Introduction
Melanoma is characterized by rapid progression and poor survival [1]. Early-stage localized melanoma patients could be effectively treated through surgical resection, but survival outcome for patients with distant metastases is always less favorable [2]. Recently, kinase inhibitors (e.g., vemurafenib, dabrafenib and trametinib) that target specific pathways have been approved by the Food and Drug Administration (FDA) [3]. Although the noteworthy improvement of antitumor response to these agents has been observed, they are rarely durable [2].
Owing to the emergence of immunotherapy, especially immune checkpoint inhibitor (ICI) therapy, prognosis for melanoma patients has dramatically improved [4–6]. However, only a subset of patients could demonstrate a remarkable response to ICI therapy. The pivotal point of this problem is the lack of effective indicators to identify patients who are more responsive to immunotherapy agents. The current broadly used biomarker of immune treatment response is tumor mutation load (TML). Other multiple microenvironment-based factors, such as immune checkpoints expression, proportion of tumor infiltration lymphocytes (TIL), and interferon gamma (IFNγ) signature, also play vital roles in response to immunotherapy. Patients who harbor fewer markers may benefit less from such treatment. This raises the question whether there exist other factors that simultaneously affect more than one of the above listed biomarkers, which could provide better predictive value for immunotherapy.
MUC16, which is a member of the mucin family and encodes cancer antigen 125 (CA125). MUC16 was determined as the monitor indicator for diagnosis of gynecological cancer [7–9]. Several recent studies have reported that MUC16 could inhibit antitumor immune responses by attenuating natural killer (NK) cells and promoting regulatory T cells activity [10–12]. Meanwhile, other studies demonstrated that MUC16 may be implicated in enhancing the activity of pro-inflammatory pathways in tumor [13, 14]. Our previous study reported the association of MUC16 mutations with high TML and favorable outcome in gastric cancer (GC). Furthermore, our results revealed that mutated MUC16 has important implications for immunotherapy [15]. However, owing to the limited number of GC samples from patients who received immune treatment, we could not validate the relevance of these results. To our knowledge, the effect of MUC16 mutations on melanoma TML, microenvironment, prognosis, and immunotherapeutic efficacy has not yet been investigated.
In this study, we explored whether the presence of MUC16 mutations was associated with TML, tumor-immune microenvironment, survival outcome, and ICI treatment efficacy in melanoma. Evidence derived from our study would have implications for guiding immunotherapy.
Results
MUC16 mutational status of melanoma
MUC16 was one of the most frequently mutated genes in melanoma. Of the 467 samples in the Cancer Genome Atlas (TCGA) cohort, 341 (73.1%) harbored MUC16 mutations. Plenty of frequently mutated genes were correlated with TML in TCGA, and presence of MUC16 mutations showed the most significant correlation (P = 2.25E-44; Supplementary Figure 1). We found that patients with mutated MUC16 had a significantly higher TML than those without it (Figure 1A). Mutation distribution of MUC16 and its family members in relation to genomic integrity maintenance genes was illustrated in Figure 1B. Waterfall plot showed that patients who harbored MUC16 mutations also had some mutations in genome repair genes (145 of 341, 42.5%; Figure 1B). Consistent results and mutational patterns of mucin family members and genome repair genes in the International Cancer Genome Consortium (ICGC) cohort were exhibited in Supplementary Figure 2.
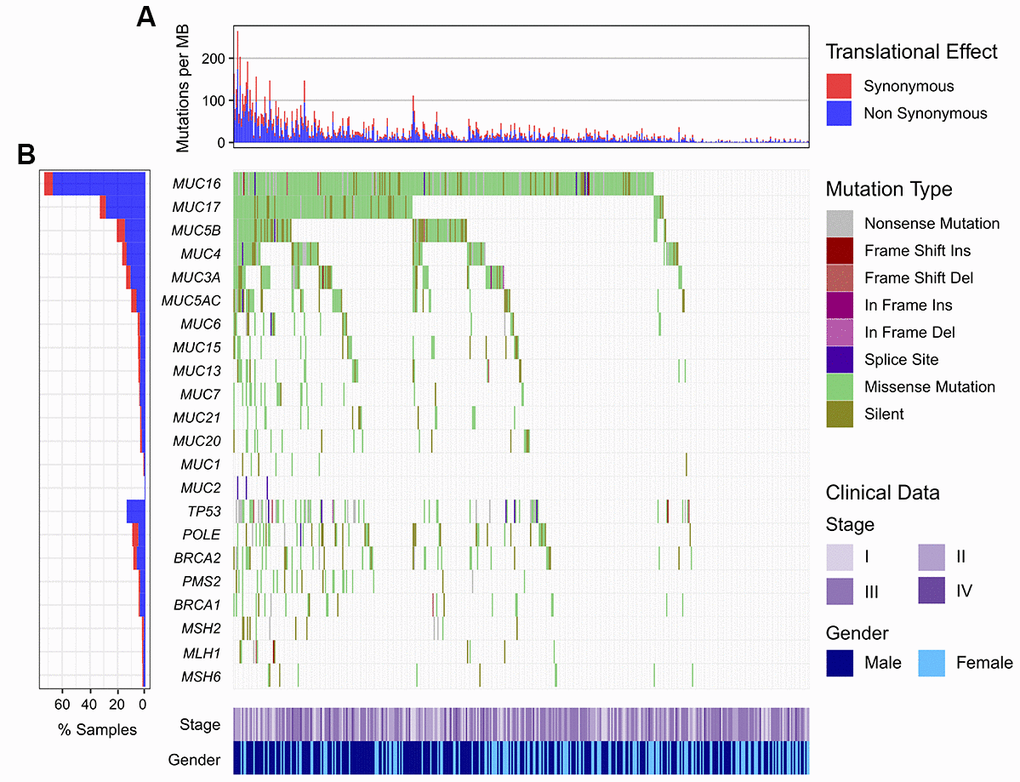
Figure 1. Mutational patterns of MUC16 and mucin family members in relation to DNA repair-related genes in the TCGA cohort. (A) Numbers of mutations per megabase in each sample. (B) Representation for mutation patterns of mucin and DNA repair genes.
The difference in TML between the 2 cohorts was not statistically significant (median TML: TCGA cohort 3.78 vs. ICGC cohort 4.27; Wilcoxon rank sum test, P = 0.11; Supplementary Figure 3).
