CI = 1.00-2.66, p=0.05), respectively.
Table 3. Cox regression analyses of genetic risk factors for PFS in MBC patients.
| Covariates | Level | Univariate Cox Model | Multivariate Cox Model | |||
| HR (95% CI) | p-value | HR (95% CI) | p-value | |||
| PI3K/AKT pathway | WT | Ref | Ref | |||
| mutations | PIK3CA-HD | 2.99 (1.64, 5.44) | 0.0004 | 2.00 (1.02, 3.93) | 0.045 | |
| PIK3CA-KD | 1.22 (0.72, 2.09) | 0.45 | 0.75 (0.39, 1.42) | 0.37 | ||
| PIK3CA-OD | 0.64 (0.23, 1.80) | 0.39 | 0.40 (0.14, 1.18) | 0.10 | ||
| P/A | 0.95 (0.49, 1.85) | 0.88 | 0.80 (0.40, 1.57) | 0.51 | ||
| TP53 aberration | No | Ref | Ref | |||
| Yes | 2.25 (1.43, 3.54) | 0.0004 | 2.22 (1.35, 3.66) | 0.002 | ||
| ERBB2 aberration | No | Ref | Ref | |||
| Yes | 1.54 (0.93, 2.55) | 0.09 | 1.45 (0.80, 2.65) | 0.23 | ||
| FAT1 aberration | No | Ref | Ref | |||
| Yes | 2.06 (1.06, 4.00) | 0.03 | 1.48 (0.74, 2.98) | 0.27 | ||
| ESR1 aberration | No | Ref | Ref | |||
| Yes | 1.16 (0.56, 2.43) | 0.69 | 1.25 (0.58, 2.71) | 0.60 | ||
| DNMT3A aberration | No | Ref | Ref | |||
| Yes | 1.05 (0.54, 2.04) | 0.88 | 0.93(0.45, 1.91) | 0.84 | ||
| FGFR aberration | No | Ref | Ref | |||
| Yes | 1.87 (1.05, 3.33) | 0.03 | 2.17 (1.17, 4.02) | 0.01 | ||
| NF1 aberration | No | Ref | Ref | |||
| Yes | 1.03 (0.38, 2.81) | 0.96 | 0.6 (0.30, 1.56) | 0.37 | ||
| TMB index | Low | Ref | Ref | |||
| High | 1.94 (1.19, 3.16) | 0.008 | 1.62 (1.00, 2.66) | 0.05 | ||
| Unknown | 0.73 (0.36, 1.47) | 0.38 | 0.88 (0.43, 1.79) | 0.72 | ||
KM curves after PMS
Since ER/PR status, TP53 mutations, FGFR aberrations and TMB levels were imbalanced between PIK3CA-HD mutant and PIK3CA-KD mutant MBC patients, we performed a 1:1 propensity score matching (PSM) strategy to avoid the potential bias. Kaplan-Merier (KM) curves after PSM still showed a marginally significant difference between PIK3CA-HD mutant and PIK3CA-KD mutant MBCs (Supplementary Figure 1, Logrank p=0.13), suggesting PIK3CA-HD mutation itself to be an important risk factor for poor prognosis for MBCs in late-line therapy.
Discussion
For progressed MBCs after the early-line (≤ 2-line) therapies, there is no consensus on the late-line therapy. Potential choices for hormone receptor (HR)-positive MBCs include mTOR inhibitor, FGFR inhibitor, Estradiol and Progestin [23, 24]. Candidates for metastatic TNBCs might be PD1/PD-L1 inhibitors, anti-VEGFR, etc [25, 26]. Novel TKIs and PD-L1 inhibitors might be useful in HER2-positive patients [27, 28]. In this study, we found that the conception of HR/HER2 status was gradually obscured in late-line therapy. Instead, the genetic aberrations and immune checkpoints became more and more important.
