Introduction
Lung cancer is one of the most common types of malignancies worldwide and is the leading cause of cancer-related mortality. According to Globacan, more than 1 million patients die from lung cancer worldwide annually [1, 2]. Lung adenocarcinoma (LUAD) is a histological type of non-small cell lung cancer that accounts for a large proportion of cases with this disease [3]. Among the major malignancies, the economic burden of LUAD remained high for decades [4]. With the accumulation of studies on the etiology and pathogenesis of LUAD, the immune response is considered crucial to the development and progression of LUAD; this means that an improvement in the tumor immune response could be beneficial for LUAD patients, thus improving the clinical symptoms and overall survival (OS) [5]. Immunotherapy has been recently developed and widely used in patients with lung cancer, including cytotoxic T lymphocyte-associated antigen 4 inhibitors and immune checkpoint inhibitors [6–8]. Despite the advancements in targeted therapy and immunotherapy, the prognosis of patients with LUAD remains poor [9]. In the era of precision medicine, it is necessary to elucidate the relationship between LUAD and immune infiltration and explore a reliable immune-related biomarker that can predict the prognosis of LUAD and become a novel target for LUAD immunotherapy.
The Janus kinase (JAK) family is one of the families of non-receptor tyrosine kinases [10]. These JAKs are critically involved in cell growth, hematopoiesis, and immunoregulation by participating in the signal transduction of hematopoietic cytokines and growth factors [11]. JAK1 is a member of the JAK family, which plays an essential and non-redundant role in the proliferation, differentiation, and metastasis of cancer cells by mediating the expression of interleukin-6 (IL-6)/JAK1/signal transducer and activator of transcription-3 (STAT3) or JAK1/phosphatidylinositol-3-kinase (PI3K) signaling pathway [12, 13]. JAK1-deficient or mutant cell lines show greater tumorigenicity, which has been reported in patients with hepatocellular carcinoma, acute lymphoblastic leukemia, lung cancer, and gastric cancer [14–16]. A previous study suggested that JAK1 mediates many immune regulatory processes, including those that are involved in tumor-driven immune escape, cancer immunosuppression, and sustained inflammation in the tumor microenvironment [17]. Programmed death-ligand 1 induced by tumor-associated macrophages promotes the progression of lung cancer through the JAK1/STAT3 signaling pathway, and the JAK1 inhibitor significantly improves the efficacy of anti-programmed cell death protein 1 immunotherapy in pancreatic cancer by inhibiting the activation of cytotoxic T lymphocytes [18, 19]. Moreover, a lower JAK1 expression is associated with T-cell infiltration and a poor prognosis in breast cancer [20]. However, the potential role of JAK1 in tumor immune infiltration and the prognosis of LUAD remains unclear.
Since early December 2019, coronavirus disease 2019 (COVID-19), which first occurred in Wuhan City, Hubei Province, China, is among the types of pneumonia with unknown etiology [21]. It is caused by a novel positive-sense RNA virus, named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which belongs to the Betacoronavirus genus in the Coronaviridae family [22]. The outbreak of COVID-19 was declared as a public health emergency of international concern [23]. After its initial discovery, SARS-CoV-2 rapidly spread all over the world, with over 6.2 million confirmed cases COVID-19 and more than 370,000 deaths as of June 1, 2020 [24]. Moreover, recent researches showed that cancer patients were at increased risk of developing COVID-19 with worse outcomes than individuals without cancer, and a significant increase of mortality rate in lung cancer patients with SARS-CoV-2 infection was detected [25, 26]. Despite the clinical features of lung cancer patients, with a higher risk to respiratory infections, immunosuppressed patients, and most of them with a previous of COPD and metastatic disease, the underlying mechanisms in the development and poor prognosis of LUAD patients with COVID-19 are still unclear.
In this study, JAK1 expression in LUAD and the correlations between JAK1 expression and the clinical features and prognosis of LUAD were analyzed using the Oncomine, UALCAN, and Kaplan-Meier plotter databases. The potential molecular mechanism of JAK1 was determined by conducting a GSEA using the Cancer Genome Atlas Lung Adenocarcinoma (TCGA-LUAD) dataset. The association between JAK1 and tumor immune cell infiltration was investigated using the TIMER and CIBERSORT databases. In addition, the JAK1 expression and proportions of immune cells in patients with LUAD with SARS-Cov-2 infection were analyzed. The present study revealed the potential role of JAK1 in LUAD immunology and its prognostic value, which will enhance the understanding on the underlying mechanisms of LUAD.
Results
Decreased expression of JAK1 in LUAD
The expression levels of JAK1 in tumor and adjacent normal tissues of various cancers were analyzed based on the Oncomine database. As shown in Figure 1A, the result revealed that the expression levels of JAK1 were lower in lung cancer tissues than in normal lung tissues. In addition, the expression levels of JAK1 mRNA were higher in brain and CNS, cervical, colorectal, esophageal, head and neck, kidney, and myeloma cancers. To further assess the expression of JAK1 in multiple malignancies, we analyzed the expression levels of JAK1 mRNA in various types of cancer using the UALCAN database. The expression level of JAK1 was higher in clear cell renal cell carcinoma, papillary renal cell carcinoma, and thyroid carcinoma. By contrast, the expression of JAK1 was significantly decreased in bladder carcinoma, breast carcinoma, colon adenocarcinoma, chromophobe renal cell carcinoma, liver hepatocellular carcinoma, LUAD, lung squamous cell carcinoma, prostate cancer, and adenocarcinoma of the rectum (Figure 1B). These data show that JAK1 has a differential expression and may be regarded as an oncogenic gene associated with the development of various tumors. As shown in Figure 1C, the quantitative evaluation of the expression level of JAK1 in LUAD samples from RNA-seq data of TCGA and microarray data showed that the JAK1 mRNA expression level in LUAD tissues was significantly lower than that in normal tissues. Overall, these results suggest that the expression of JAK1 is decreased in various types of cancer, particularly in LUAD, demonstrating that JAK1 may suppress LUAD tumorigenesis.