MUC16 mutations are associated with high TML in both cohorts
In the TCGA cohort, melanoma patients with MUC16 mutations had a significantly higher TML than those without MUC16 mutations (median TML: 4.19 vs. 1.25; Wilcoxon rank sum test, P < 0.001; Figure 2A). In MUC16 mutated patients, we found that BRCA1/2 (56 [16.4%] patients with mutations), TP53 (57 [16.7%] patients with mutations), POLE (44 [12.9%] patients with mutations), and MMR genes (total 49 [14.4%] patients with mutations) were significantly co-mutated (Fisher exact test, all P < 0.01; Supplementary Table 1). Mutations in these genes caused a significantly higher mutation load (OR > 3, P < 0.01; Figure 2C). TML could be suitably divided into high and low subgroups with a cutoff value of 4.22 (Supplementary Figure 4). To rule out the possibility that higher TML was generated by mutations in genome repair genes rather than directly by MUC16 mutations, we performed multivariate logistic regression model with mutations in DDR and MMR genes, and clinical confounding factors taken into consideration. Association of MUC16 mutations with higher TML was still statistically significant after adjusting for these confounding variables (OR: 15.61, 95% CI: 6.15-52.87, P < 0.001; Figure 2C).
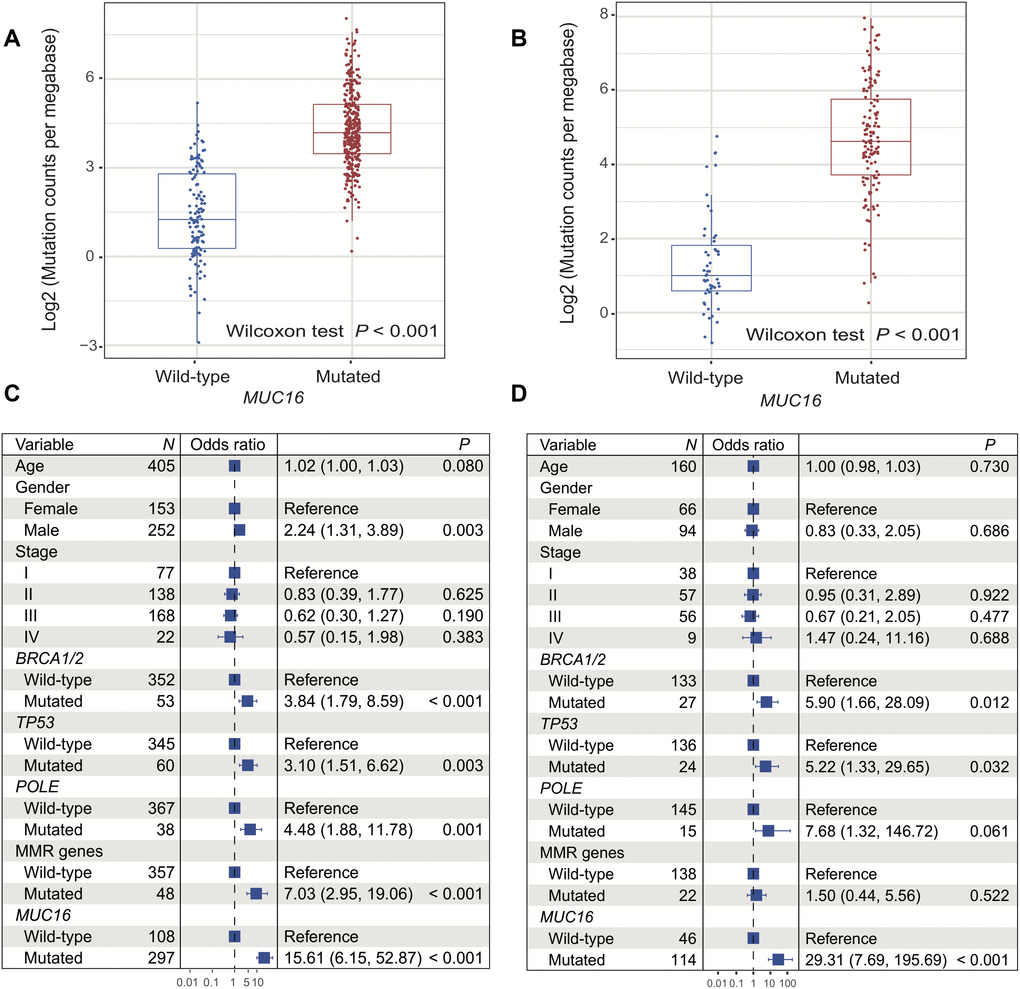
Figure 2. Correlation of MUC16 mutations with tumor mutational load in 2 cohorts. (A, B) Mutation burden of melanoma samples with and without MUC16 mutation (left: TCGA; right: ICGC). (C, D) Multivariate logistic regression models were conducted to explore association of MUC16 mutations with TML (left: TCGA; right: ICGC).
Additionally, high TML was also observed in tumor samples with MUC16 mutations of other immune activated tumor types (e.g., cancers of the lung, colorectal, kidney, bladder, and head and neck) in the TCGA cohort (Wilcoxon rank sum test, all P < 0.001; Supplementary Figure 5).
Of the 183 melanoma patients in the ICGC cohort, the significantly high TML was also observed in patients with MUC16 mutations (median TML: 4.63 vs. 1.01; Wilcoxon rank sum test, P < 0.001; Figure 2B). Genomic integrity maintenance genes, including BRCA1/2 (26 [19.7%] samples with mutations), TP53 (27 [20.5%] samples with mutations), POLE (16 [12.1%] samples with mutations), and MMR genes (total 22 [16.7%] samples with mutations) were also significantly co-mutated in these MUC16 mutated patients (Fisher exact test, all P < 0.01; Supplementary Table 1). Multivariate logistic regression model that included these gene mutations and clinical variables was performed to control confounders. In this model, the association between MUC16 mutation and high TML was still statistically significant (OR: 29.31, 95% CI: 7.69-195.69, P < 0.001; Figure 2D).
In addition to association of MUC16 mutation status with TML, we also discovered that the total count of MUC16 mutations was associated with high TML in these 2 cohorts (TCGA: Spearman R = 0.85, P < 0.001; ICGC: Spearman R = 0.885, P < 0.001) (Supplementary Figure 6).