After early-line therapies, MBC tumors become more heterogenous and have more somatic gene aberrations. PIK3CA, ESR1 and GATA3 mutations increased in progressed patients after chemotherapy [15]. The rate of ESR1 mutations increased after AI treatment [29]. After the treatment of CDK4/6 inhibitors, RB1 mutation could be detected by ctDNA testing [30]. Based on these observations, we hypothesize that after multi-line therapies, the genotype of MBC tumors will change significantly, which may lead to the resistance of MBC tumors to the standard therapies that were designed based on HR/HER2 status. In addition, MBC tumors might become “warmer” after multi-line therapies, because the new mutations might generate and present many novel TAAs. Therefore, in terms of late-line therapy, we need to consider more about tumor genotype and immune checkpoints.
In recent years, the usage of antibodies to block the immune checkpoint PD-1 / PD-L1 has become a promising treatment strategy for cancer patients. However, many patients have failed to respond to PD-1 / PD-L1 treatment. Therefore, plenty of researches have focused on the biomarkers to distinguish the responders and non-responders for PD-1/PD-L1 antibody treatment [31]. IMpassion 130 study showed that the low positivity rate of 1% PD-L1 expression rate could be sufficient for patients to have a better response to PD-L1 antibody in triple-negative breast cancers (TNBC) [32]. In addition, a higher non-synonymous mutation or candidate neoantigen burden in tumors (TMB) improved the treatment response of PD-1 antibody in lung cancers [19]. In this study, PIK3CA-HD mutations were concentrated in HR-positive patients, and more than half of these patients were TMB-H (Figure 3). Thus, for this group of patients, PD-1 antibody might be a reasonable choice.
In this study, we found that in MBC patients with PI3K/AKT pathway aberration, TP53, ERBB2, FAT1, and FGFR aberrations increased significantly (Table 2). In particular, FGFR aberration and TMB-high patients concentrated in PIK3CA-HD mutant MBC patients, suggesting this specific genotype might be particularly related to poor prognosis and immune checkpoints disruption. This study categorized PI3k/AKT mutations precisely, and clearly identified the specific genotype (PIK3CA-HD) for novel treatment strategies, such as PD-1 inhibitors, FGFR inhibitors, etc.
Here, we found that both PIK3CA-HD mutations and FGFR aberration seemed to concentrate in HR-positive MBC patients. Is there any potential molecular mechanism underlying the coincidence? However, there is no report about the interaction between PIK3CA-HD mutations and FGFR aberration. In FGFR2mutant endometrial cancer, the sub-therapeutic doses of PI3K inhibitors could enhance the efficacy of anti-FGFR therapies [33]. Such a synergic anti-tumor effect suggested PIK3CA mutation and FGFR aberration might be independent to each other. If they have interaction, inhibiting each of them might also affect the other one. Only when they were independent, PI3K inhibitors and anti-FGFR therapies have synergic effects.
PIK3CA-HD mutant MBC patients showed resistance to mTOR inhibitor Everolimus. More than 70% of patients who received Everolimus in late-line therapy progressed within six months. Due to the limited sample number, we did not demonstrate a concrete result here. However, we still suggested that for PIK3CA-HD mutant MBC patients, mTOR inhibitor Everolimus might be not as effective as in other PI3K/AKT pathway aberrant patients. PD-1 inhibitor plus FGFR inhibitor might be a promising option.
This study involved 193 MBC patients who received late-line therapies from three affiliated hospitals in Central South University. The limitation was the relatively small sample size for each PI3K/AKT pathway aberrant category. The sample size is even smaller after PSM. In the foreseeable future, we would accumulate many more samples to further validate our hypothesis. For now, we raised our hypothesis and provided clues for reasonable treatment strategies, which might be beneficial for MBC patients who have no choice.
