Introduction
Age is the number one risk factor for several human health issues, those referred to as age-related diseases [1, 2]. These age-related diseases grow in importance as the median age of the world’s population continues to increase [3, 4]. As a result, the molecular mechanisms influencing aging have long been a topic of interest for study [5–7]. Recent developments in the study of the human metabolome have allowed for wide-spread study of changes in the metabolome in association with aging and age-related disease [1, 8, 9].
Acylcarnitines are carrier forms of fatty acids required for import of long-chain (LC) fatty acids into mitochondria for β-oxidation to occur [10]. The most common reported changes is an increase in blood concentration of LC acylcarnitines in individuals with age-related diseases [11–14]. Dysregulation of acylcarnitine homeostasis has been tied to a variety of age-related diseases, including cardiovascular disease [11, 12, 15], type II diabetes mellitus [13, 16, 17], osteoarthritis [18], chronic obstructive pulmonary disease [19], macular degeneration [14], glaucoma [20] and Alzheimer’s disease [21–23]. In addition to the association with age-related diseases, abnormal acylcarnitine levels are associated with activation of inflammation [24] and mitochondrial dysfunction [25, 26]. Loss of mitochondrial function both contributes to the process of aging and is, itself, an indirect result of aging. Mitochondrial dysfunction has been documented to play a role in development of most age-related diseases; however, it occurs independently of disease [27–29]. Additionally, healthy older individuals require more time to reestablish acylcarnitine homeostasis after stimulation with insulin, and higher acylcarnitine levels are found in aged men with reduced physical ability when compared with similarly aged men with normal physical ability [30, 31]. Gender differences in acylcarnitine associations with age have not been well studied, but some differences between men and women have been reported [32, 33]. Additionally, gender differences in lipid metabolites related to lifespan have been reported [34].
Neither LC acyl-CoA nor free LC fatty acids can migrate across the inner membrane of the mitochondria, so the formation of acylcarnitines is critical to metabolism of LC fatty acids [35]. This system, termed the carnitine shuttle [see Reuter & Evans [25] for review], normally maintains carnitine and acylcarnitine within a narrow range [35]. This ensures normal functioning of fatty acid β-oxidation as well as adequate availability of CoA. Acylcarnitines are also transported into plasma [25], and as a consequence, plasma acylcarnitine levels can serve as an indicator of mitochondrial function [36].
Despite the known relationship between acylcarnitine homeostasis and mitochondrial function, investigations of acylcarnitines have focused primarily on diseased populations, and little attention has been given to differences in acylcarnitine homeostasis between healthy men and women. The current study was designed to test whether acylcarnitines vary by age and gender in a healthy, non-diseased population. Results from high-resolution metabolomics (HRM) analyses show that LC and very long-chain (VLC) acylcarnitines increase with age and have gender-specific differences in healthy individuals.
Results
Study population demographics
All 78 female adults and 85 male adults were healthy, without history of smoking, known disease or metabolic risk factors for disease (Figure 1A). The mean age was 43.5 years, ranging from 20 to 90 y (Figure 1B). The population included multiple races and ethnicities, but the population size was too small for separate analyses. There was no difference in age distribution of males and females.
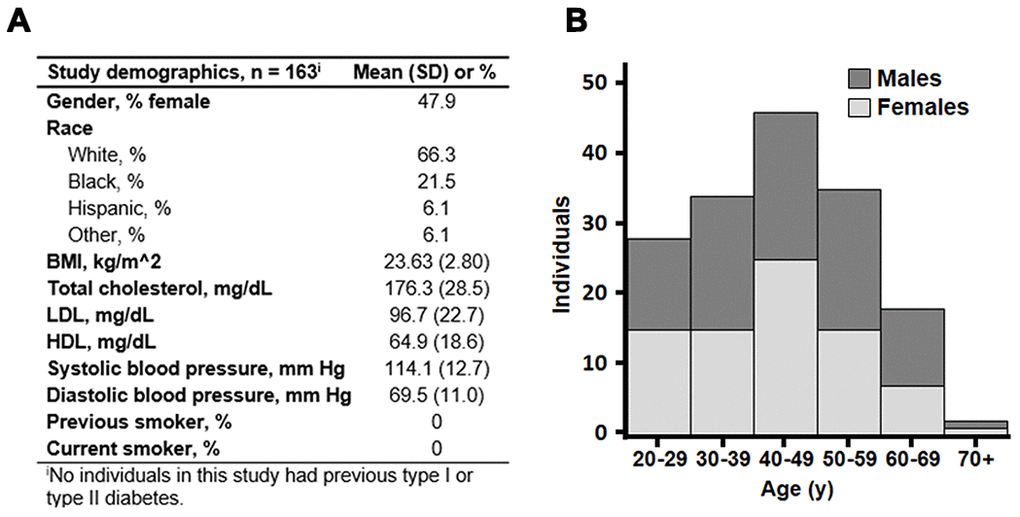
Figure 1. Background characteristics of 163 healthy adults of the predictive medicine cohort. (A) Mean values with standard deviation (SD) of gender, race and clinical measures are shown. (B) Age distribution of the subset. Stacked bars are shown with men in dark gray and women in light gray. Mean age was 43.5 years, and ages ranged from 20 to 90 years.
Metabolome-wide association study (MWAS) with age
After filtering for m/z features present in at least 80% of samples, 26045 features (hereafter referred as “metabolic features” or “metabolites”) were detected by C18 and anion exchange (AE) columns, with 25680 unique m/z within 10 ppm. MWAS (p < 0.05) showed that 1915 features associated with age (154 features at FDR <0.2), with 986 associating positively and 929 associating negatively (Figure 2A). In the current study, we retained all features with p < 0.05 for annotation of possible acylcarnitines and subsequent analysis. Full listings of these are provided (C18, Supplementary Table 1; AE, Supplementary Table 2) and respective annotations are given (Supplementary Tables 3 and 4).
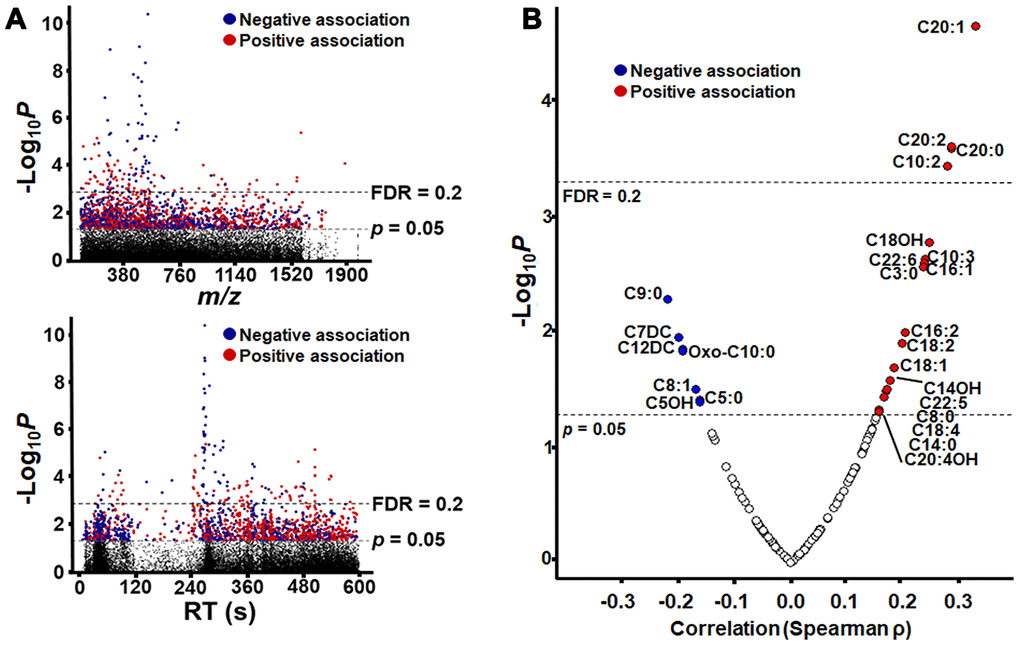
Figure 2. Metabolome-Wide Association Study (MWAS) of plasma metabolites correlated with age. (A) Type 1 Manhattan plot showing -log10p for correlation of each metabolite plotted by m/z (mass-to-charge ratio) and type 2 Manhattan plot showing -log10p for correlation of each metabolite plotted by chromatographic retention time (RT) in seconds, as separated the C18 column. Plots are shown with significance (n = 1505, p = 0.05) and false discovery rate (n= 140, FDR = 0.2) thresholds by dashed lines, and the detailed information of metabolic features is provided in Supplementary Table 1. (B) Plot of acylcarnitine correlation strength and direction (Spearman ρ) by –Log10 p. Acylcarnitines with p < 0.05 are labeled by the chain length, saturation and modification of the acyl group (see Table 1 for details). For acylcarnitines detected on both C18 and anion exchange columns, only the C18 data is represented in the plot. The plot is shown with significance (n = 26, p = 0.05) and false discovery rate (n= 4, FDR = 0.2) thresholds by dashed lines In all plots, significant negative correlations are shown in blue, and significant positive correlations are shown in red.