MUC16 mutations are associated with immune-active microenvironment
CIBERSORT method revealed that infiltration of CD8 T cells was significantly higher in patients with MUC16 mutations (P < 0.05), and resting NK cells exhibited the opposite behavior (P < 0.05) (Figure 3A). Consistently, we found that patients with MUC16 mutations had considerably high infiltration of pro-inflammatory M1 macrophages (P < 0.001) and low infiltration of immune-suppressive M2 macrophages (P < 0.01) (Figure 3A).
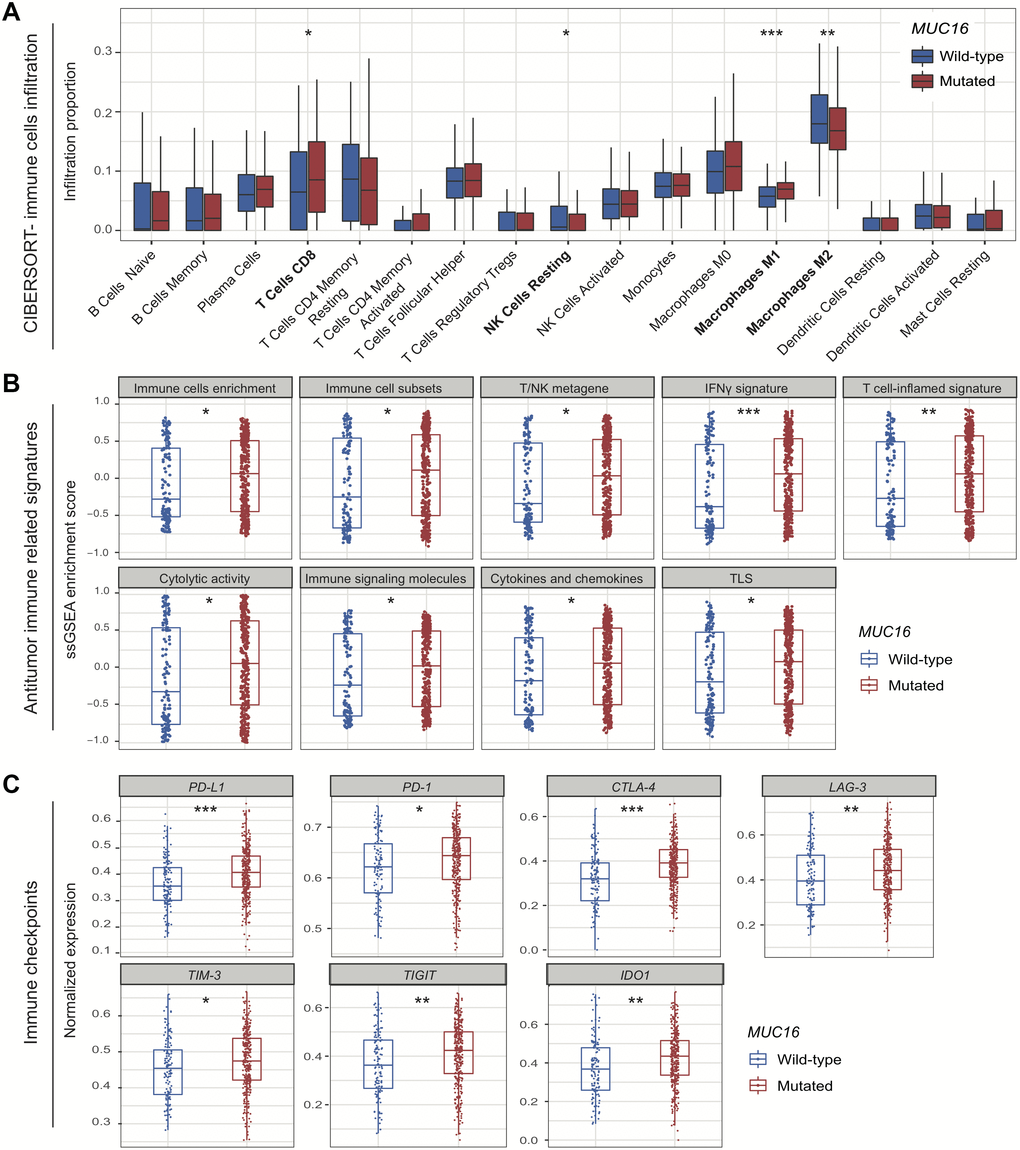
Figure 3. Association of MUC16 mutations with tumor microenvironment. (A) Distinct immune cells infiltration in MUC16 mutated and wild-type subgroups. (B) Distribution of immune-related signatures of samples stratified by MUC16 mutational status. (C) Distinct expression of immune checkpoints in MUC16 mutated and wild-type samples. * P < 0.05; ** P < 0.01; *** P < 0.001.
Patients with MUC16 mutations had significant enrichment in total immune cells, immune cell subsets (i.e., T cells, B cells, and NK cells), and T/NK metagene (i.e., T cells and NK cells activity) (all P < 0.05; Figure 3B). We also observed high enrichment in IFNγ signal and its related T cell-inflamed gene signature that were previously reported to predict immunotherapy response (both P < 0.01; Figure 3B) [16]. Besides, increased cytolytic activity, enrichment in cytokines and chemokines, and enhanced tertiary lymphoid structures (TLS) were all found in patients with MUC16 mutations (all P < 0.05; Figure 3B).
We observed that expression of PD-L1, PD-1, and CTLA-4 was significantly upregulated in patients with MUC16 mutations (Wilcoxon rank sum test, all P < 0.05; Figure 3C). Other checkpoints, including LAG-3, TIM-3, TIGIT, and IDO1 also exhibited consistent results (all P < 0.05; Figure 3C).
Pathways significantly associated with MUC16 mutations
In gene set enrichment analysis (GSEA) analysis, immune response-related pathways, such as antigen processing and presentation, graft versus host disease, and allograft rejection (normalized enrichment score range: 2.25-2.54; all FDR = 0.001) were among the top enriched pathways of patients with MUC16 mutations based on KEGG dataset (Supplementary Figure 7A). Pathways for antigen processing and presentation of peptide antigen, response to interferon gamma, and interferon gamma mediated signaling pathway (normalized enrichment score range: 2.46-2.52; all FDR = 0.002) were the top 3 enriched circuits of patients with MUC16 mutations based on GO dataset (Supplementary Figure 7B).