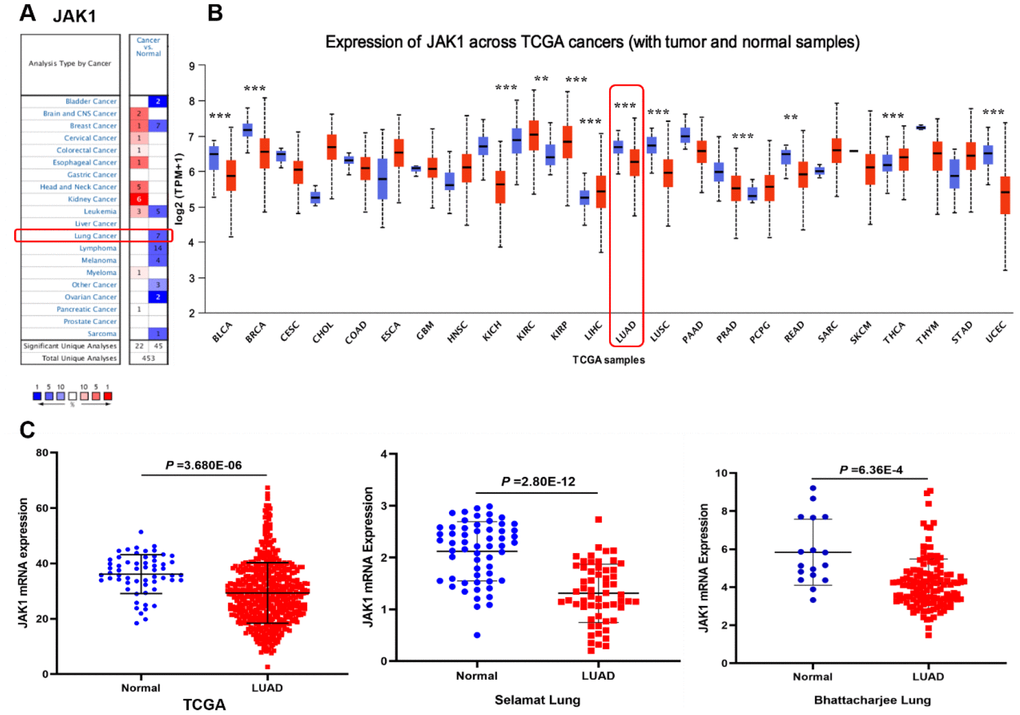
Figure 1. Expression level of JAK1 in different human cancers. (A) Increased or decreased expression of JAK1 in datasets of different types of cancers compared with normal tissues in the Oncomine database. (B) JAK1 expression levels in different cancers from the TCGA database detected by using the UALCAN database. Bule represents Normal, Red represents Tumor. (C) Comparison of JAK1 expression levels between LUAD tissues and corresponding normal tissues in different datasets including TCGA, Selamat lung, and Bhattacharjee lung datasets.
Associations between JAK1 and clinical characteristics of LUAD patients
Since the JAK1 expression was significantly decreased in LUAD samples compared with that in the adjacent normal tissues, we focused on the detection of JAK1 expression in LUAD patients with different conditions by conducting further UALCAN database analysis. Results showed that JAK1 expression was much lower in patients with LUAD (aged 41–60 years and 61–80 years) than in normal individuals (Figure 2A). After conducting further analysis of the expression of JAK1 based on gender and race, we found that JAK1 was highly associated with sex and race, and the LUAD male patients had the lowest JAK2 expression levels (Figure 2B, 2C). Another significant clinical feature was the influence of nodal metastasis and individual cancer stages on JAK1 expression. Results indicated that JAK1 expression was closely associated with cancer stage and nodal metastasis (Figure 2D, 2E). A differential analysis of the expression of JAK1 based on TP53 mutation status was also conducted. We found that patients with LUAD with a TP53 mutation status had lower JAK1 expression (Figure 2F). All of these results indicated that a low JAK1 expression was widely associated with the development of LUAD. The detailed clinical characteristics of patients with LUAD are presented in Table 1.