Table 1. Correlation of acylcarnitines with age.
| m/z | RT (s) | SLC | Database | Common name | Median intensity | Correlation (ρ) | p-value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| C18 positive | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 162.1116 | 42 | 2b | HMDB00062 | Carnitine (C0) | 3.60E6 | -0.0437 | 0.5797 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 454.3869 | 470 | 3 | LMFA07070010 | Eicosenoylcarnitine (C20:1) | 2.60E5 | 0.3249 | 2.32E-5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 452.3712 | 391 | 3 | LMFA07070011 | Eicosadienoylcarnitine (C20:2) | 1.78E5 | 0.2827 | 2.56E-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 456.4025 | 465 | 3 | HMDB06460 | Arachidyl carnitine (C20:0) | 9.27E4 | 0.282 | 2.65E-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 312.2154 | 93 | 3 | HMDB13325 | Decadienoylcarnitine (C10:2) | 3.01E5 | 0.2754 | 3.73E-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 444.3665 | 348 | 3 | LMFA07070028 | Hydroxystearoylcarnitine (C18OH) | 7.27E4 | 0.2439 | 1.70E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 310.1998 | 76 | 3 | LMFA07070016 | Decatrienoylcarnitine (C10:3)* | 1.03E6 | 0.2363 | 2.39E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 472.3387 | 335 | 3 | HMDB06510 | Docosahexaenoylcarnitine (C22:6) | 2.42E4 | 0.2344 | 2.60E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 398.3245 | 334 | 3 | HMDB06317 | Hexadecenoylcarnitine (C16:1)* | 6.73E5 | 0.2341 | 2.63E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 218.1375 | 258 | 2b | HMDB00824 | Propionylcarnitine (C3:0) | 5.14E4 | 0.2327 | 2.79E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 396.3088 | 313 | 3 | HMDB13334 | Hexadecadienoylcarnitine (C16:2)* | 2.30E5 | 0.2005 | 0.0103 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 422.3244 | 323 | 3 | HMDB06318 | Octadecatrienoylcarnitine (C18:3) | 2.19E5 | 0.1947 | 0.0128 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 388.3037 | 286 | 3 | HMDB13166 | Hydroxymyristoylcarnitine (C14OH) | 1.21E5 | 0.1735 | 0.0268 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 474.3564 | 347 | 3 | HMDB06321 | Docosapentaenoylcarnitine (C22:5) | 4.26E4 | 0.1683 | 0.0318 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 420.3089 | 314 | 3 | HMDB06463 | Octadecatetraenoylcarnitine (C18:4) | 2.62E4 | 0.1553 | 0.0478 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 464.3352 | 309 | 3 | LMFA07070036 | Hydroxyicosatetraenoylcarnitine (C20:4OH) | 4.94E3 | 0.1539 | 0.0498 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 262.1637 | 258 | 2b | LMFA07070041 | Hydroxyvalerylcarnitine (C5OH) | 1.51E4 | -0.1602 | 0.0411 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 246.1687 | 265 | 2b | HMDB13128 | Valerylcarnitine (C5:0)* | 5.07E4 | -0.1612 | 0.0398 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 374.2519 | 311 | 3 | HMDB13327 | Dodecanedioylcarnitine (C12DC) | 8.53E4 | -0.1914 | 0.0144 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 304.1741 | 257 | 3 | HMDB13328 | Pimelylcarnitine (C7DC) | 2.97E4 | -0.1976 | 0.0114 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 302.231 | 301 | 3 | HMDB13288 | Nonanoylcarnitine (C9:0) | 6.07E4 | -0.2174 | 5.30E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Anion exchange | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 310.2014 | 57 | 3 | LMFA07070016 | Decatrienoylcarnitine (C10:3)* | 3.30E5 | 0.2421 | 1.85E-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 398.3265 | 76 | 3 | HMDB13207 | Hexadecenoylcarnitine (C16:1)* | 3.68E5 | 0.1945 | 0.0128 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 426.3576 | 111 | 3 | HMDB06351 | Octadecenoylcarnitine (C18:1) | 2.55E6 | 0.1808 | 0.0209 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 396.3108 | 65 | 3 | HMDB13334 | Hexadecadienoylcarnitine (C16:2)* | 1.21E5 | 0.1723 | 0.0279 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 369.2828 | 59 | 3 | HMDB13331 | Tetradecadienoylcarnitine (C14:2) | 1.20E5 | 0.1682 | 0.0318 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 288.2169 | 63 | 2b | HMDB00791 | Octanoylcarnitine (C8:0) | 5.17E5 | 0.1632 | 0.0374 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 372.3109 | 71 | 3 | HMDB05066 | Myristoylcarnitine (C14:0) | 2.51E5 | 0.1547 | 0.0486 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 286.2013 | 570 | 2b | HMDB13324 | Octenoylcarnitine (C8:1) | 2.72E4 | -0.1681 | 0.032 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 246.1698 | 581 | 2b | HMDB13128 | Valerylcarnitine (C5:0)* | 7.13E3 | -0.1778 | 0.0232 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 330.2272 | 584 | 3 | HMDB13202 | Ketodecanoylcarnitine (Oxo-C10) | 2.08E3 | -0.1905 | 0.0149 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Correlation analyzed by Spearman (ρ) and significance of correlation (p-value) are shown for each acylcarnitine detected on both C18 and anion exchange columns. Mass-to-charge ratio (m/z), retention time (RT), Schymanski level of confidence (SLC), and Human Metabolome Database (HMDB) or Lipid Maps (LMFA) database identifier are given for each acylcarnitine. Although no significant correlation was found for free carnitine (C0), values are given for reference. A SLC of 3 indicates a putative identification using LCMS data, whereas a SLC of 2b indicates a probable match using additional diagnostic MS/MS data. All acylcarnitines were detected as an M + H adduct. *, acylcarnitine identified by both C18 and anion exchange. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Annotation of features with accurate mass match to M + H adducts of acylcarnitines with xMSannotator [37] resulted in 132 annotated acylcarnitines, 30 of which displayed a significant association with age (P<0.05, Table 1). For some of the higher abundance features, MS/MS spectra showed nominal mass fragments at 85 and 144, characteristic of acylcarnitines, and the features are therefore discussed as the corresponding acylcarnitines with accurate mass match (see also Figure 5, below). Among the m/z features matching acylcarnitines, 4 were identified by both C18 and AE columns, resulting in 26 uniquely annotated acylcarnitines that varied with age. Nineteen of these correlated positively with age, and 7 correlated negatively with age (Figure 2B). Carnitine (m/z 162.1116, 42 s) did not vary with age (p = 0.58).
Acylcarnitine associations with age
The 26 acylcarnitines found to associate with age in plasma were compared for similarity between individuals through one-way hierarchical clustering analysis (HCA), with the study population arranged by age (Figure 3). Results showed LC and VLC acylcarnitines (≥ 16 carbon chain) [38] (Figure 3) clustered together in a major lower cluster and several medium-chain (MC) and short-chain (SC) acylcarnitines clustered into an upper cluster (see also Figure 2B).
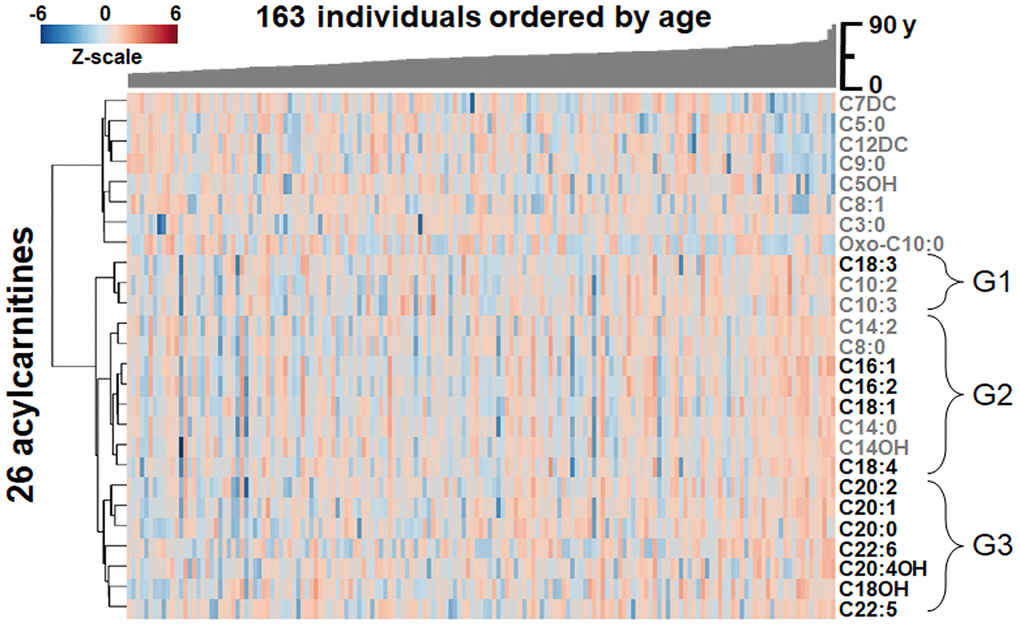
Figure 3. Heat map of one-way hierarchical clustering analysis (HCA) of the 26 acylcarnitines significantly associating with age. Along the x-axis, individuals are organized by age, with youngest on the left. The y-axis is comprised of the one-way HCA of acylcarnitines. Each column represents an individuals’ metabolic profile of the 35 acylcarnitines. Degree of deviation of acylcarnitine concentration below the mean of the study population are indicated by saturation of blue coloration, and degree of deviation of acylcarnitine concentration above the mean of the study population are indicated by saturation of red coloration. Short-chain and medium-chain acylcarnitines are labeled in gray, and long-chain and very-long-chain acylcarnitines are highlighted by labeling in black. For acylcarnitines detected on both C18 and anion exchange columns, only the C18 data was included in the HCA. The lower major acylcarnitine cluster is labeled by its subgroups, G1-3.