Conclusion
MBC patients with PIK3CA mutations in helical domain had a specific ctDNA profile with high TMB and high FGFR aberration rate, which might lead to poor PFS for late-line therapy. PD-1/PD-L1 inhibitor and FGFR inhibitor could be promising as a late-line option for MBC patients with this specific genotype.
Materials and Methods
Patients
This study included MBC patients who were within 2 weeks after progression of the early-line (≤2) salvage treatment. All patients had a confirmed pathology diagnosis (histology type: invasive ductal carcinomas or lobular carcinomas). According to RECIST 1.1 standards, patients had at least one measurable distant disease lesion. All patients had an Eastern Cooperative Oncology Group/World Health Organization Performance Status (ECOG/WHO PS) of 0 or 1. Patients with inflammatory BC, multiple primary malignancies, immunodeficiency or organ transplantation history were excluded. Patients who have received mTOR inhibitor treatment were also not eligible for the study.
Study design
This multicenter study was conducted across three local cancer centers. MBC patients who failed in early-line (≤2 lines) standard chemotherapies or standard chemotherapies combined with anti-HER2 therapies or endocrine therapies received big-panel NGS for ctDNA testing. After ctDNA testing, patients received late-line therapy.
The primary endpoint was PFS, defined as the number of days from the beginning date of late-line therapy to progressive disease, defined as: 1) >20% growth of measurable target lesions and the absolute increase of target lesions >5mm, or 2) presence of new lesions [34]. There was no predetermined per-patient follow-up period. Patients visited clinics every week. Disease progression was assessed at every visit by imaging tools (MRI, CT, ECT and ultrasound, etc) [34].
ctDNA testing and TMB calculation
Peripheral blood samples were collected within 7 days before treatment. We performed ctDNA testing according to protocol described previously [5, 35, 36]. The main steps included DNA extraction, target capture, NGS and sequencing data analysis. Tumor mutation burden (TMB) is an important prognostic factor [19]. Usually, TMB is calculated from whole exome sequencing data or big gene panels [17, 37]. TMB was determined by analyzing the somatic mutations per mega-base (Mb). TMB analysis interrogated SNVs and small indels with the variant allele frequency (VAF) ≥3%. TMB-U (unknown) is defined as the maximum VAF <3%. A cut-off of the top 25% of the TMB of all BC samples from Geneplus database was 9 mutations (Muts) / Mb. In this study, TMB greater than 9 Muts / Mb was defined as TMB-H (high). TMB less than 9 Muts / Mb was defined as TMB-L (low).
Statistical analysis
To search for significant ctDNA aberrations, the R package “ComplexHeatmap” was used to rank the hot genetic aberrations in PI3K/AKT pathway aberrant subgroups. A two-sided log-rank test was used to test the influence of PI3K/AKT pathway aberrations in terms of late-line treatment PFS in MBCs. To reduce the potential bias between subgroups, we also performed 1:1 propensity score matching (PSM) analysis as described previously [38]. Both the univariate and multivariate Cox proportional hazards regression analyses were used to evaluate the prognostic factors for PFS. All statistical analyses were conducted by using SAS 9.4 and R 3.6.0 software. All tests of hypotheses were two-tailed and conducted at a significance level of 0.05 and at a marginal significance level of 0.15.
Ethics
The study protocol and informed consent form were censored by the independent ethics committee for each center. In our study, each participant filled and signed an approved written informed consent provided by the independent ethics committee.
Statement of translational relevance
12 years ago, Barbareschi M's team reported PIK3CA mutations in the helical domain exon 9 were associated with significantly worse prognoses in breast cancer. That study used frozen samples from 163 surgery patients. Here, we demonstrate the effect of PIK3CA helic domain mutations in metastatic breast cancer with late-line therapy by using plasma ctDNA. MBC patients with PIK3CA mutations in helical domain had a specific ctDNA profile with high TMB and high FGFR aberration rate, leading to poor PFS for late-line therapy. Our findings directly reflect the effect of genetic aberrations on treatment outcome and suggest potential strategies for MBC patients. PD-1/PD-L1 inhibitor and FGFR inhibitor could be promising as a late-line option for MBC patients with this specific genotype.