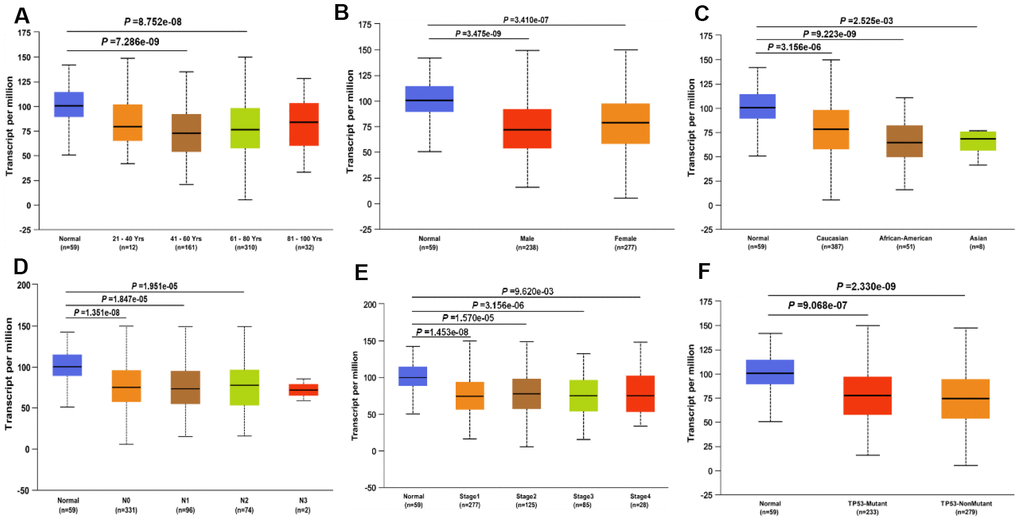
Figure 2. JAK1 transcription in subgroups of patients with LUAD, divided based on gender, age, ethnicity and other clinical features. (A) age. (B) gender. (C) ethnicity. (D) Lymph -node status. (E) Disease stage. (F) TP53 mutation status. Color images are available online.
Table 1. Clinical characteristics of patients with lung adenocarcinoma.
| Variable | Case, n | Percentage (%) |
| Age (years) | ||
| 21-40 | 12 | 2.33 |
| 41-60 | 161 | 31.26 |
| 61-80 | 310 | 60.19 |
| 81-100 | 32 | 6.22 |
| Gender | ||
| Female | 277 | 53.79 |
| male | 238 | 46.21 |
| Ethnicity | ||
| Caucasian | 387 | 86.77 |
| African-American | 51 | 11.44 |
| Asian | 8 | 1.79 |
| Lymph-node status | ||
| N0 | 331 | 65.8 |
| N1 | 96 | 19.09 |
| N2 | 74 | 14.71 |
| N3 | 2 | 0.4 |
| Disease stage | ||
| Stage 1 | 277 | 53.79 |
| Stage 2 | 125 | 24.27 |
| Stage 3 | 85 | 16.5 |
| Stage 4 | 28 | 5.44 |
| TP53 mutation status | ||
| TP53-mutant | 233 | 45.5 |
| TP53-nonmutant | 279 | 54.5 |
Protein expression and DNA methylation levels of JAK1 in patients with LUAD
To determine whether the JAK1 protein was also differentially expressed in LUAD tissues, a protein expression analysis was conducted using the data from a clinical proteomic tumor analysis consortium in the UALCAN database. Consistent with the RNA expression data, results indicated that the protein expression of JAK1 was lower in LUAD tissues than in normal tissues, and was negatively associated with tumor grade (Figure 3A, 3B). As the tumor grade increased, the expression of JAK1 decreased significantly. These evidences strongly indicated that both mRNA and protein expressions of JAK1 were significantly decreased in LUAD. To further identify the mechanisms underlying the downregulation of JAK1 in LUAD, the methylation levels of JAK1 in the LUAD dataset were analyzed. As shown in Figure 3C, the DNA methylation levels of JAK1 between normal lung samples and LUAD tissues were compared, and a significant difference was observed (p < 0.001). In addition, the gene expression of JAK1 was negatively correlated with its methylation level (Figure 3D). These results revealed that the hypermethylation of JAK1 in patients with LUAD may contribute to downregulation of JAK1 in LUAD; however, this finding requires further investigation in the future.
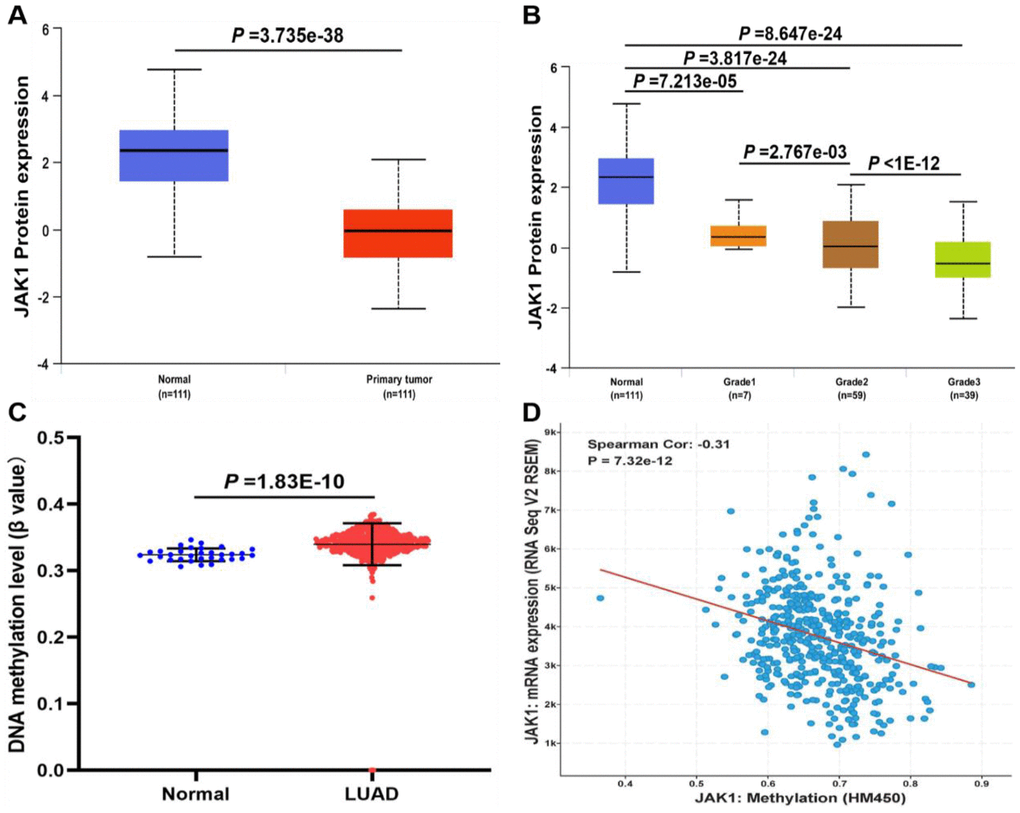
Figure 3. The protein expression and DNA methylation of JAK1 in patients with LUAD. (A) Protein expression levels of JAK1 in normal and LUAD samples. (B) protein expression levels of JAK1 in normal individuals or LUAD patients with tumor grade 1, 2, 3. (C) DNA methylation levels of JAK1 in normal individuals and LUAD patients. (D) correlation analysis of JAK1 mRNA expression with JAK1 methylation.