The lower cluster of 12 LC and VLC acylcarnitines contained three subclusters, labeled G1-G3 in Figure 3. G1 contained C18:3 as well as C10:2 and C10:3 (Figure 3). G2, the largest of the three, contained 8 acylcarnitines with 14- and 16-carbon acyl groups as well as C8:0, C18:4 and C18:1 (Figure 3). G3 contained 7 acylcarnitines with 20- and 22-carbon acyl groups as well as C18OH (Figure 3). In all three subgroups, abundances of LC and VLC acylcarnitines were increased with age.
The top three acylcarnitines by strength of association were present in G3 and each had 20-carbon acyl groups: eicosenoylcarnitine (C20:1; ρ = 0.3249, p < 0.0001; Figure 4A), eicosadienoylcarnitine (C20:2; ρ = 0.2827, p = 0.0003; Figure 4B) and arachidylcarnitine (C20:0; ρ = 0.2820, p = 0.0003; Figure 4C). The following three acylcarnitines by strength of association were present in G2 and included decadienoylcarnitine (C10:2; ρ = 0.2754, p = 0.0004; Figure 4D), hydroystearoylcarnitine (C18OH; ρ = 0.2439, p = 0.0017; Figure 4E) and decatrienoylcarnitine (C10:3; ρ = 0.2421, p = 0.0019; Figure 4F). The relative intensity values (Table 1) showed that the most highly associated acylcarnitines had relatively high abundances compared to other acylcarnitines associated with age.
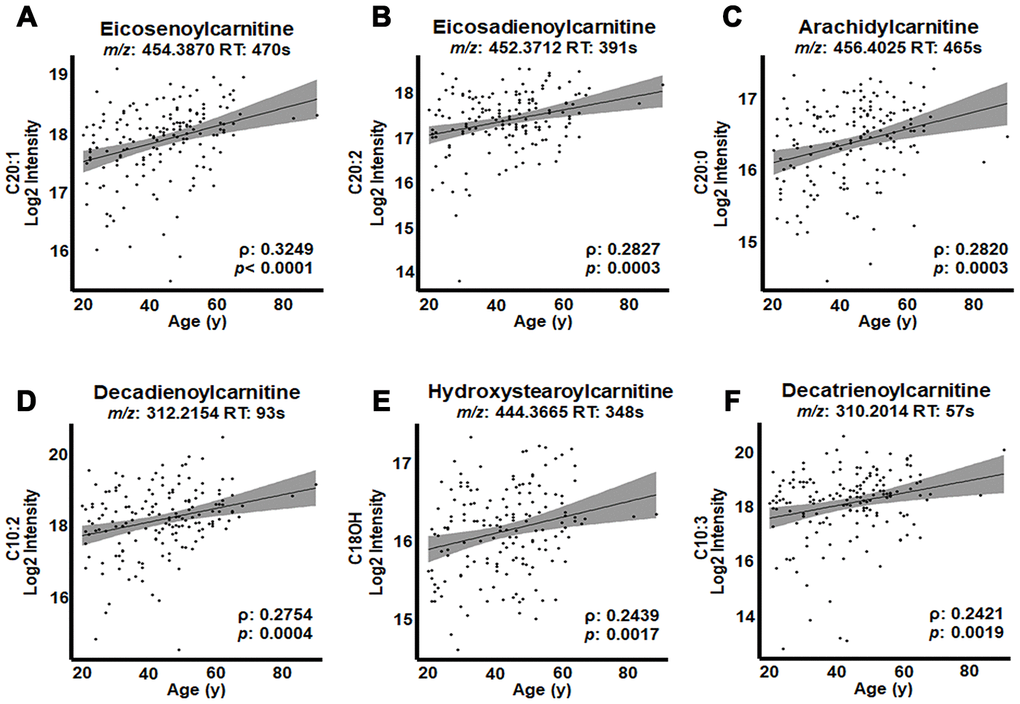
Figure 4. Highest correlations of acylcarnitines with age in human plasma. Log2 transformed intensity values for ions, identified by mass-to-charge ratio (m/z) and retention time (RT) for individual plasma samples are plotted against individual ages. Confidence intervals (95%) are shown in gray. (A) Eicosenoylcarnitine (C20:1), (B) eicosadienoylcarnitine (C20:2), (C) arachidylcarnitine (C20:0) and (D) decadienoylcarnitine (C10:2) were significant at FDR = 0.2. (E) Hydroxystearoylcarnitine (C18OH) and (F) decatrienoylcarnitine (C10:3) were significant at p < 0.05.
In contrast to the pattern of the LC and VLC acylcarnitines, which increased with age, the MC and SC cluster decreased in association with age (Figures 2B, 3). These were all relatively low abundance signals (Table 1). The MC and SC cluster included odd-chain (OC; C5, C7, C9) acylcarnitines, all of which decreased in association with age (Figure 3). The MC and SC cluster also contained features matching acylcarnitine derivatives of two dicarboxylic acids (DC) (Figures 2B, 3). Propionylcarnitine (C3:0) was present in the MC and SC cluster but did not decrease with age unlike the other OC acylcarnitines (see also Figure 2B). MS/MS fragmentation supported the identity of valerylcarnitine (C5:0; Figure 5A), hydroxyvalerylcarnitine (C5OH; Figure 5B), octenoylcarnitine (C8:1; Figure 5C) and octanoylcarnitine (C8:0; Figure 5D) relative to database spectra for methylbutyroylcarnitine. C5:0 matched several fragments for methylbutyroylcarnitine (Figure 5A). C5OH, C8:1 and C8:0 matched with methylbutyroylcarnitine, with the exception of one major fragment. The mass difference for fragments matched the mass differences present between the given acylcarnitine and methylbutyroylcarnitine (Figure 5B–5D).
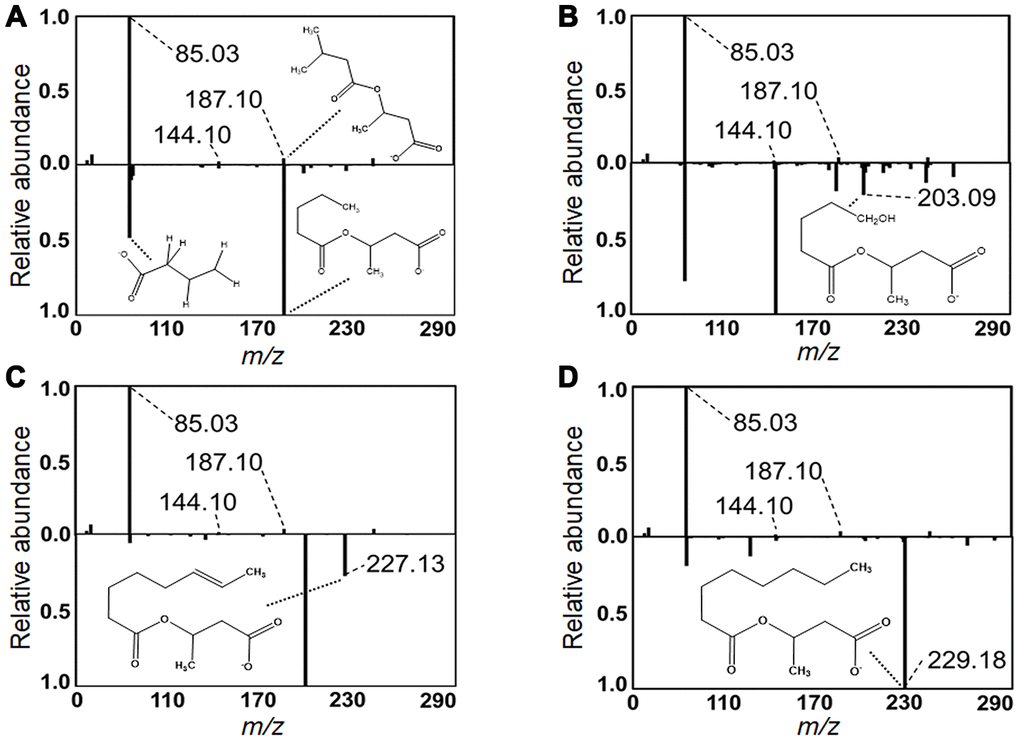
Figure 5. Identification of acylcarnitines by MS/MS. Experimental MS/MS fragmentations of (A) valerylcarnitine, (B) hydroxyvalerylcarnitine, (C) octenoylcarnitine and (D) octanoylcarnitine are juxtaposed below a library MS/MS fragmentation of methylbutyroylcarnitine. Diagnostic fragments common between library and experimental fragmentations are labeled. Pertinent MS/MS peaks are labeled for mass-to-charge ratio (m/z) by broken line. Additionally, distinctive fragments equivalent in mass difference to that of the mass difference between the represented acylcarnitine and methylbutyroylcarnitine are labeled with proposed fragment structure displayed. MS/MS peaks and matching proposed fragment structure are labeled by dotted line.