MUC16 mutations are linked with favorable prognosis in 2 cohorts
In the TCGA cohort, melanoma patients with MUC16 mutations had a significantly better overall survival (OS) than those without it (median OS: 104.5 [95% CI, 77.1-131.9] vs. 49.3 [95% CI, 42.6-55.9] months; Log rank test P < 0.001; Figure 4A). Multivariate Cox regression model remained statistically significant with confounding factors taken into account (HR: 0.44, 95% CI: 0.31-0.61, P < 0.001; Figure 4B).
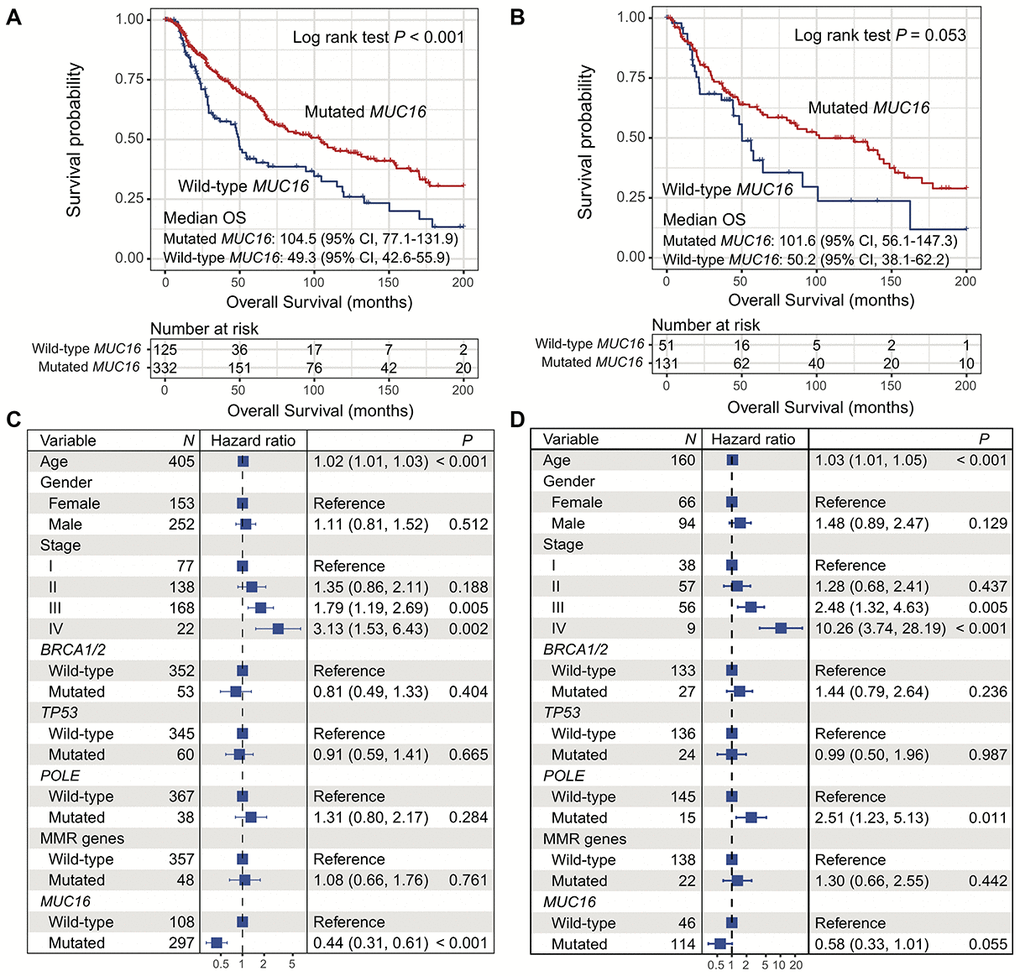
Figure 4. Correlation of MUC16 mutations with overall survival in 2 cohorts. (A, B) Kaplan-Meier survival analysis based on MUC16 mutational status (left: TCGA; right: ICGC). (C, D) Forest plot representation of association of MUC16 mutations with prognosis (left: TCGA; right: ICGC).
Consistently, there was a statistically correlation between the presence of MUC16 mutations and better OS in patients in the ICGC cohort (median OS: 101.6 [95% CI, 56.1-147.3] vs. 50.2 [95% CI, 38.1-62.2] months; Log rank test P = 0.053; Figure 4B). The result still remained statistically significant after adjusting for confounding variables (HR: 0.58, 95% CI: 0.33-1.01, P = 0.055; Figure 4D).
MUC16 mutations were associated with better ICI response and survival in male patients
Consistent with our results for the patients in the TCGA cohort, we found that the presence of MUC16 mutations were most significantly associated with TML in the ICI-treated cohort (P = 5.55E-17; Supplementary Figure 8A). Patients with MUC16 mutations had significantly higher TML and neoantigen load than those without MUC16 mutations (median TML: 4.46 vs. 1.53; median neoantigen load: 5.41 vs. 2.36; Wilcoxon rank sum test, both P < 0.001) (Supplementary Figure 8B, 8C).
In the ICI-treated cohort, patients with MUC16 mutations had a higher response rate to the therapy than those without MUC16 mutations in male patients (response rate: 45.6% vs. 18.5%; Fisher exact test, P = 0.017; Figure 5A). However, this association was not observed in female patients (response rate: 35.0% vs. 50.0%; Fisher exact test, P = 0.281; Figure 5B) and overall patients (response rate: 41.2% vs. 32.4%; Fisher exact test, P = 0.361; Figure 5C). Consistent with above findings, we observed that responders had a significantly higher MUC16 mutation rate than non-responders in male patients (mutation rate: 83.9% vs. 58.5%; Fisher exact test, P = 0.017; Supplementary Figure 9A), but not in female patients (mutation rate: 58.3% vs. 72.2%; Fisher exact test, P = 0.281; Supplementary Figure 9B) and overall patients (mutation rate: 72.8% vs. 64.0%; Fisher exact test, P = 0.361; Supplementary Figure 9C).