Acknowledgments
We thank the technical support from Beijing Geneplus Institution. We also thank the support from foundations listed below.
Conflicts of Interest
The authors declare that there are no conflicts of interest.
Funding
Hunan Provincial Department of Science and Technology, Social Development Branch (2018SK2124) and Hunan Provincial Health Commission Foundation (B2019085).
References
- 1. Loibl S, Treue D, Budczies J, Weber K, Stenzinger A, Schmitt WD, Weichert W, Jank P, Furlanetto J, Klauschen F, Karn T, Pfarr N, von Minckwitz G, et al. Mutational Diversity and Therapy Response in Breast Cancer: A Sequencing Analysis in the Neoadjuvant GeparSepto Trial. Clin Cancer Res. 2019; 25:3986–95. https://doi.org/10.1158/1078-0432.CCR-18-3258 [PubMed]
- 2. Campbell RA, Bhat-Nakshatri P, Patel NM, Constantinidou D, Ali S, Nakshatri H. Phosphatidylinositol 3-kinase/AKT-mediated activation of estrogen receptor alpha: a new model for anti-estrogen resistance. J Biol Chem. 2001; 276:9817–24. https://doi.org/10.1074/jbc.M010840200 [PubMed]
- 3. Ligresti G, Militello L, Steelman LS, Cavallaro A, Basile F, Nicoletti F, Stivala F, McCubrey JA, Libra M. PIK3CA mutations in human solid tumors: role in sensitivity to various therapeutic approaches. Cell Cycle. 2009; 8:1352–58. https://doi.org/10.4161/cc.8.9.8255 [PubMed]
- 4. Bader AG, Kang S, Zhao L, Vogt PK. Oncogenic PI3K deregulates transcription and translation. Nat Rev Cancer. 2005; 5:921–29. https://doi.org/10.1038/nrc1753 [PubMed]
- 5. Hu ZY, Xie N, Tian C, Yang X, Liu L, Li J, Xiao H, Wu H, Lu J, Gao J, Hu X, Cao M, Shui Z, et al. Identifying Circulating Tumor DNA Mutation Profiles in Metastatic Breast Cancer Patients with Multiline Resistance. EBioMedicine. 2018; 32:111–18. https://doi.org/10.1016/j.ebiom.2018.05.015 [PubMed]
- 6. Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, Yan H, Gazdar A, Powell SM, Riggins GJ, Willson JK, Markowitz S, Kinzler KW, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004; 304:554. https://doi.org/10.1126/science.1096502 [PubMed]
- 7. Ramirez-Ardila DE, Helmijr JC, Look MP, Lurkin I, Ruigrok-Ritstier K, van Laere S, Dirix L, Sweep FC, Span PN, Linn SC, Foekens JA, Sleijfer S, Berns EM, Jansen MP. Hotspot mutations in PIK3CA associate with first-line treatment outcome for aromatase inhibitors but not for tamoxifen. Breast Cancer Res Treat. 2013; 139:39–49. https://doi.org/10.1007/s10549-013-2529-7 [PubMed]
- 8. Meyer DS, Koren S, Leroy C, Brinkhaus H, Müller U, Klebba I, Müller M, Cardiff RD, Bentires-Alj M. Expression of PIK3CA mutant E545K in the mammary gland induces heterogeneous tumors but is less potent than mutant H1047R. Oncogenesis. 2013; 2:e74. https://doi.org/10.1038/oncsis.2013.38 [PubMed]
- 9. Barbareschi M, Buttitta F, Felicioni L, Cotrupi S, Barassi F, Del Grammastro M, Ferro A, Dalla Palma P, Galligioni E, Marchetti A. Different prognostic roles of mutations in the helical and kinase domains of the PIK3CA gene in breast carcinomas. Clin Cancer Res. 2007; 13:6064–69. https://doi.org/10.1158/1078-0432.CCR-07-0266 [PubMed]