Tests for correlations among the acylcarnitines in Figure 3 showed that the high-abundance LC and VLC metabolites in G3 were highly positively correlated with each other, with the one exception of C22:6, which did not correlate with any other LC or VLC acylcarnitines (Supplementary Table 5). In contrast, these LC and VLC acylcarnitines were not correlated, either positively or negatively, with carnitine or acetylcarnitine. Propionylcarnitine was associated with arachidylcarnitine C20:0 (ρ = 0.2968, p = 0.0001). Few associations occurred among MC and SC acylcarnitines (Supplementary Table 5). Acylcarnitines with a 10-carbon acyl group as well as C14OH had associations with LC and VLC acylcarnitines (Supplementary Table 5). OC acylcarnitines had few correlations; only C9:0 correlated with C5:0 (ρ = 0.3561, p < 0.0001) and C7DC (ρ = 0.3583, p < 0.0001; Supplementary Table 5). Hydroxyvalerylcarnitine (C5OH) showed no correlation with any other OC acylcarnitines.
Gender differences in acylcarnitine associations with age
The 6 acylcarnitines with greatest positive rho value were examined for their associations with age when individuals were separated by gender. This separation by gender did not show stronger correlation of acylcarnitines with age (Table 2) than observed for the combined analyses (Fig 5). Comparisons of genders show that C20:1 and C20:2 exhibited stronger correlations with age in males than females while C20:0, C10:2, C18OH and C10:3 exhibited slightly stronger correlations with age in females than males.
Table 2. Comparison of correlations of acylcarnitines with age by gender.
| Name | Structure | Female (ρ, p) | Male (ρ, p) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Eicosenoylcarnitine | C20:1 | 0.2828, 0.0121 | 0.3551, 8.55E-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Eicosadienoylcarnitine | C20:2 | 0.1344, 0.2409 | 0.3693, 5.04E-4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Arachidylcarnitine | C20:0 | 0.3078, 6.11E-3 | 0.2397, 0.0271 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decadienoylcarnitine | C10:2 | 0.3194, 4.37E-3 | 0.2323, 0.0324 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hydroxystearoylcarnitine | C18OH | 0.3455, 2.95E-3 | 0.1457, 0.1845 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decatrienoylcarnitine | C10:3 | 0.2456, 0.0330 | 0.2304, 0.0365 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| For the 6 acylcarnitines with the highest correlation with age in the whole population, correlation with age by gender was tested. Spearman’s ρ and p-value are shown for correlations in females and males. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
The xMWAS analysis of these 6 acylcarnitines against the remainder of the metabolome in each gender resulted in different clustering within each network (Fig 6). The female network grouped into 5 distinct clusters, with separate clusters for C18OH, C20:0, C20:1 and C20:2 (Clusters 1, 2, 4 and 5, respectively), and C10:2 and C10:3 (Cluster 3) were consolidated into one cluster of metabolites (Figure 6A). Relative to females, the male network grouped more tightly resulting in 4 clusters, with C20:1 and C20:2 in Cluster 4 (Figure 6B). Additionally, the three 20 carbon acylcarnitines showed associations with more metabolites in the males compared to females.
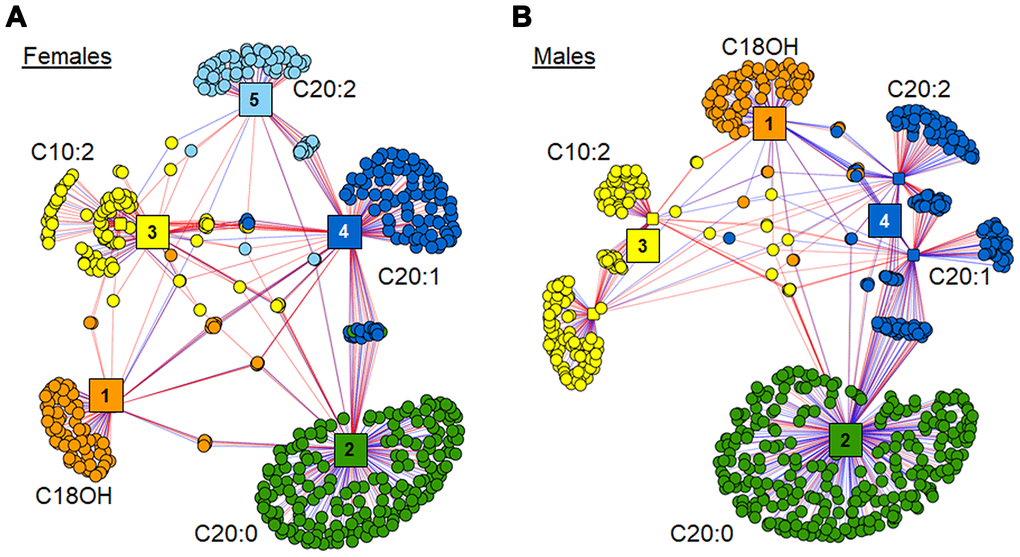
Figure 6. Association of top 6 age-associated acylcarnitines with metabolome. (A) xMWAS network of top 6 age-associated acylcarnitines as associated with metabolome within females. Cluster 1 (orange) has features predominantly associated with hydroxystearoylcarnitine (C18OH). Cluster 2 (green) is comprised of features associated most closely with arachidylcarnitine (C20:0). Cluster 3 (yellow) contains features clustered around decadienoylcarnitine (C10:2) and decatrienoylcarnitine (C10:3). Cluster 4 (dark blue) has features mainly associated with eicosenoylcarnitine (C20:1). Cluster 5 (light blue) has features mainly associated with eicosadienoylcarnitine (C20:2). See Supplementary Table 6 for detailed annotation of metabolites included in the female network. (B) xMWAS network of top 6 age-associated acylcarnitines as associated with metabolome within males. Clusters 1-3 form around the same acylcarnitines as their respective acylcarnitines in the female subset. Cluster 4 (blue) has features mainly associated with both C20:1 and C20:2. See Supplementary Table 7 for detailed annotation of metabolites included in the male network. Positive associations are shown in red, while negative associations are shown with blue lines.
Metabolites associated with these 6 acylcarnitines shared little commonality between females and males, with less than 11% of those metabolites included in the female network also being present in the male network. Annotations of the features in the female and male xMWAS networks are provided in Supplementary Tables 6 and 7, respectively. Pathway enrichment analyses of the metabolic features associated in both the female and male xMWAS networks resulted in distinctly different pathways associating with age-associated acylcarnitines in women and in men. Cytochrome P450 metabolism (females: p = 0.001; males: p = 0.030) and vitamin E metabolism (females: p = 0.006; males: p = 0.001) changed in association with age-associated acylcarnitines in both genders (Table 3). In females, glycerophospholipid metabolism (p = 0.004), leukotriene metabolism (p = 0.004), fatty acid activation (p = 0.017), glycosphingolipid metabolism (p = 0.020), prostaglandin formation (p = 0.028), tryptophan metabolism (p = 0.032) and fatty acid biosynthesis (p = 0.039) changed in association with age-associated acylcarnitines (Table 3). In males, the carnitine shuttle (p = 0.002), cholesterol biosynthesis (p = 0.011) and steroid biosynthesis (p = 0.030) changed in association with age-associated acylcarnitines (Table 3).