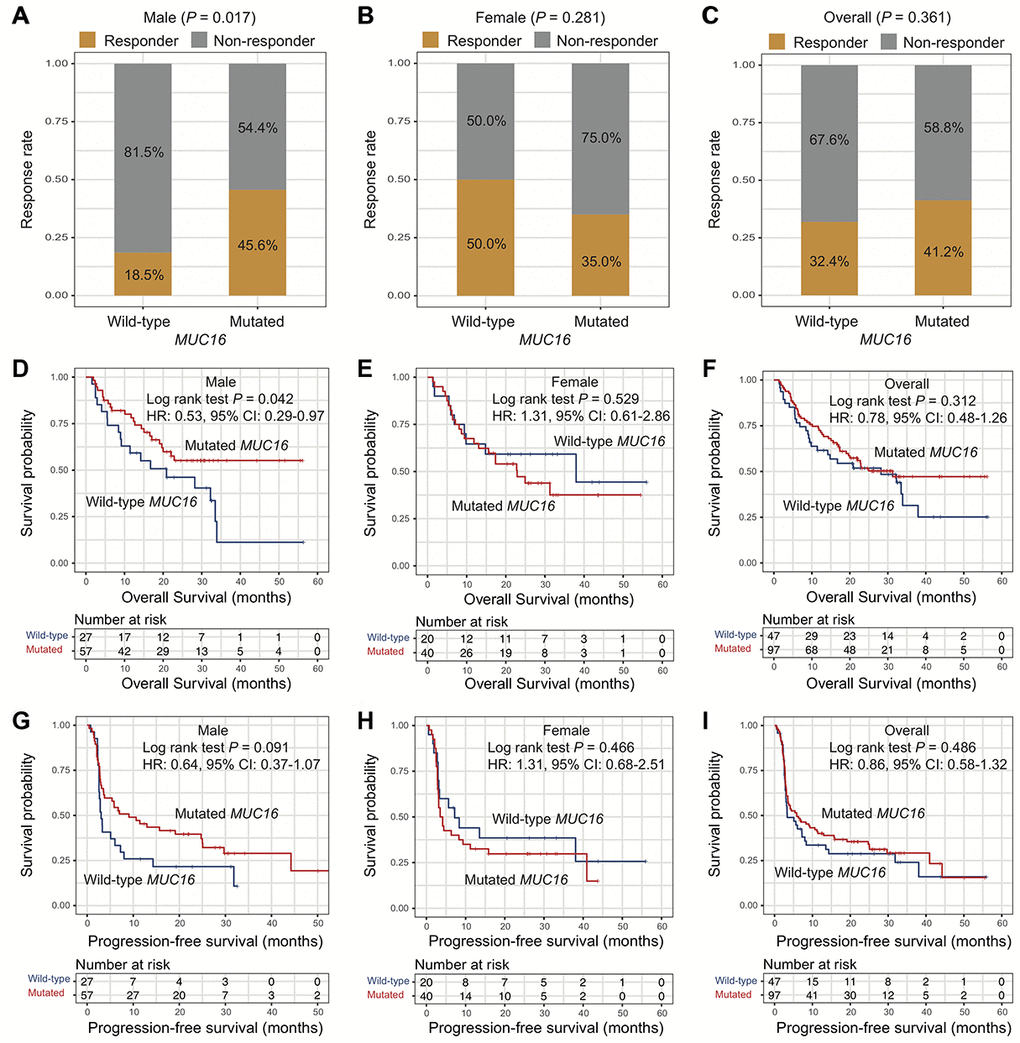
Figure 5. Association of MUC16 mutations with ICI therapy efficacy. (A–C) Association of MUC16 mutations with response rate to ICI therapy in male, female, and overall patients. (D–F) Overall survival plot of ICI treated patients stratified by MUC16 mutational status in male, female, and overall patients. (G–I) Progression-free survival plot of ICI treated patients stratified by MUC16 mutational status in male, female, and overall patients.
Besides, we found that male patients with MUC16 mutations had a better OS than those without MUC16 mutations (median OS: not calculable [the median OS of melanoma male patients with MUC16 mutations could not be calculated owing to more than half patients in this group were alive] vs. 20.9 [95% CI, 9.17-NA (not available)] months; Log rank test P = 0.042; Figure 5D). Male patients with MUC16 mutations exhibited a better trend of progression-free survival (PFS) than those without MUC16 mutations although this difference did not reach statistical significance (median PFS: 9.07 [95% CI, 3.60-25.1] vs. 3.03 [95% CI, 2.57-8.00] months; Log rank test P = 0.091; Figure 5G). In female and overall patients, OS and PFS were not statistically significant when stratified by MUC16 mutational status (all Log rank test P > 0.05) (Figure 5E, 5F, 5H, 5I).
Discussion
Recently, multiple studies from immunotherapy clinical trials have shown promising findings that harbored a potential to predict or control tumor progression in melanoma [17, 18]. However, inconsistent conclusions of these results highlight the urgent need to determine a more suitable sub-population or biomarkers for immunotherapy in melanoma, which becomes a momentous challenge in the area of immuno-oncology.
Previous clinical trials have revealed that patients with high TML could benefit more from ICI agents in melanoma and non-small cell lung cancer (NSCLC) [19, 20]. However, accurately assessment of TML faces several limitations in clinical practice. The sampling approaches, distinct sequencing platforms, threshold definition, and costs of whole exome sequencing blockade the broadly implementation of TML evaluation [21]. Recent studies have reported that rather than measuring total mutation counts in exome, mutation status of a single specific gene could serve as a surrogate for evaluating TML or ICI therapy efficacy [15, 21–23]. From the TCGA melanoma cohort, we found that patients with MUC16 mutation had the most significant association with high TML. Noticeably, besides melanoma, our study showed other cancers benefited from immunotherapy (e.g., cancers of lung, colorectal, kidney, bladder, and head and neck) also exhibited the consistent association of MUC16 mutation with high TML. All these findings suggested patients with MUC16 mutations may be more responsive to immunotherapies in melanoma and other relevant tumors.
High expression of immune checkpoints such as PD-L1 was approved by FDA as an essential criterion for pembrolizumab treatment in NSCLC [24]. In our study, we found that the other checkpoints (e.g., PD-1, CTLA-4, TIM-3 and LAG-3) except for PD-L1, were all significantly upregulated in patients with MUC16 mutations. Several studies have reported the immune homeostatic effect of tumor infiltration CD8 T cells and macrophages in tumor-immune microenvironment [25–27]. In this study, we found that patients with MUC16 mutations had significantly high enrichment in CD8 T cells and M1 macrophages. Conversely, immune suppressive M2 macrophages were enriched in MUC16 wild-type group. Patients with MUC16 mutations also harbored a considerably high enrichment of IFNγ and T cell-inflamed signal, which could accurately predict immunotherapeutic efficacy [16, 28]. These findings indicated that MUC16 mutations were related to immune-activated microenvironment and potentially high response rate to immunotherapy.