Prognostic value of JAK1 in patients with LUAD
To explore the prognostic value of JAK1 expression levels in patients with LUAD, the Kaplan–Meier plotter database was used. As shown in Figure 4A, low expression of JAK1 was associated with worse OS in patients with LUAD (OS, hazard ratio [HR] = 0.43, 95% confidence interval [CI] = 0.33–0.55, log-rank p = 4.2e–12). Similarly, low JAK1 expression was significantly associated with reduced progression-free survival (PFS) (Figure 4B, PFS, HR = 0.56, 95% CI = 0.41–0.77, log-rank p = 0.00025). Furthermore, low expression of JAK1 mRNA also indicated adverse PPS (Figure 4C). Overall, these results suggest that JAK1 expression is an important factor affecting the survival of patients with LUAD.
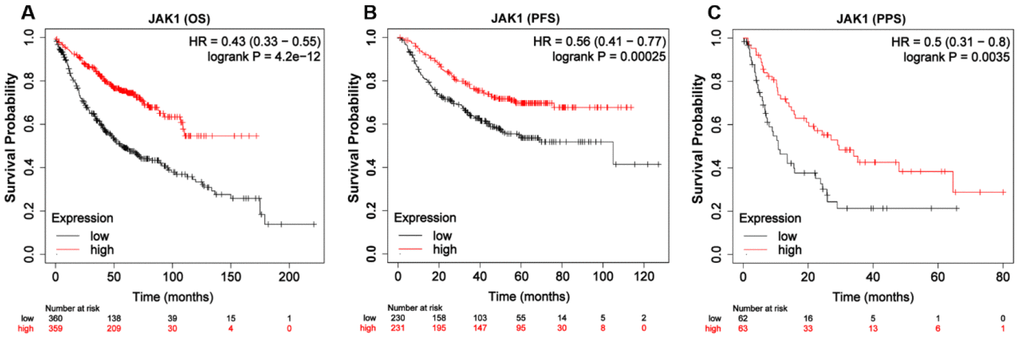
Figure 4. Kaplan-Meier survival curves of LUAD patients according to the mRNA expression of JAK1 using Kaplan-Meier plotter tool. (A) Over survival curve of patients with LUAD base on JAK1 expression levels. (B) Progression-free survival curve of patients with LUAD base on JAK1 expression levels. (C) Post-progression survival curve of patients with LUAD base on JAK1 expression levels.
Identification of JAK1-associated signaling pathways using gene set enrichment analysis
To explore the potential molecular function of JAK1 in LUAD carcinogenesis, GSEA of high- and low-JAK1 expression datasets acquired from the TCGA database was performed to explore the signaling pathways associated with JAK1. Based on the selected cutoff criteria, a total of 64 signaling pathways were significantly enriched; among them, the top 20 signaling pathways were selected and are presented in Table 2. The significantly upregulated pathways enriched in the high JAK1 group involved in cell adhesion and tumorigenesis were as follows: regulation of the actin cytoskeleton, focal adhesion, ERBB signaling pathway, MAPK signaling pathway, Toll-like receptor signaling pathway, and non-small cell lung cancer. Meanwhile, the pathways associated with immune responses were as follows: JAK-STAT signaling pathway, B-cell receptor signaling pathway, T-cell receptor signaling pathway, chemokine signaling pathway, natural killer cell mediated cytotoxicity, and leukocyte transendothelial migration. The enrichment results are presented in Figure 5.