- 10. Cheng J, Fu S, Wei C, Tania M, Khan MA, Imani S, Zhou B, Chen H, Xiao X, Wu J, Fu J. Evaluation of PIK3CA mutations as a biomarker in Chinese breast carcinomas from Western China. Cancer Biomark. 2017; 19:85–92. https://doi.org/10.3233/CBM-160380 [PubMed]
- 11. Arsenic R, Treue D, Lehmann A, Hummel M, Dietel M, Denkert C, Budczies J. Comparison of targeted next-generation sequencing and Sanger sequencing for the detection of PIK3CA mutations in breast cancer. BMC Clin Pathol. 2015; 15:20. https://doi.org/10.1186/s12907-015-0020-6 [PubMed]
- 12. Janku F, Wheler JJ, Naing A, Falchook GS, Hong DS, Stepanek VM, Fu S, Piha-Paul SA, Lee JJ, Luthra R, Tsimberidou AM, Kurzrock R. PIK3CA mutation H1047R is associated with response to PI3K/AKT/mTOR signaling pathway inhibitors in early-phase clinical trials. Cancer Res. 2013; 73:276–84. https://doi.org/10.1158/0008-5472.CAN-12-1726 [PubMed]
- 13. Zhao L, Vogt PK. Helical domain and kinase domain mutations in p110alpha of phosphatidylinositol 3-kinase induce gain of function by different mechanisms. Proc Natl Acad Sci USA. 2008; 105:2652–57. https://doi.org/10.1073/pnas.0712169105 [PubMed]
- 14. Emanuel BS, Zackai EH, Medne L. Emanuel Syndrome. 2007 Apr 20 [updated 2017 Aug 31]. In: Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJH, Stephens K, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2019. Available from http://www.ncbi.nlm.nih.gov/books/NBK1263/ [PubMed]
- 15. Zhou D, Ouyang Q, Liu L, Liu J, Tang Y, Xiao M, Wang Y, He Q, Hu ZY. Chemotherapy Modulates Endocrine Therapy-Related Resistance Mutations in Metastatic Breast Cancer. Transl Oncol. 2019; 12:764–74. https://doi.org/10.1016/j.tranon.2019.02.014 [PubMed]
- 16. Miller TW, Hennessy BT, González-Angulo AM, Fox EM, Mills GB, Chen H, Higham C, García-Echeverría C, Shyr Y, Arteaga CL. Hyperactivation of phosphatidylinositol-3 kinase promotes escape from hormone dependence in estrogen receptor-positive human breast cancer. J Clin Invest. 2010; 120:2406–13. https://doi.org/10.1172/JCI41680 [PubMed]
- 17. Budczies J, Allgäuer M, Litchfield K, Rempel E, Christopoulos P, Kazdal D, Endris V, Thomas M, Fröhling S, Peters S, Swanton C, Schirmacher P, Stenzinger A. Optimizing panel-based tumor mutational burden (TMB) measurement. Ann Oncol. 2019; 30:1496–506. https://doi.org/10.1093/annonc/mdz205 [PubMed]
- 18. Gubin MM, Artyomov MN, Mardis ER, Schreiber RD. Tumor neoantigens: building a framework for personalized cancer immunotherapy. J Clin Invest. 2015; 125:3413–21. https://doi.org/10.1172/JCI80008 [PubMed]
- 19. Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015; 348:124–28. https://doi.org/10.1126/science.aaa1348 [PubMed]
- 20. Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, Walsh LA, Postow MA, Wong P, Ho TS, Hollmann TJ, Bruggeman C, Kannan K, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014; 371:2189–99. https://doi.org/10.1056/NEJMoa1406498 [PubMed]