Results revealed that the presence of MUC16 mutations was significantly associated with high ICI response rate and overall survival in male patients from an ICI treated melanoma cohort. This suggests that sex difference may be a potential variable in determining immunotherapeutic efficacy for patients with mutated MUC16. Consistent with our findings, a recent meta-analysis reported that male patients gained higher benefit from ICI therapy than female patients (HR: male 0.72 [95% CI, 0.65-0.79] vs. female 0.86 [95% CI, 0.79-0.93]; P = 0.002) [29]. In our study, we found that male patients harbored significantly higher TMB than female patients (Figure 2C), which was also reported in a previous study [30]. This finding suggested that male patients may have higher neoantigen and immunogenicity, which speculatively justifies association of MUC16 mutation with higher response rate and better ICB therapy outcome in male patients as compared with female patients.
Several limitations remained in our study. Firstly, melanoma samples with gene expression profile were only acquired from one cohort. Secondly, the mechanisms for correlation between MUC16 mutations and higher mutation load are elusive, which requires further investigation.
Our study discovered that patients with MUC16 mutations had significantly high TML, immune-activated tumor microenvironment and favorable survival outcome. Importantly, the presence of MUC16 mutations was significantly associated with better immunotherapeutic efficacy in male patients. Therefore, MUC16 mutations may serve as a surrogate for predicting efficacy of immune checkpoints based therapies, and future clinical trials are needed to validate our findings.
Materials and Methods
Genomic and clinical information of melanoma patients
Somatic mutation data of 467 melanoma samples in the Cancer Genome Atlas (TCGA) cohort were acquired from Genome Data Commons (https://portal.gdc.cancer.gov). The validation dataset contained 183 samples and was obtained from International Cancer Genome Consortium (ICGC) (https://dcc.icgc.org). Gene expression profiles of 465 patients were obtained from TCGA cohort.
The ICI therapy cohort was obtained from the study by Liu et al. [31], which is the largest publicly available melanoma ICI-treated patient cohort. This cohort contained 144 patients treated with either anti-PD-1 or anti-CTLA-4 agents. In this study, patients with complete or partial response were considered responders; other statuses (i.e., progressive disease, stable disease, and mixed response) were considered non-responders.
MUC16 mutation versus TML
Genomic instability or high mutation load is largely correlated with mutations in DNA damage repair (DDR) and mismatch repair (MMR) related genes [32]. In addition to univariate analysis of association of MUC16 mutations with TML, we performed multivariate logistic regression with mutations in DDR genes (e.g., BRCA1/2, TP53, and POLE) and MMR genes (i.e., MLH1, MSH2, MSH6, and PMS2), and clinical factors as confounding variables to eliminate the false positive possibility. TML was defined as log2 transformation of mutation counts per megabase. We applied a univariate clustering approach (i.e., Ckmeans.1d.dp algorithm) available from R package Ckmeans.1d.dp (version 4.2.2) [33] to determine the optimal cutoff value of high versus low TML followed by recently broadly used value (i.e., 17 mutation counts per megabase, log2 transformed level [4.09]).
Gene set enrichment analysis
We applied single sample gene set enrichment analysis (ssGSEA) approach embedded in R package GSVA (version 1.32.0) to evaluate overall enrichment of specific immune signatures of each sample [46]. GSEA was implemented by fgsea package (version 1.10.0). Signaling pathways in Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) were used as the background database.
Statistical analyses
R software (version 3.6.1) was used in this study to perform relevant statistical analyses. Mutation patterns were presented via GenVisR package (version 1.16.0) [47]. We drew and compared survival curves using Kaplan-Meier approach and Log rank test respectively with Rsurvival (version 2.44-1.1) and survminer (version 0.4.5) packages. Multivariate logistic and Cox regression models were built using forestmodel package (version 0.5.0). Associations of MUC16 mutations with continuous and categorical variables were estimated with Wilcoxon rank sum test and Chi-square test, separately.
Author Contributions
KC and XL designed this study; KC, XL and QW developed the methodology and acquired the related data; XL, QW, MY and YY performed data analysis and interpretation; XL, KC and QW drafted and revised the manuscript; KC and XL supervised this study.
Acknowledgments
We thank Prof. Wei Zhang at the Wake Forest Baptist Comprehensive Cancer Center for his valued recommendations.
Conflicts of Interest
All authors declare no conflicts of interest.
Funding
This study was supported by the Program for Changjiang Scholars and Innovative Research Team in University in China [IRT_14R40 to K. C.], and the National Natural Science Foundation of China [no. 31801117 to X. L.].
References
- 1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66:7–30. https://doi.org/10.3322/caac.21332 [PubMed]
- 2. Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell. 2015; 161:1681–96. https://doi.org/10.1016/j.cell.2015.05.044 [PubMed]
- 3. McArthur GA, Ribas A. Targeting oncogenic drivers and the immune system in melanoma. J Clin Oncol. 2013; 31:499–506. https://doi.org/10.1200/JCO.2012.45.5568 [PubMed]