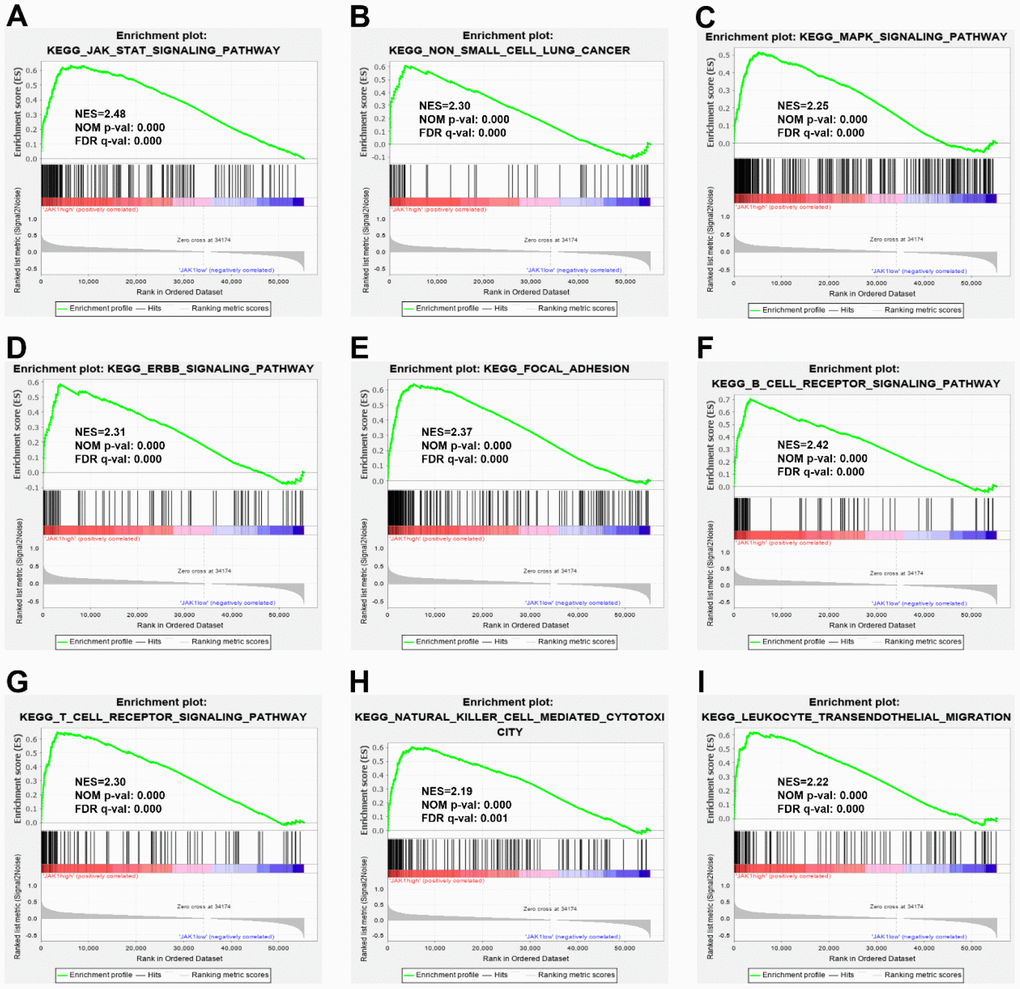
Figure 5. Enrichment plots from GSEA. GSEA results indicated that the JAK-STAT signaling pathway (A), the non-small cell lung cancer (B), the MARK signaling pathway (C), the ERBB signaling pathway (D), the focal adhesion, (E), the B cell receptor signaling pathway (F), the T cell receptor signaling pathway (G), the natural killer mediated cytotoxicity (H), and leukocyte transendothelial migration (I) were differentially enriched in LUAD samples with high JAK1 expression. NES, normalized enrichment score; NOM p-val, normalized P-value; FDR q-val, false discovery rate q-value; GSEA, Gene Set Enrichment Analysis.
Table 2. Gene sets enriched in the high JAK1 expression phenotype group.
| Gene set name | Size | NES | NOM p-val | FDR q-val | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_JAK_STAT_SIGNALING_PATHWAY | 155 | 2.48 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_B_CELL_RECEPTOR_SIGNALING_PATHWAY | 75 | 2.42 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_FOCAL_ADHESION | 199 | 2.37 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_ERBB_SIGNALING_PATHWAY | 87 | 2.31 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_T_CELL_RECEPTOR_SIGNALING_PATHWAY | 108 | 2.3 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_NON_SMALL_CELL_LUNG_CANCER | 54 | 2.3 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_REGULATION_OF_ACTIN_CYTOSKELETON | 213 | 2.28 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM | 76 | 2.28 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_GLIOMA | 65 | 2.26 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_NOD_LIKE_RECEPTOR_SIGNALING_PATHWAY | 62 | 2.26 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_FC_EPSILON_RI_SIGNALING_PATHWAY | 79 | 2.25 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_MAPK_SIGNALING_PATHWAY | 267 | 2.25 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_CHEMOKINE_SIGNALING_PATHWAY | 188 | 2.25 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_COLORECTAL_CANCER | 62 | 2.23 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_LEUKOCYTE_TRANSENDOTHELIAL_MIGRATION | 116 | 2.22 | 0 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_CHRONIC_MYELOID_LEUKEMIA | 73 | 2.21 | 0.002 | 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_TOLL_LIKE_RECEPTOR_SIGNALING_PATHWAY | 102 | 2.21 | 0 | 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_ACUTE_MYELOID_LEUKEMIA | 57 | 2.2 | 0.002 | 0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_NATURAL_KILLER_CELL_MEDIATED_CYTOTOXICITY | 132 | 2.19 | 0 | 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG_SMALL_CELL_LUNG_CANCER | 84 | 2.19 | 0 | 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NES: Normalized Enrichment Score; NOM: Nominal; FDR: False Discovery Rate. Gene sets with NOM p-value < 0.05 and FDR q-value < 0.1 are considered as statistically significant. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Association between JAK1 and tumor-infiltrating immune cells in LUAD
Considering that JAK1 is a crucial mediator of immune cell activation, a correlation analysis between JAK1 expression and six types of immune-infiltrating cells, including B cells, CD8+ T cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells, was performed. As shown in Figure 6A, JAK1 expression levels were significantly positively correlated with infiltrating levels of B cells (r = 0.163, p = 3.21e–04), CD8+ T cells (r = 0.333, p = 4.48e–14), CD4+ T cells (r = 0.37, p = 3.73e–17), macrophage (r = 0.34, p = 1.4e–14), neutrophils (r = 0.431, p = 2.98e–23), and dendritic cells (r = 0.475, p = 7.70e–29) in LUAD. Taken together, the above evidence indicates that JAK1 may be involved in the immune response of patients with LUAD by affecting the immune cells.