Table 3. Metabolic pathways associated with top 6 age-related acylcarnitines by gender.
| Metabolic Pathway | Number of metabolites at p < 0.05 | Number of metabolites from pathway detected | p-value |
| Females | |||
| Cytochrome P450 drug metabolism | 5 | 26 | 0.001 |
| Glycerophospholipid metabolism | 4 | 28 | 0.004 |
| Leukotriene metabolism | 6 | 51 | 0.004 |
| Vit E metabolism | 4 | 30 | 0.006 |
| Fatty acid activation | 3 | 24 | 0.017 |
| Glycosphingolipid metabolism | 3 | 25 | 0.020 |
| Prostaglandin formation | 4 | 40 | 0.028 |
| Tryptophan metabolism | 4 | 41 | 0.032 |
| Fatty acid biosynthesis | 3 | 29 | 0.039 |
| Males | |||
| Vit E metabolism | 6 | 30 | 0.001 |
| Carnitine shuttle | 5 | 28 | 0.002 |
| Cholesterol biosynthesis | 4 | 31 | 0.011 |
| Steroid biosynthesis | 5 | 53 | 0.030 |
| Cytochrome P450 drug metabolism | 3 | 25 | 0.030 |
Discussion
Our results show that plasma LC and VLC acylcarnitines increase with age in healthy individuals. In individuals with no age-related disease nor metabolic risk factors, all age-associated acylcarnitines with acyl groups of carbon chains 16 carbons or longer increased in association with age. All OC acylcarnitines except for C3:0 were decreased with age. The results also show that among the most strongly age-associated acylcarnitines, C20:2 and C18OH show difference in strength of association between genders, and the genders differ considerably in lipid pathways which vary in association with age-associated acylcarnitines.
Changes in acylcarnitines with age alone point toward decreased mitochondrial function with age. Elevated acylcarnitines, especially LC and VLC acylcarnitines, serve as markers of mitochondrial deficiency in fatty acid oxidation [25, 26]. Downstream associations of increased LC acylcarnitines with other lipid metabolic pathways, such as glycerophospholipid, glycosphingolipid, fatty acid and cholesterol metabolism, further suggests that the effect of age occurs through changes in mitochondrial function [39]. Inverse association of hydroxyvalerylcarnitine (C5OH) with age as well as the lack of association of C5OH with any other age-related OC acylcarnitines suggests changes in other mitochondrial pathways such as branched chain amino acid (BCAA) metabolism [40, 41]. Negative association of dicarboxylic acylcarnitines (C7DC and C12DC) with age suggests decreased β-oxidation of LC dicarboxylic acylcarnitines in the peroxisome with age [42]. Increased LC and VLC with age and dysregulation of BCAA metabolism and mitochondrial and peroxisomal lipid metabolism are consistent with the effect of age being upstream of the mitochondria. Such upstream effects on mitochondrial lipid and amino acid metabolism and peroxisomal activity are known to occur through changes in mTOR/PPARα activity with age [43, 44].
The increase in acylcarnitines in healthy individuals with increased age alone suggests a need for improved matching of case and control groups in studies of age-related disease. For instance, in some studies on increased acylcarnitines in age-related disease, case groups were older than controls [12–14, 23]. Other studies used populations in which there was no significant difference in age of cases and controls, but the case group mean age was over 5 years greater than that of the control group [16, 19, 22]. The consistency of these studies with regard to associations of acylcarnitines and disease supports the correctness of the authors conclusions; none-the-less, the magnitude of effects might be impacted by non-disease-related, age-associated changes of acylcarnitines in the controls.
Two LC acylcarnitines, C16:1 (hexadecenoylcarnitine) and C18:1 (octadecenoylcarnitine), positively correlated with age in the present study, have been reported to be positively associated with age-related disease in some of these studies. Bouchouirab et al. [13] reported decreased plasma clearance of C18:1 in response to insulin or postprandially in individuals with type II diabetes when compared to controls who were non-diabetic with no family history of diabetes. In this study, the mean age of individuals with type II diabetes was nearly twice that of the controls group. Similarly, Adams et al. [16] reported increased fasting C18:1 in obese, African American women with type II diabetes compared to that of obese, African American women without disease. In this study, the mean age of diabetic individuals was 5 years older than that of controls, and the maximum age studied for the diabetic group was 18 years greater than that of the control group. Lastly, a recent study of neovascular age-related macular degeneration [14] reported increased C16:1 and C18:1 in diseased individuals compared to controls. Their study controlled for age in the analysis, and the mean age for both groups was above 70 years old. Our present study mostly had individuals 20-70 years of age, and this limits conclusions which may be drawn from comparisons.
The difference in direction of association of C3:0 compared to all other OC acylcarnitines may be a result of decreased flux with age because propionyl-CoA is a product of OC fatty acid β-oxidation [45]. Increased C3:0 with age may be indicative of an age-related change in metabolism of OC LC fatty acids, which have been shown to associate negatively with heart disease and type II diabetes [46, 47]; however, we observed no association of OC LC acylcarnitines with age. Alternatively, the decrease in C5:0, C7DC and C9:0 with age may be a result of decreased dairy consumption with age, as dairy products are an important source of OC fatty acids in the diet [48, 49]. This increase in C3:0 with age could also be due to age-related differences in composition of the microbiome or in dietary fiber intake [50]; these parameters were not assessed in our study population. Future study of metabolic flux using stable isotope-labeled OC fatty acids could shed light on whether this observation is due to an age-related change in metabolism or an external factor such as dietary fiber or dairy intake.
As with previously noted differences in acylcarnitine association with age in men and women, we also found differences in strength of association of several acylcarnitines with age between genders. Previously, Muilwijk et al. [33] reported an increase in concentrations of several acylcarnitines with age in individuals who were without age-related disease but whose clinical measures were not controlled. They noted that increases observed in acylcarnitines with age were higher in women, and several more significant changes in acylcarnitines with age were reported for women than were for men. Yu et al. [32] reported C18:1 as a correlate with age in both men and women; however, in women the relationship was much stronger. In neither of these studies were the differences in strengths of correlation a focus. Similarly, our data show that there is difference between the genders in how strongly acylcarnitines associate with age. Our study shows different associations of acylcarnitines with the remainder of metabolism. Generally, women exhibited more lipid metabolic pathways which were altered in association with age-associated acylcarnitines. In addition to this, women exhibited changes in leukotriene metabolism, prostaglandin formation and tryptophan metabolism in association with age-associated acylcarnitines. Association of these inflammatory-related pathways with age-associated acylcarnitines in women but not in men support previous findings that gender differences in inflammatory pathways tend to magnify in old age [51–54].
The present study used stringent selection criteria for health of individuals studied; however, the population was small and cannot be considered representative of the general population, especially for racial and ethnic comparisons. Additionally, this was a cross-sectional study of individuals at different ages and does not address longitudinal changes within individuals as a result of aging. Studies are needed to evaluate changes in acylcarnitines which occur as a result of loss of mitochondrial function and other physiological measures of aging, such as telomeric shortening, loss of proteostasis, deregulated nutrient-sensing or physical frailty [55, 56].
In summary, the present results show that abundant acylcarnitines increase in plasma with age in healthy individuals. The results emphasize the importance of strict control for age in metabolomic studies of age-related diseases in order to account for metabolic alterations which occur as an adaptive response in healthy aging regardless of disease state. The study also shows important gender differences in glycerolipids and other metabolic networks linked to acylcarnitines that vary by age. Thus, the results provide justification for detailed studies of lipid metabolism in aging, specifically to understand gender differences which could impact underlying gender-specific disease mechanisms.
Materials and Methods
Chemicals
HPLC grade acetonitrile and methanol, LC-MS water and 98% formic acid were obtained from Sigma-Aldrich (St. Louis, MO). A mixture of 14 stable isotopic chemicals used as an internal standard [57] included [13C6]-D-glucose, [15N]-indole, [2-15N]-L-lysine dihydrochloride, [13C5]-L-glutamic acid, [13C7]-benzoic acid, [3,4-13C2]-cholesterol, [15N]-L-tyrosine, [trimethyl-13C3]-caffeine, [15N2]-uracil, [3,3-13C2]-cystine, [1,2-13C2]-palmitic acid, [15N,13C5]-L-methionine, [15N]-choline chloride and 2’-deoxyguanosine-15N2,13C10-5’-monophosphate from Cambridge Isotope Laboratories, Inc (Andover, PA).
Human plasma samples
A subset of samples (n = 163) from the Predictive Medicine Research (PREMED) cohort (ClinicalTrials.gov Identifier: NCT00336570) was used; the subset represented all available samples and did not appear to have selection bias. The study was reviewed and approved by the Emory University Investigational Review Board (IRB00024767). PREMED subjects were healthy individuals between 20 and 90 years of age. Participants were originally studied to define a “normal” clinical value or range of values for plasma contents in healthy individuals to evaluate methods for detecting early multiorgan disease (NCT00336570). Included participants possessed low BMI, LDL cholesterol and blood pressure as well as no history of smoking. Participants were excluded by use of lipid lowering medication, presence of any number of chronic or acute diseases or disorders, listed in full detail at ClinicalTrials.gov. Blood plasma was collected with EDTA, and samples were stored at -80°C prior to LC-MS analysis.