- 21. Alborelli I, Leonards K, Rothschild SI, Leuenberger LP, Savic Prince S, Mertz KD, Poechtrager S, Buess M, Zippelius A, Läubli H, Haegele J, Tolnay M, Bubendorf L, et al. Tumor mutational burden assessed by targeted NGS predicts clinical benefit from immune checkpoint inhibitors in non-small cell lung cancer. J Pathol. 2020; 250:19–29. https://doi.org/10.1002/path.5344 [PubMed]
- 22. Wang F, Wei XL, Wang FH, Xu N, Shen L, Dai GH, Yuan XL, Chen Y, Yang SJ, Shi JH, Hu XC, Lin XY, Zhang QY, et al. Safety, efficacy and tumor mutational burden as a biomarker of overall survival benefit in chemo-refractory gastric cancer treated with toripalimab, a PD-1 antibody in phase Ib/II clinical trial NCT02915432. Ann Oncol. 2019; 30:1479–86. https://doi.org/10.1093/annonc/mdz197 [PubMed]
- 23. Ballinger TJ, Meier JB, Jansen VM. Current Landscape of Targeted Therapies for Hormone-Receptor Positive, HER2 Negative Metastatic Breast Cancer. Front Oncol. 2018; 8:308. https://doi.org/10.3389/fonc.2018.00308 [PubMed]
- 24. Iwamoto T, Taira N, Fujisawa T, Araki K, Sakamaki K, Sangai T, Kikawa Y, Shien T, Takao S, Sato M, Goto Y, Yoshida T, Takahashi M, et al. Hormonal Therapy Resistant Estrogen-receptor Positive Metastatic Breast Cancer Cohort (HORSE-BC) Study : Current Status of Treatment Selection in Japan. Acta Med Okayama. 2018; 72:369–74. https://doi.org/10.18926/AMO/56172 [PubMed]
- 25. Dirix LY, Takacs I, Jerusalem G, Nikolinakos P, Arkenau HT, Forero-Torres A, Boccia R, Lippman ME, Somer R, Smakal M, Emens LA, Hrinczenko B, Edenfield W, et al. Avelumab, an anti-PD-L1 antibody, in patients with locally advanced or metastatic breast cancer: a phase 1b JAVELIN Solid Tumor study. Breast Cancer Res Treat. 2018; 167:671–86. https://doi.org/10.1007/s10549-017-4537-5 [PubMed]
- 26. Cao J, Zhang J, Wang Z, Wang B, Lv F, Wang L, Hu X. Hypothyroidism as a potential biomarker of efficacy of famitinib, a novel VEGFR-2 inhibitor in metastatic breast cancer. Cancer Chemother Pharmacol. 2014; 74:389–98. https://doi.org/10.1007/s00280-014-2505-x [PubMed]
- 27. Kodumudi KN, Ramamoorthi G, Snyder C, Basu A, Jia Y, Awshah S, Beyer AP, Wiener D, Lam L, Zhang H, Greene MI, Costa RL, Czerniecki BJ. Sequential Anti-PD1 Therapy Following Dendritic Cell Vaccination Improves Survival in a HER2 Mammary Carcinoma Model and Identifies a Critical Role for CD4 T Cells in Mediating the Response. Front Immunol. 2019; 10:1939. https://doi.org/10.3389/fimmu.2019.01939 [PubMed]
- 28. Chia S, Bedard PL, Hilton J, Amir E, Gelmon K, Goodwin R, Villa D, Cabanero M, Tu D, Tsao M, Seymour L. A Phase Ib Trial of Durvalumab in Combination with Trastuzumab in HER2-Positive Metastatic Breast Cancer (CCTG IND.229). Oncologist. 2019; 24:1439–45. https://doi.org/10.1634/theoncologist.2019-0321 [PubMed]