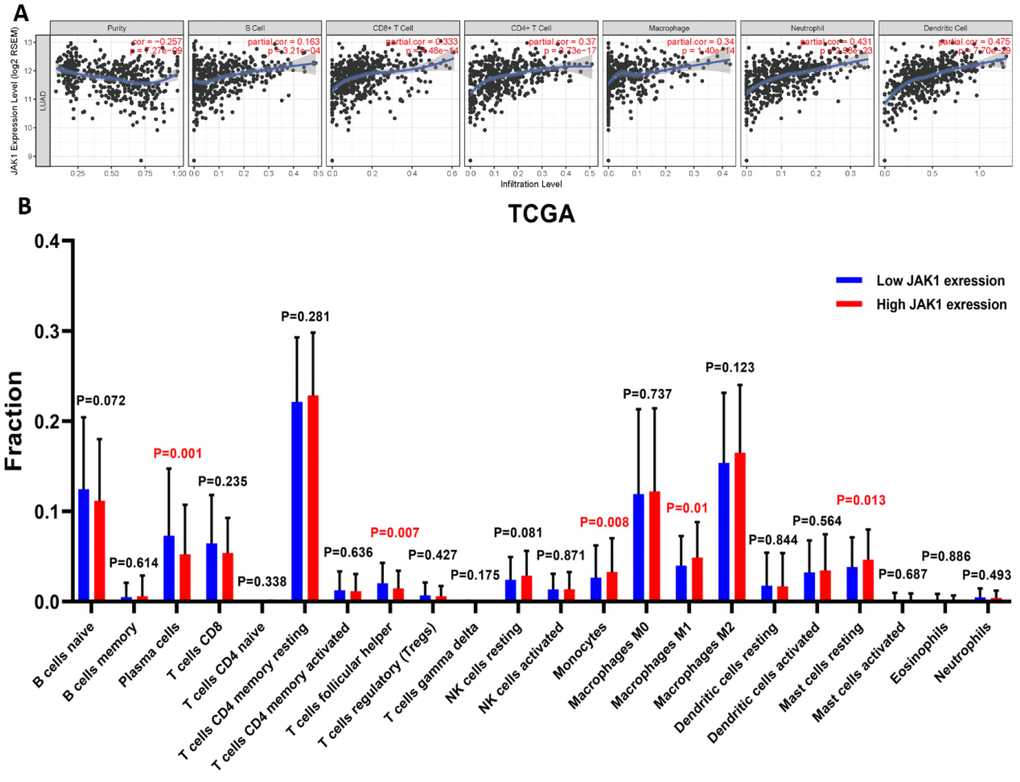
Figure 6. Correlation analysis of JAK1 expression with immune filtration level in patients with LUAD using TIMER and CIBERSORT databases.(A) JAK1 correlation with tumor purity, B cell, CD8+ T cell, CD4+ T cell, macrophage, neutrophil, dendritic cell in LUAD. (B) Proportions of the 22 tumor-infiltrating immune cell types in high and low JAK1 expression groups.
Next, we determined whether the immune infiltration status in patients with LUAD was changed between the high and low JAK1 expression groups. The high and low JAK1 expression samples among 497 TCGA LUAD samples were divided into high and low expression groups, respectively. The fraction of the 22 subpopulations of immune cell types in the high and low JAK1 expression groups was clearly determined from the results of the CIBERSORT analysis. Results showed that significantly decreased monocyte and macrophage M1 were observed in the low JAK1 expression group from the TCGA database (Figure 6B).
Moreover, JAK1 was positively correlated with the infiltrating level of macrophages (Figure 6A). The CIBERSORT analysis further revealed that M1 macrophages were significantly upregulated in the high JAK1 expression group, whereas M2 macrophages showed no significant change (Figure 6B). To further analyze the association between JAK1 and different macrophages, the correlations between JAK1 and the markers of M1 macrophages in LUAD were investigated using the TIMER database. Results showed that the M1 macrophage markers such as PTGS2, IRF5, and NOS2 were significantly associated with the expression levels of JAK1 (Supplementary Figure 1). These results indicate that JAK1 may have a potential role in the regulation of the polarization of M1 macrophages. Based on these results, we analyzed the prognostic value of JAK1 expression in LUAD based on the macrophages using the Kaplan–Meier database. As indicated in Supplementary Figure 2A, 2B, low JAK1 expression was significantly associated with unfavorable survival prognosis of LUAD in enriched macrophages (HR = 0.55), whereas a significant difference was not observed in the reduction of the number of macrophages (p > 0.05). Altogether, these results indicate that JAK1 may be involved in the immune response, leading to a worse prognosis in patients with LUAD.