High-resolution metabolomics (HRM)
Plasma samples were analyzed as described previously [58]. Briefly, 50 μL plasma samples were treated 2:1 (v/v) with acetonitrile, and 2.5 μL internal standard of a mixture of 14 stable isotope standards was added. Proteins were precipitated and pelleted by incubation at 4°C for 30 min followed by centrifugation for 10 min at 21000 x g at 4°C. Supernatants were placed in autosampler vials and maintained at 4°C in an autosampler. Samples were analyzed by liquid chromatography-Fourier transform mass spectrometry at 60,000 resolution (Accela-LTQ Velos Orbitrap; m/z range from 85-2000) in triplicate. A dual chromatography setup was utilized, using AE and C18. Electrospray ionization was performed in positive ion mode. Data collection occurred continuously throughout 10 min of chromatographic separation.
Raw files were converted to .cdf files using Xcalibur file converter from Thermo Fisher (San Diego, CA). Data extraction was performed using apLCMS [59] and xMSanalyzer [60], generating m/z features, a m/z feature consisting of mass-to-charge ratio (m/z), retention time (RT) and ion intensity. Feature and sample filtering retained features with a median CV of 50% or less, a minimum mean Pearson correlation coefficient of 0.7 between technical replicates of each sample, and which were detected in at least 30% of samples. Two pooled human reference plasma samples were concurrently analyzed, as described by [61]. Briefly, NIST SRM1950 was analyzed at the beginning and end of the study. A second pooled reference sample (Q-Standard 3; Qstd3) was analyzed at the beginning of each batch of 20 samples. Qstd3 was prepared from plasma pooled from 2 separate lots from Equitech-Bio, Inc (Kerrville, Texas).
Metabolite annotation and identification
The m/z features were annotated for possible identities against the Human Metabolome Database (http://www.hmdb.ca/) and the LIPID MAPS Lipidomics Gateway (http://www.lipidmaps.org/) using xMSannotator [37], which scores annotations based on correlation modularity clustering and isotopic, adduct and mass defect grouping. This provides annotation at a confidence equivalent to a Schymanski level of confidence (SLC) 3 as defined by Schymanski et al. [62]. All metabolites were matched at < 10 ppm accuracy. All acylcarnitines were detected in the form of a M + H adduct. All age-associated acylcarnitines were tested for probable structure using MS/MS fragmentation with the Accela-LTQ Velos Orbitrap. Acylcarnitines for which probable structure could be identified by diagnostic fragments compared against values from the METLIN MS/MS metabolite database [63] were designated as SLC 2b as defined by Schymanski et al. [62].
Metabolome-wide association study of age
Using the R package, xmsPANDA (https://github.com/kuppal2/xmsPANDA), a metabolome-wide association study (MWAS) was performed to identify m/z features which associate with age in healthy individuals. Intensities for m/z features from xMSanalyzer, which had intensities for at least 80% of samples were log2 transformed and quantile normalized. Missing values were replaced by half of the minimum reported intensity for that feature. Normalized m/z features were tested by Spearman’s rank correlation for association with age, with a p < 0.05 considered significant. A Benjamini/Hochberg false discovery rate (FDR) method was used to correct for multiple comparisons, with an FDR threshold of 0.2 [64]. Hierarchical clustering analysis (HCA) was performed using MetaboAnalyst [65].
All acylcarnitines found to associate with age from the C18 column were analyzed for correlation between acylcarnitine subtypes. Age-associated acylcarnitines from the C18 column were chosen over those from the AE column due to greater coverage of acylcarnitines. In addition to those found to be age-associated, free carnitine (C0) and acetylcarnitine (C2) were included for analysis. Correlations were analyzed using Spearman’s rank correlation test, and a Bonferroni correction was applied [66], resulting in p < 1.98 * 104 being considered significant.
The top 6 age-associated acylcarnitines were tested for associations with other components of the metabolome in each gender using xMWAS based on partial least-squares regression [67]. After using the data normalization and filtering methods described above, the top 6 age-associated acylcarnitines were tested for associations with 26040 metabolites within both female (n = 78) and male (n = 85) subsets of the population. In generation of both networks, metabolites were filtered so that the 10000 metabolites with the lowest relative standard deviation were included in the partial least squares analysis. Thresholds for inclusion in the network were |r| > 0.30 and p < 0.01. Pairwise results from MWAS of the metabolites used for generation of network structures were used for pathway enrichment analysis using mummichog (v1) [68]. For each gender, enriched pathways were filtered for those that included at least 3 significantly associated metabolites at p < 0.05.
Supplementary Materials
Acknowledgments
Dr. Young-Mi Go and Dr. Dean P. Jones share equal senior authorship in this collaborative research.
Conflicts of Interest
The authors declare no conflicts and interest.
Funding
This work was funded by NIEHS Grants R01 ES023485 (DPJ and YMG) and P30 ES019776 (DPJ), and NIH S10 OD018006 (DPJ).
References
- 1. Hoffman JM, Lyu Y, Pletcher SD, Promislow DE. Proteomics and metabolomics in ageing research: from biomarkers to systems biology. Essays Biochem. 2017; 61:379–88. https://doi.org/10.1042/EBC20160083 [PubMed]
- 2. Franceschi C, Garagnani P, Morsiani C, Conte M, Santoro A, Grignolio A, Monti D, Capri M, Salvioli S. The continuum of aging and age-related diseases: common mechanisms but different rates. Front Med (Lausanne). 2018; 5:61. https://doi.org/10.3389/fmed.2018.00061 [PubMed]
- 3. World Population Ageing 2015 (ST/ESA/SER.A/390). United Nations, Department of Economic and Social Affairs, Population Division. 2015.
- 4. Jaul E, Barron J. Age-related diseases and clinical and public health implications for the 85 years old and over population. Front Public Health. 2017; 5:335. https://doi.org/10.3389/fpubh.2017.00335 [PubMed]
- 5. Cristofalo VJ, Gerhard GS, Pignolo RJ. Molecular biology of aging. Surg Clin North Am. 1994; 74:1–21. https://doi.org/10.1016/s0039-6109(16)46225-0 [PubMed]
- 6. Johnson FB, Sinclair DA, Guarente L. Molecular biology of aging. Cell. 1999; 96:291–302. https://doi.org/10.1016/s0092-8674(00)80567-x [PubMed]
- 7. Jones DP, Mody VC
Jr , Carlson JL, Lynn MJ, Sternberg PJr . Redox analysis of human plasma allows separation of pro-oxidant events of aging from decline in antioxidant defenses. Free Radic Biol Med. 2002; 33:1290–300. https://doi.org/10.1016/s0891-5849(02)01040-7 [PubMed] - 8. Mishur RJ, Rea SL. Applications of mass spectrometry to metabolomics and metabonomics: detection of biomarkers of aging and of age-related diseases. Mass Spectrom Rev. 2012; 31:70–95. https://doi.org/10.1002/mas.20338 [PubMed]
- 9. Bakula D, Ablasser A, Aguzzi A, Antebi A, Barzilai N, Bittner MI, Jensen MB, Calkhoven CF, Chen D, Grey AD, Feige JN, Georgievskaya A, Gladyshev VN, et al. Latest advances in aging research and drug discovery. Aging (Albany NY). 2019; 11:9971–81. https://doi.org/10.18632/aging.102487 [PubMed]
- 10. Bremer J. Carnitine—metabolism and functions. Physiol Rev. 1983; 63:1420–80. https://doi.org/10.1152/physrev.1983.63.4.1420 [PubMed]