- 4. Hodi FS, O’Day SJ, McDermott DF, Weber RW, Sosman JA, Haanen JB, Gonzalez R, Robert C, Schadendorf D, Hassel JC, Akerley W, van den Eertwegh AJ, Lutzky J, et al. Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med. 2010; 363:711–23. https://doi.org/10.1056/NEJMoa1003466 [PubMed]
- 5. Schachter J, Ribas A, Long GV, Arance A, Grob JJ, Mortier L, Daud A, Carlino MS, McNeil C, Lotem M, Larkin J, Lorigan P, Neyns B, et al. Pembrolizumab versus ipilimumab for advanced melanoma: final overall survival results of a multicentre, randomised, open-label phase 3 study (KEYNOTE-006). Lancet. 2017; 390:1853–62. https://doi.org/10.1016/S0140-6736(17)31601-X [PubMed]
- 6. Larkin J, Hodi FS, Wolchok JD. Combined Nivolumab and Ipilimumab or Monotherapy in Untreated Melanoma. N Engl J Med. 2015; 373:1270–71. https://doi.org/10.1056/NEJMc1509660 [PubMed]
- 7. Felder M, Kapur A, Gonzalez-Bosquet J, Horibata S, Heintz J, Albrecht R, Fass L, Kaur J, Hu K, Shojaei H, Whelan RJ, Patankar MS. MUC16 (CA125): tumor biomarker to cancer therapy, a work in progress. Mol Cancer. 2014; 13:129. https://doi.org/10.1186/1476-4598-13-129 [PubMed]
- 8. Vuento MH, Stenman UH, Pirhonen JP, Mäkinen JI, Laippala PJ, Salmi TA. Significance of a single CA 125 assay combined with ultrasound in the early detection of ovarian and endometrial cancer. Gynecol Oncol. 1997; 64:141–46. https://doi.org/10.1006/gyno.1996.4545 [PubMed]
- 9. Chao A, Tang YH, Lai CH, Chang CJ, Chang SC, Wu TI, Hsueh S, Wang CJ, Chou HH, Chang TC. Potential of an age-stratified CA125 cut-off value to improve the prognostic classification of patients with endometrial cancer. Gynecol Oncol. 2013; 129:500–04. https://doi.org/10.1016/j.ygyno.2013.02.032 [PubMed]
- 10. Gubbels JA, Felder M, Horibata S, Belisle JA, Kapur A, Holden H, Petrie S, Migneault M, Rancourt C, Connor JP, Patankar MS. MUC16 provides immune protection by inhibiting synapse formation between NK and ovarian tumor cells. Mol Cancer. 2010; 9:11. https://doi.org/10.1186/1476-4598-9-11 [PubMed]
- 11. Belisle JA, Horibata S, Jennifer GA, Petrie S, Kapur A, André S, Gabius HJ, Rancourt C, Connor J, Paulson JC, Patankar MS. Identification of siglec-9 as the receptor for MUC16 on human NK cells, B cells, and monocytes. Mol Cancer. 2010; 9:118. https://doi.org/10.1186/1476-4598-9-118 [PubMed]
- 12. Fan K, Yang C, Fan Z, Huang Q, Zhang Y, Cheng H, Jin K, Lu Y, Wang Z, Luo G, Yu X, Liu C. MUC16 C terminal-induced secretion of tumor-derived IL-6 contributes to tumor-associated treg enrichment in pancreatic cancer. Cancer Lett. 2018; 418:167–75. https://doi.org/10.1016/j.canlet.2018.01.017 [PubMed]
- 13. Morgado M, Sutton MN, Simmons M, Warren CR, Lu Z, Constantinou PE, Liu J, Francis LL, Conlan RS, Bast RC
Jr , Carson DD. Tumor necrosis factor-α and interferon-γ stimulate MUC16 (CA125) expression in breast, endometrial and ovarian cancers through NFκB. Oncotarget. 2016; 7:14871–84. https://doi.org/10.18632/oncotarget.7652 [PubMed] - 14. Wu YM, Nowack DD, Omenn GS, Haab BB. Mucin glycosylation is altered by pro-inflammatory signaling in pancreatic-cancer cells. J Proteome Res. 2009; 8:1876–86. https://doi.org/10.1021/pr8008379 [PubMed]
- 15. Li X, Pasche B, Zhang W, Chen K. Association of MUC16 mutation with tumor mutation load and outcomes in patients with gastric cancer. JAMA Oncol. 2018; 4:1691–98. https://doi.org/10.1001/jamaoncol.2018.2805 [PubMed]
- 16. Havel JJ, Chowell D, Chan TA. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer. 2019; 19:133–50. https://doi.org/10.1038/s41568-019-0116-x [PubMed]
- 17. Luke JJ, Flaherty KT, Ribas A, Long GV. Targeted agents and immunotherapies: optimizing outcomes in melanoma. Nat Rev Clin Oncol. 2017; 14:463–82. https://doi.org/10.1038/nrclinonc.2017.43 [PubMed]
- 18. Ribas A, Lawrence D, Atkinson V, Agarwal S, Miller WH
Jr , Carlino MS, Fisher R, Long GV, Hodi FS, Tsoi J, Grasso CS, Mookerjee B, Zhao Q, et al. Combined BRAF and MEK inhibition with PD-1 blockade immunotherapy in BRAF-mutant melanoma. Nat Med. 2019; 25:936–40. https://doi.org/10.1038/s41591-019-0476-5 [PubMed] - 19. Chan TA, Wolchok JD, Snyder A. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2015; 373:1984. https://doi.org/10.1056/NEJMc1508163 [PubMed]
- 20. Carbone DP, Reck M, Paz-Ares L, Creelan B, Horn L, Steins M, Felip E, van den Heuvel MM, Ciuleanu TE, Badin F, Ready N, Hiltermann TJ, Nair S, et al, and CheckMate 026 Investigators. First-line nivolumab in stage IV or recurrent non-small-cell lung cancer. N Engl J Med. 2017; 376:2415–26. https://doi.org/10.1056/NEJMoa1613493 [PubMed]
- 21. Jia Q, Wang J, He N, He J, Zhu B. Titin mutation associated with responsiveness to checkpoint blockades in solid tumors. JCI Insight. 2019; 4:e127901. https://doi.org/10.1172/jci.insight.127901 [PubMed]
- 22. Wang F, Zhao Q, Wang YN, Jin Y, He MM, Liu ZX, Xu RH. Evaluation of POLE and POLD1 mutations as biomarkers for immunotherapy outcomes across multiple cancer types. JAMA Oncol. 2019; 5:1504–06. [Epub ahead of print]. https://doi.org/10.1001/jamaoncol.2019.2963 [PubMed]
- 23. Braun DA, Ishii Y, Walsh AM, Van Allen EM, Wu CJ, Shukla SA, Choueiri TK. Clinical validation of PBRM1 alterations as a marker of immune checkpoint inhibitor response in renal cell carcinoma. JAMA Oncol. 2019; 5:1631–33. [Epub ahead of print]. https://doi.org/10.1001/jamaoncol.2019.3158 [PubMed]