Determining whether SARS-CoV-2 infection may reduce the expression of JAK1
To further explore the changes in the expression of JAK1 in LUAD cells after SARS-CoV-2 infection, the GSE147507 and GSE148729 cell lines were selected to analyze the changes in JAK1 expression in different cell lines of LUAD after SARS-CoV-2 infection. As shown in Figure 7A, 7B, the expression of JAK1 in SARS-CoV-2 infected A549 cells and Calu3 cells was significantly decreased compared with that in the control group. This finding revealed that JAK1 expression may decrease in LUAD cell lines after SARS-CoV-2 infection. Next, we explored whether the infiltration of multiple immune cells in LUAD was related to SARS-CoV-2 infection. The A549 and Calu3 samples from the GSE147507 and GSE148729 groups were divided into normal and SARS-CoV-2 infection groups. We found a significant decrease in the proportion of CD8+ T cells in the SARS-CoV-2 infection group from the potted database in the GSE147507 (Figure 7C) and GSE148729 samples (Figure 7D).
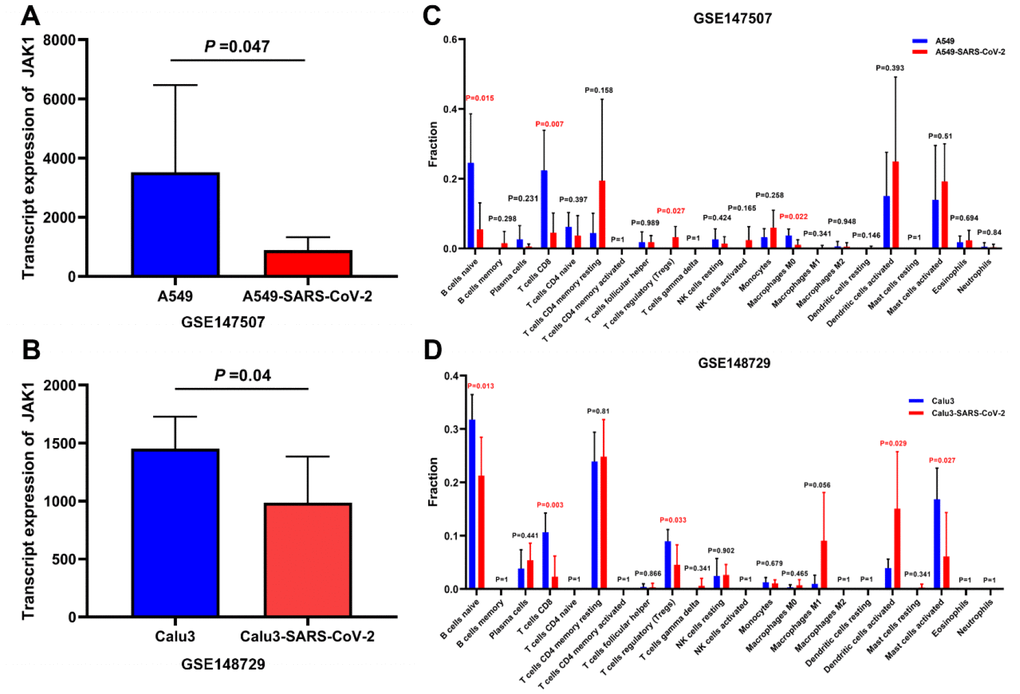
Figure 7. Analysis of the JAK1 expression and immune cells proportion in LUAD cell lines with SARS-CoV-2 infection. (A) JAK1 expression in A549 cells and A549 cells with SARS-CoV-2 infection in GSE147507. (B) JAK1 expression in Calu3 cells and Calu3 cells with SARS-CoV-2 infection. (C) Proportions of the 22 tumor-infiltrating immune cell types in A549 cells with and without SARS-CoV-2 infection. (D) Proportions of the 22 tumor-infiltrating immune cell types in Calu3 cells with and without SARS-CoV-2 infection.
Discussion
LUAD is one of the common subtypes of lung malignant neoplasms. Although the diagnostic and therapeutic strategies for LUAD have greatly improved over the past decades, the prognosis of patients with LUAD remains poor. The abnormal gene expression is involved in tumorigenesis and correlated with the prognosis of these patients [27, 28]. However, the potential role of JAK1 in tumor immune infiltration and the prognosis of LUAD remains ambiguous. In the present study, we demonstrated that JAK1 expression was remarkably decreased in LUAD tissues and was associated with gender, ethnicity, age, sex, lymph node status, disease stage, and TP53 mutation status in LUAD patients. DNA methylation is an important form of epigenetic modification of DNA, which can change the genetic expression without changing the DNA sequence, especially in hypermethylation status [29]. The methylation level of JAK1 was significantly increased and was negatively associated with JAK1 gene expression, indicating that the hypermethylation of JAK1 may cause a significant decrease in JAK1 expression. Moreover, low JAK1 expression was significantly associated with unfavorable prognosis. These results suggest that JAK1 may be involved in lung carcinogenesis and affects its prognosis.
Previous studies have reported that JAK1 is highly associated with tumorigenesis and plays important roles in the proliferation, differentiation, and apoptosis of cancer cells [13, 30]. However, the effect of JAK1 on cell proliferation and apoptosis remains unclear. JAK1 was associated with the poor survival of patients with breast cancer; JAK1 knockdown by siRNA or inhibitors can lead to the arrest of cell grow and apoptosis [20, 31]. The abnormal expression of JAK1 was observed in multiple tumor tissues. This finding indicated that JAK1 may be involved in tumorigenesis. JAK1 was significantly decreased in patients with LUAD, which means that JAK1 may play a role in the development of LUAD.