- 11. Kalim S, Clish CB, Wenger J, Elmariah S, Yeh RW, Deferio JJ, Pierce K, Deik A, Gerszten RE, Thadhani R, Rhee EP. A plasma long-chain acylcarnitine predicts cardiovascular mortality in incident dialysis patients. J Am Heart Assoc. 2013; 2:e000542. https://doi.org/10.1161/JAHA.113.000542 [PubMed]
- 12. Koh AS, Gao F, Liu J, Fridianto KT, Ching J, Tan RS, Wong JI, Chua SJ, Leng S, Zhong L, Keng BM, Huang FQ, Yuan JM, et al. Metabolomic profile of arterial stiffness in aged adults. Diab Vasc Dis Res. 2018; 15:74–80. https://doi.org/10.1177/1479164117733627 [PubMed]
- 13. Bouchouirab FZ, Fortin M, Noll C, Dubé J, Carpentier AC. Plasma palmitoyl-carnitine (AC16:0) is a marker of increased postprandial nonesterified incomplete fatty acid oxidation rate in adults with type 2 diabetes. Can J Diabetes. 2018; 42:382–88.e1. https://doi.org/10.1016/j.jcjd.2017.09.002 [PubMed]
- 14. Mitchell SL, Uppal K, Williamson SM, Liu K, Burgess LG, Tran V, Umfress AC, Jarrell KL, Cooke Bailey JN, Agarwal A, Pericak-Vance M, Haines JL, Scott WK, et al. The carnitine shuttle pathway is altered in patients with neovascular age-related macular degeneration. Invest Ophthalmol Vis Sci. 2018; 59:4978–85. https://doi.org/10.1167/iovs.18-25137 [PubMed]
- 15. Rizza S, Copetti M, Rossi C, Cianfarani MA, Zucchelli M, Luzi A, Pecchioli C, Porzio O, Di Cola G, Urbani A, Pellegrini F, Federici M. Metabolomics signature improves the prediction of cardiovascular events in elderly subjects. Atherosclerosis. 2014; 232:260–64. https://doi.org/10.1016/j.atherosclerosis.2013.10.029 [PubMed]
- 16. Adams SH, Hoppel CL, Lok KH, Zhao L, Wong SW, Minkler PE, Hwang DH, Newman JW, Garvey WT. Plasma acylcarnitine profiles suggest incomplete long-chain fatty acid beta-oxidation and altered tricarboxylic acid cycle activity in type 2 diabetic african-american women. J Nutr. 2009; 139:1073–81. https://doi.org/10.3945/jn.108.103754 [PubMed]
- 17. Mihalik SJ, Goodpaster BH, Kelley DE, Chace DH, Vockley J, Toledo FG, DeLany JP. Increased levels of plasma acylcarnitines in obesity and type 2 diabetes and identification of a marker of glucolipotoxicity. Obesity (Silver Spring). 2010; 18:1695–700. https://doi.org/10.1038/oby.2009.510 [PubMed]
- 18. Tootsi K, Kals J, Zilmer M, Paapstel K, Ottas A, Märtson A. Medium- and long-chain acylcarnitines are associated with osteoarthritis severity and arterial stiffness in end-stage osteoarthritis patients: a case-control study. Int J Rheum Dis. 2018; 21:1211–18. https://doi.org/10.1111/1756-185X.13251 [PubMed]
- 19. Novotna B, Abdel-Hamid M, Koblizek V, Svoboda M, Hejduk K, Rehacek V, Bis J, Salajka F. A pilot data analysis of a metabolomic HPLC-MS/MS study of patients with COPD. Adv Clin Exp Med. 2018; 27:531–39. https://doi.org/10.17219/acem/68763 [PubMed]
- 20. Leruez S, Marill A, Bresson T, de Saint Martin G, Buisset A, Muller J, Tessier L, Gadras C, Verny C, Gohier P, Amati-Bonneau P, Lenaers G, Bonneau D, et al. A metabolomics profiling of glaucoma points to mitochondrial dysfunction, senescence, and polyamines deficiency. Invest Ophthalmol Vis Sci. 2018; 59:4355–61. https://doi.org/10.1167/iovs.18-24938 [PubMed]
- 21. Ciavardelli D, Piras F, Consalvo A, Rossi C, Zucchelli M, Di Ilio C, Frazzini V, Caltagirone C, Spalletta G, Sensi SL. Medium-chain plasma acylcarnitines, ketone levels, cognition, and gray matter volumes in healthy elderly, mildly cognitively impaired, or alzheimer’s disease subjects. Neurobiol Aging. 2016; 43:1–12. https://doi.org/10.1016/j.neurobiolaging.2016.03.005 [PubMed]
- 22. Cristofano A, Sapere N, La Marca G, Angiolillo A, Vitale M, Corbi G, Scapagnini G, Intrieri M, Russo C, Corso G, Di Costanzo A. Serum levels of acyl-carnitines along the continuum from normal to alzheimer’s dementia. PLoS One. 2016; 11:e0155694. https://doi.org/10.1371/journal.pone.0155694 [PubMed]
- 23. Lin CN, Huang CC, Huang KL, Lin KJ, Yen TC, Kuo HC. A metabolomic approach to identifying biomarkers in blood of alzheimer’s disease. Ann Clin Transl Neurol. 2019; 6:537–45. https://doi.org/10.1002/acn3.726 [PubMed]
- 24. Rutkowsky JM, Knotts TA, Ono-Moore KD, McCoin CS, Huang S, Schneider D, Singh S, Adams SH, Hwang DH. Acylcarnitines activate proinflammatory signaling pathways. Am J Physiol Endocrinol Metab. 2014; 306:E1378–87. https://doi.org/10.1152/ajpendo.00656.2013 [PubMed]
- 25. Reuter SE, Evans AM. Carnitine and acylcarnitines: pharmacokinetic, pharmacological and clinical aspects. Clin Pharmacokinet. 2012; 51:553–72. https://doi.org/10.1007/BF03261931 [PubMed]
- 26. McGill MR, Li F, Sharpe MR, Williams CD, Curry SC, Ma X, Jaeschke H. Circulating acylcarnitines as biomarkers of mitochondrial dysfunction after acetaminophen overdose in mice and humans. Arch Toxicol. 2014; 88:391–401. https://doi.org/10.1007/s00204-013-1118-1 [PubMed]
- 27. Shigenaga MK, Hagen TM, Ames BN. Oxidative damage and mitochondrial decay in aging. Proc Natl Acad Sci USA. 1994; 91:10771–78. https://doi.org/10.1073/pnas.91.23.10771 [PubMed]
- 28. Ames BN, Shigenaga MK, Hagen TM. Mitochondrial decay in aging. Biochim Biophys Acta. 1995; 1271:165–70. https://doi.org/10.1016/0925-4439(95)00024-x [PubMed]
- 29. Lesnefsky EJ, Chen Q, Hoppel CL. Mitochondrial metabolism in aging heart. Circ Res. 2016; 118:1593–611. https://doi.org/10.1161/CIRCRESAHA.116.307505 [PubMed]
- 30. Consitt LA, Koves TR, Muoio DM, Nakazawa M, Newton CA, Houmard JA. Plasma acylcarnitines during insulin stimulation in humans are reflective of age-related metabolic dysfunction. Biochem Biophys Res Commun. 2016; 479:868–74. https://doi.org/10.1016/j.bbrc.2016.09.116 [PubMed]
- 31. Lum H, Sloane R, Huffman KM, Kraus VB, Thompson DK, Kraus WE, Bain JR, Stevens R, Pieper CF, Taylor GA, Newgard CB, Cohen HJ, Morey MC. Plasma acylcarnitines are associated with physical performance in elderly men. J Gerontol A Biol Sci Med Sci. 2011; 66:548–53. https://doi.org/10.1093/gerona/glr006 [PubMed]
- 32. Yu Z, Zhai G, Singmann P, He Y, Xu T, Prehn C, Römisch-Margl W, Lattka E, Gieger C, Soranzo N, Heinrich J, Standl M, Thiering E, et al. Human serum metabolic profiles are age dependent. Aging Cell. 2012; 11:960–67. https://doi.org/10.1111/j.1474-9726.2012.00865.x [PubMed]
- 33. Muilwijk M, Vaz FM, Celis-Morales C, Peters RJ, van Valkengoed IG. The association of acylcarnitines and amino acids with age in dutch and south-asian Surinamese living in amsterdam. J Clin Endocrinol Metab. 2018; 103:3783–91. https://doi.org/10.1210/jc.2018-00809 [PubMed]
- 34. Ma S, Yim SH, Lee SG, Kim EB, Lee SR, Chang KT, Buffenstein R, Lewis KN, Park TJ, Miller RA, Clish CB, Gladyshev VN. Organization of the mammalian metabolome according to organ function, lineage specialization, and longevity. Cell Metab. 2015; 22:332–43. https://doi.org/10.1016/j.cmet.2015.07.005 [PubMed]
- 35. Longo N, Frigeni M, Pasquali M. Carnitine transport and fatty acid oxidation. Biochim Biophys Acta. 2016; 1863:2422–35. https://doi.org/10.1016/j.bbamcr.2016.01.023 [PubMed]
- 36. Peng KY, Watt MJ, Rensen S, Greve JW, Huynh K, Jayawardana KS, Meikle PJ, Meex RC. Mitochondrial dysfunction-related lipid changes occur in nonalcoholic fatty liver disease progression. J Lipid Res. 2018; 59:1977–86. https://doi.org/10.1194/jlr.M085613 [PubMed]
- 37. Uppal K, Walker DI, Jones DP. xMSannotator: an R package for network-based annotation of high-resolution metabolomics data. Anal Chem. 2017; 89:1063–67. https://doi.org/10.1021/acs.analchem.6b01214 [PubMed]