- 29. Robinson DR, Wu YM, Vats P, Su F, Lonigro RJ, Cao X, Kalyana-Sundaram S, Wang R, Ning Y, Hodges L, Gursky A, Siddiqui J, Tomlins SA, et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat Genet. 2013; 45:1446–51. https://doi.org/10.1038/ng.2823 [PubMed]
- 30. Xu B, Krie A, De P, Williams C, Elsey R, Klein J, Leyland-Jones B. Utilizing Tumor and Plasma Liquid Biopsy in Treatment Decision Making for an Estrogen Receptor-Positive Advanced Breast Cancer Patient. Cureus. 2017; 9:e1408. https://doi.org/10.7759/cureus.1408 [PubMed]
- 31. Tunger A, Sommer U, Wehner R, Kubasch AS, Grimm MO, Bachmann MP, Platzbecker U, Bornhäuser M, Baretton G, Schmitz M. The Evolving Landscape of Biomarkers for Anti-PD-1 or Anti-PD-L1 Therapy. J Clin Med. 2019; 8:8. https://doi.org/10.3390/jcm8101534 [PubMed]
- 32. Schmid P, Adams S, Rugo HS, Schneeweiss A, Barrios CH, Iwata H, Diéras V, Hegg R, Im SA, Shaw Wright G, Henschel V, Molinero L, Chui SY, et al, and IMpassion130 Trial Investigators. Atezolizumab and Nab-Paclitaxel in Advanced Triple-Negative Breast Cancer. N Engl J Med. 2018; 379:2108–21. https://doi.org/10.1056/NEJMoa1809615 [PubMed]
- 33. Packer LM, Geng X, Bonazzi VF, Ju RJ, Mahon CE, Cummings MC, Stephenson SA, Pollock PM. PI3K Inhibitors Synergize with FGFR Inhibitors to Enhance Antitumor Responses in FGFR2mutant Endometrial Cancers. Mol Cancer Ther. 2017; 16:637–48. https://doi.org/10.1158/1535-7163.MCT-16-0415 [PubMed]
- 34. Litière S, de Vries EG, Seymour L, Sargent D, Shankar L, Bogaerts J, and RECIST Committee. The components of progression as explanatory variables for overall survival in the Response Evaluation Criteria in Solid Tumours 1.1 database. Eur J Cancer. 2014; 50:1847–53. https://doi.org/10.1016/j.ejca.2014.03.014 [PubMed]
- 35. Phallen J, Sausen M, Adleff V, Leal A, Hruban C, White J, Anagnostou V, Fiksel J, Cristiano S, Papp E, Speir S, Reinert T, Orntoft MW, et al. Direct detection of early-stage cancers using circulating tumor DNA. Sci Transl Med. 2017; 9:9. https://doi.org/10.1126/scitranslmed.aan2415 [PubMed]
- 36. Yang X, Chu Y, Zhang R, Han Y, Zhang L, Fu Y, Li D, Peng R, Li D, Ding J, Li Z, Zhao M, Zhang K, et al. Technical Validation of a Next-Generation Sequencing Assay for Detecting Clinically Relevant Levels of Breast Cancer-Related Single-Nucleotide Variants and Copy Number Variants Using Simulated Cell-Free DNA. J Mol Diagn. 2017; 19:525–36. https://doi.org/10.1016/j.jmoldx.2017.04.007 [PubMed]
- 37. Chalmers ZR, Connelly CF, Fabrizio D, Gay L, Ali SM, Ennis R, Schrock A, Campbell B, Shlien A, Chmielecki J, Huang F, He Y, Sun J, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017; 9:34. https://doi.org/10.1186/s13073-017-0424-2 [PubMed]
- 38. Liu J, Li J, Wang H, Wang Y, He Q, Xia X, Hu ZY, Ouyang Q. Clinical and genetic risk factors for Fulvestrant treatment in post-menopause ER-positive advanced breast cancer patients. J Transl Med. 2019; 17:27. https://doi.org/10.1186/s12967-018-1734-x [PubMed]