- 24. Garon EB, Rizvi NA, Hui R, Leighl N, Balmanoukian AS, Eder JP, Patnaik A, Aggarwal C, Gubens M, Horn L, Carcereny E, Ahn MJ, Felip E, et al, and KEYNOTE-001 Investigators. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med. 2015; 372:2018–28. https://doi.org/10.1056/NEJMoa1501824 [PubMed]
- 25. Chen J, Ye X, Pitmon E, Lu M, Wan J, Jellison ER, Adler AJ, Vella AT, Wang K. IL-17 inhibits CXCL9/10-mediated recruitment of CD8+ cytotoxic T cells and regulatory T cells to colorectal tumors. J Immunother Cancer. 2019; 7:324. https://doi.org/10.1186/s40425-019-0757-z [PubMed]
- 26. Ubil E, Caskey L, Holtzhausen A, Hunter D, Story C, Earp HS. Tumor-secreted Pros1 inhibits macrophage M1 polarization to reduce antitumor immune response. J Clin Invest. 2018; 128:2356–69. https://doi.org/10.1172/JCI97354 [PubMed]
- 27. Valeta-Magara A, Gadi A, Volta V, Walters B, Arju R, Giashuddin S, Zhong H, Schneider RJ. Inflammatory breast cancer promotes development of M2 tumor-associated macrophages and cancer mesenchymal cells through a complex chemokine network. Cancer Res. 2019; 79:3360–71. https://doi.org/10.1158/0008-5472.CAN-17-2158 [PubMed]
- 28. Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR, Albright A, Cheng JD, Kang SP, Shankaran V, Piha-Paul SA, Yearley J, Seiwert TY, et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest. 2017; 127:2930–40. https://doi.org/10.1172/JCI91190 [PubMed]
- 29. Conforti F, Pala L, Bagnardi V, De Pas T, Martinetti M, Viale G, Gelber RD, Goldhirsch A. Cancer immunotherapy efficacy and patients’ sex: a systematic review and meta-analysis. Lancet Oncol. 2018; 19:737–46. https://doi.org/10.1016/S1470-2045(18)30261-4 [PubMed]
- 30. Gupta S, Artomov M, Goggins W, Daly M, Tsao H. Gender disparity and mutation burden in metastatic melanoma. J Natl Cancer Inst. 2015; 107:djv221. https://doi.org/10.1093/jnci/djv221 [PubMed]
- 31. Liu D, Schilling B, Liu D, Sucker A, Livingstone E, Jerby-Amon L, Zimmer L, Gutzmer R, Satzger I, Loquai C, Grabbe S, Vokes N, Margolis CA, et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat Med. 2019; 25:1916–27. https://doi.org/10.1038/s41591-019-0654-5 [PubMed]
- 32. Latham A, Srinivasan P, Kemel Y, Shia J, Bandlamudi C, Mandelker D, Middha S, Hechtman J, Zehir A, Dubard-Gault M, Tran C, Stewart C, Sheehan M, et al. Microsatellite instability is associated with the presence of lynch syndrome pan-cancer. J Clin Oncol. 2019; 37:286–95. https://doi.org/10.1200/JCO.18.00283 [PubMed]
- 33. Wang H, Song M. Ckmeans.1d.dp: optimal k-means clustering in one dimension by dynamic programming. R J. 2011; 3:29–33. [PubMed]
- 34. Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015; 12:453–57. https://doi.org/10.1038/nmeth.3337 [PubMed]
- 35. Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013; 4:2612. https://doi.org/10.1038/ncomms3612 [PubMed]
- 36. Nagalla S, Chou JW, Willingham MC, Ruiz J, Vaughn JP, Dubey P, Lash TL, Hamilton-Dutoit SJ, Bergh J, Sotiriou C, Black MA, Miller LD. Interactions between immunity, proliferation and molecular subtype in breast cancer prognosis. Genome Biol. 2013; 14:R34. https://doi.org/10.1186/gb-2013-14-4-r34 [PubMed]
- 37. Dong ZY, Zhong WZ, Zhang XC, Su J, Xie Z, Liu SY, Tu HY, Chen HJ, Sun YL, Zhou Q, Yang JJ, Yang XN, Lin JX, et al. Potential predictive value of TP53 and KRAS mutation status for response to PD-1 blockade immunotherapy in lung adenocarcinoma. Clin Cancer Res. 2017; 23:3012–24. https://doi.org/10.1158/1078-0432.CCR-16-2554 [PubMed]
- 38. Rooney MS, Shukla SA, Wu CJ, Getz G, Hacohen N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell. 2015; 160:48–61. https://doi.org/10.1016/j.cell.2014.12.033 [PubMed]
- 39. Finkin S, Yuan D, Stein I, Taniguchi K, Weber A, Unger K, Browning JL, Goossens N, Nakagawa S, Gunasekaran G, Schwartz ME, Kobayashi M, Kumada H, et al. Ectopic lymphoid structures function as microniches for tumor progenitor cells in hepatocellular carcinoma. Nat Immunol. 2015; 16:1235–44. https://doi.org/10.1038/ni.3290 [PubMed]
- 40. Tsai KK, Zarzoso I, Daud AI. PD-1 and PD-L1 antibodies for melanoma. Hum Vaccin Immunother. 2014; 10:3111–16. https://doi.org/10.4161/21645515.2014.983409 [PubMed]
- 41. Van Allen EM, Miao D, Schilling B, Shukla SA, Blank C, Zimmer L, Sucker A, Hillen U, Foppen MH, Goldinger SM, Utikal J, Hassel JC, Weide B, et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science. 2015; 350:207–11. https://doi.org/10.1126/science.aad0095 [PubMed]
- 42. Long L, Zhang X, Chen F, Pan Q, Phiphatwatchara P, Zeng Y, Chen H. The promising immune checkpoint LAG-3: from tumor microenvironment to cancer immunotherapy. Genes Cancer. 2018; 9:176–89. https://doi.org/10.18632/genesandcancer.180 [PubMed]
- 43. Das M, Zhu C, Kuchroo VK. Tim-3 and its role in regulating anti-tumor immunity. Immunol Rev. 2017; 276:97–111. https://doi.org/10.1111/imr.12520 [PubMed]
- 44. Dougall WC, Kurtulus S, Smyth MJ, Anderson AC. TIGIT and CD96: new checkpoint receptor targets for cancer immunotherapy. Immunol Rev. 2017; 276:112–20. https://doi.org/10.1111/imr.12518 [PubMed]
- 45. Blair AB, Kleponis J, Thomas DL
2nd , Muth ST, Murphy AG, Kim V, Zheng L. IDO1 inhibition potentiates vaccine-induced immunity against pancreatic adenocarcinoma. J Clin Invest. 2019; 129:1742–55. https://doi.org/10.1172/JCI124077 [PubMed] - 46. Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics. 2013; 14:7. https://doi.org/10.1186/1471-2105-14-7 [PubMed]
- 47. Skidmore ZL, Wagner AH, Lesurf R, Campbell KM, Kunisaki J, Griffith OL, Griffith M. GenVisR: genomic visualizations in R. Bioinformatics. 2016; 32:3012–14. https://doi.org/10.1093/bioinformatics/btw325 [PubMed]