- 38. Beermann C, Jelinek J, Reinecker T, Hauenschild A, Boehm G, Klör HU. Short term effects of dietary medium-chain fatty acids and n-3 long-chain polyunsaturated fatty acids on the fat metabolism of healthy volunteers. Lipids Health Dis. 2003; 2:10. https://doi.org/10.1186/1476-511X-2-10 [PubMed]
- 39. Aon MA, Bhatt N, Cortassa SC. Mitochondrial and cellular mechanisms for managing lipid excess. Front Physiol. 2014; 5:282. https://doi.org/10.3389/fphys.2014.00282 [PubMed]
- 40. Adeva-Andany MM, López-Maside L, Donapetry-García C, Fernández-Fernández C, Sixto-Leal C. Enzymes involved in branched-chain amino acid metabolism in humans. Amino Acids. 2017; 49:1005–28. https://doi.org/10.1007/s00726-017-2412-7 [PubMed]
- 41. Shao D, Villet O, Zhang Z, Choi SW, Yan J, Ritterhoff J, Gu H, Djukovic D, Christodoulou D, Kolwicz SC
Jr , Raftery D, Tian R. Glucose promotes cell growth by suppressing branched-chain amino acid degradation. Nat Commun. 2018; 9:2935. https://doi.org/10.1038/s41467-018-05362-7 [PubMed] - 42. Ferdinandusse S, Denis S, Van Roermund CW, Wanders RJ, Dacremont G. Identification of the peroxisomal beta-oxidation enzymes involved in the degradation of long-chain dicarboxylic acids. J Lipid Res. 2004; 45:1104–11. https://doi.org/10.1194/jlr.M300512-JLR200 [PubMed]
- 43. Pawlak M, Lefebvre P, Staels B. Molecular mechanism of PPARα action and its impact on lipid metabolism, inflammation and fibrosis in non-alcoholic fatty liver disease. J Hepatol. 2015; 62:720–33. https://doi.org/10.1016/j.jhep.2014.10.039 [PubMed]
- 44. Nofal M, Zhang K, Han S, Rabinowitz JD. mTOR inhibition restores amino acid balance in cells dependent on catabolism of extracellular protein. Mol Cell. 2017; 67:936–46.e5. https://doi.org/10.1016/j.molcel.2017.08.011 [PubMed]
- 45. Pfeuffer M, Jaudszus A. Pentadecanoic and heptadecanoic acids: multifaceted odd-chain fatty acids. Adv Nutr. 2016; 7:730–34. https://doi.org/10.3945/an.115.011387 [PubMed]
- 46. Khaw KT, Friesen MD, Riboli E, Luben R, Wareham N. Plasma phospholipid fatty acid concentration and incident coronary heart disease in men and women: the EPIC-norfolk prospective study. PLoS Med. 2012; 9:e1001255. https://doi.org/10.1371/journal.pmed.1001255 [PubMed]
- 47. Santaren ID, Watkins SM, Liese AD, Wagenknecht LE, Rewers MJ, Haffner SM, Lorenzo C, Hanley AJ. Serum pentadecanoic acid (15:0), a short-term marker of dairy food intake, is inversely associated with incident type 2 diabetes and its underlying disorders. Am J Clin Nutr. 2014; 100:1532–40. https://doi.org/10.3945/ajcn.114.092544 [PubMed]
- 48. Sebastian RS, Goldman JD, Enns CW, LaComb RP. Fluid Milk Consumption in the United States: What We Eat In America, NHANES 2005-2006 (Dietary Data Brief No. 3). United States Department of Agriculture, Agricultural Research Service. 2010.
- 49. Jenkins B, West JA, Koulman A. A review of odd-chain fatty acid metabolism and the role of pentadecanoic acid (c15:0) and heptadecanoic acid (c17:0) in health and disease. Molecules. 2015; 20:2425–44. https://doi.org/10.3390/molecules20022425 [PubMed]
- 50. Weitkunat K, Schumann S, Nickel D, Hornemann S, Petzke KJ, Schulze MB, Pfeiffer AF, Klaus S. Odd-chain fatty acids as a biomarker for dietary fiber intake: a novel pathway for endogenous production from propionate. Am J Clin Nutr. 2017; 105:1544–51. https://doi.org/10.3945/ajcn.117.152702 [PubMed]
- 51. Funk CD. Prostaglandins and leukotrienes: advances in eicosanoid biology. Science. 2001; 294:1871–75. https://doi.org/10.1126/science.294.5548.1871 [PubMed]
- 52. Sorgdrager FJ, Naudé PJ, Kema IP, Nollen EA, Deyn PP. Tryptophan metabolism in inflammaging: from biomarker to therapeutic target. Front Immunol. 2019; 10:2565. https://doi.org/10.3389/fimmu.2019.02565 [PubMed]
- 53. Nevalainen T, Kananen L, Marttila S, Jylhä M, Hervonen A, Hurme M, Jylhävä J. Transcriptomic and epigenetic analyses reveal a gender difference in aging-associated inflammation: the vitality 90+ study. Age (Dordr). 2015; 37:9814. https://doi.org/10.1007/s11357-015-9814-9 [PubMed]
- 54. Das A. “Inflammaging” and estradiol among older U.S. Women: a nationally representative longitudinal study. Biodemography Soc Biol. 2017; 63:295–308. https://doi.org/10.1080/19485565.2017.1403304 [PubMed]
- 55. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013; 153:1194–217. https://doi.org/10.1016/j.cell.2013.05.039 [PubMed]
- 56. Braganza A, Corey CG, Santanasto AJ, Distefano G, Coen PM, Glynn NW, Nouraie SM, Goodpaster BH, Newman AB, Shiva S. Platelet bioenergetics correlate with muscle energetics and are altered in older adults. JCI Insight. 2019; 5:e128248. https://doi.org/10.1172/jci.insight.128248 [PubMed]
- 57. Go YM, Kim CW, Walker DI, Kang DW, Kumar S, Orr M, Uppal K, Quyyumi AA, Jo H, Jones DP. Disturbed flow induces systemic changes in metabolites in mouse plasma: a metabolomics study using ApoE⁻/ ⁻ mice with partial carotid ligation. Am J Physiol Regul Integr Comp Physiol. 2015; 308:R62–72. https://doi.org/10.1152/ajpregu.00278.2014 [PubMed]
- 58. Go YM, Walker DI, Soltow QA, Uppal K, Wachtman LM, Strobel FH, Pennell K, Promislow DE, Jones DP. Metabolome-wide association study of phenylalanine in plasma of common marmosets. Amino Acids. 2015; 47:589–601. https://doi.org/10.1007/s00726-014-1893-x [PubMed]
- 59. Yu T, Park Y, Johnson JM, Jones DP. apLCMS—adaptive processing of high-resolution LC/MS data. Bioinformatics. 2009; 25:1930–36. https://doi.org/10.1093/bioinformatics/btp291 [PubMed]
- 60. Uppal K, Soltow QA, Strobel FH, Pittard WS, Gernert KM, Yu T, Jones DP. xMSanalyzer: automated pipeline for improved feature detection and downstream analysis of large-scale, non-targeted metabolomics data. BMC Bioinformatics. 2013; 14:15. https://doi.org/10.1186/1471-2105-14-15 [PubMed]
- 61. Go YM, Walker DI, Liang Y, Uppal K, Soltow QA, Tran V, Strobel F, Quyyumi AA, Ziegler TR, Pennell KD, Miller GW, Jones DP. Reference standardization for mass spectrometry and high-resolution metabolomics applications to exposome research. Toxicol Sci. 2015; 148:531–43. https://doi.org/10.1093/toxsci/kfv198 [PubMed]
- 62. Schymanski EL, Jeon J, Gulde R, Fenner K, Ruff M, Singer HP, Hollender J. Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ Sci Technol. 2014; 48:2097–98. https://doi.org/10.1021/es5002105 [PubMed]
- 63. Guijas C, Montenegro-Burke JR, Domingo-Almenara X, Palermo A, Warth B, Hermann G, Koellensperger G, Huan T, Uritboonthai W, Aisporna AE, Wolan DW, Spilker ME, Benton HP, Siuzdak G. METLIN: a technology platform for identifying knowns and unknowns. Anal Chem. 2018; 90:3156–64. https://doi.org/10.1021/acs.analchem.7b04424 [PubMed]
- 64. Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society: Series B (Methodological). 1995; 57:289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
- 65. Chong J, Soufan O, Li C, Caraus I, Li S, Bourque G, Wishart DS, Xia J. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018; 46:W486–94. https://doi.org/10.1093/nar/gky310 [PubMed]
- 66. Sedgwick P. Multiple hypothesis testing and bonferroni’s correction. BMJ. 2014; 349:g6284. https://doi.org/10.1136/bmj.g6284 [PubMed]
- 67. Uppal K, Ma C, Go YM, Jones DP, Wren J. xMWAS: a data-driven integration and differential network analysis tool. Bioinformatics. 2018; 34:701–02. https://doi.org/10.1093/bioinformatics/btx656 [PubMed]
- 68. Li S, Park Y, Duraisingham S, Strobel FH, Khan N, Soltow QA, Jones DP, Pulendran B. Predicting network activity from high throughput metabolomics. PLoS Comput Biol. 2013; 9:e1003123. https://doi.org/10.1371/journal.pcbi.1003123 [PubMed]