To further investigate the molecular functions and potential mechanisms of JAK1 in LUAD, the GSEA was conducted to explore the pathways enriched in high JAK1 expression samples and identify the top 20 upregulated pathways associated with cell adhesion, tumorigenesis, and the immune response. The significantly enriched pathway related to cell adhesion included the regulation of the actin cytoskeleton and focal adhesion. Perturbation of the actin cytoskeleton is involved in the impairment of the mitochondria and apoptosis, which results in the inhibition of liver cancer cell growth [32]. The upregulated molecules linked to the migratory signals of the actin cytoskeleton promote the invasion and metastasis of cancer cells [33]. Tumorigenesis-associated pathways, such as the ERBB signaling pathway, MAPK signaling pathway, Toll-like receptor signaling pathway, and non-small cell lung cancer, were also predicted in the high JAK1 expression group. In addition, the signaling pathways enriched in the high JAK1 group involved in immune responses were JAK-STAT signaling pathway, B-cell receptor signaling pathway, T-cell receptor signaling pathway, chemokine signaling pathway, natural killer cell-mediated cytotoxicity, and leukocyte transendothelial migration. Chen et al. found that JAK1 is positively correlated with immune cell infiltration, in which the dysregulation of JAK1 was associated with infiltrating macrophages in breast cancer [20]. Overall, these results suggest that JAK1 is highly associated with tumorigenesis and immune-related pathways. Therefore, JAK1 may play a role in the carcinogenesis of LUAD by affecting these signaling pathways, thus resulting in a poor prognosis in patients with LUAD.
The results of TIMER analyses showed that JAK1 was positively associated with the infiltrating levels of CD4+ T cells, CD8+ T cells, macrophages, and dendritic cells. The estimated fraction of 22 tumor-infiltrating immune cells in LUAD was analyzed using the CIBERSORT database. The results indicated that the proportions of monotypes and M1 macrophages were significantly decreased with reduction of JAK1 expression. These results suggest that JAK1 expression is closely associated with the expression of tumor-infiltrating immune cells and may be involved in the immune response to LUAD development, resulting in a poor prognosis in patients with LUAD.
Notably, the results of CIBERSORT analysis showed that the proportion of M1 macrophages in the low JAK1 expression group was significantly decreased compared with that in the high JAK1 expression group. By contrast, no significant differences were observed in the expression of M2 macrophages. To further investigate the association between JAK1 expression and M1 macrophages, the correlation between JAK1 and the marker genes of M1 macrophages was analyzed. Results showed that JAK1 has a significantly positive association with the gene markers of M1 macrophages. This evidence suggests that low JAK1 expression may downregulate the polarization of macrophages to M1 macrophages, which could contribute to the development of LUAD. M1 macrophages produce reactive oxygen and promote a Th1 response with strong tumoricidal activity [34]. A previous study reported that promoting the polarization of M1 macrophages could inhibit the proliferation, invasion, migration, and angiogenesis of tumor cells; facilitate apoptosis; and M1macrophage densities in the tumor islets, stroma, orislets and stroma were positively associated with NSCLC patient’s survival time [35, 36]. It is possible that low JAK1 expression reduces the polarization of M1 macrophages, thus accelerating the progression of LUAD and leading to a poor prognosis.
The outbreak of COVID-19 worldwide has led to a large number of human deaths and severe economic recession [24]. Data from China thus far have shown that mechanical ventilation or ICU admission is more frequently required in cancer patients with COVID-19 infection than in the general population, and higher mortality rate and severity was observed in lung cancer patients with COVID-19 [36]. Decreased JAK1 expression in LUAD patients was associated with unfavorable prognosis, but it was not clear that whether JAK1 was associated with increased mortality rate and severity in LUAD patients with COVID-19. To explore the potential role of JAK1 in the development of LUAD after SARS-CoV-2 infection, the GSE147507 and GSE148729 were used. Results showed that the expression levels of JAK1 decreased in both cell lines after SARS-CoV-2 infection. Elevated serum of interferon (IFN)-γ has been found in patients ARDS in COVID-19, and it exerts multiple effects through the JAK1/JAK2 signaling resulting to the activation of a inflammatory factors downstream cascade [37, 38]. It means that SARS-CoV-2 infection may enhance the production of cytokines and aggravate the pulmonary inflammation through the JAK1 related signaling pathways, thus leading to an increased mortality rate in LUAD patients with COVID-19. We also found that the proportion of CD8+ T cells was also significantly decreased after SARS-CoV-2 infection (Figure 7). JAK1 was involved in the activation of T cells, and its low expression was correlated with a decrease in CD8+ T cells in patients with COVID-19 pneumonia, especially in those with severe and critical cases [39, 40]. These results indicate that SARS-CoV-2 infection may reduce the proportion of CD8+ T cells in LUAD cell lines by inhibiting the expression of JAK1. Lung cancer patients are immunocompromised, especially those receiving antitumor therapy. Some studies have shown that patients with COVID-19 pneumonia have immunodeficiency and hypo-immunity, which may increase the severity of the disease and result in death [41]. Therefore, we focused on JAK1 to explore its possible effect on immune cells in LUAD and thus play a role in the development of this condition. This study is limited by the clinical data of LUAD patients with COVID-19 pneumonia, and further study is needed in the future. However, our findings provide evidence and clues for the treatment and prognosis of patients with LUAD with COVID-19 pneumonia.
In conclusion, JAK1 expression decreased significantly in patients with LUAD, which was correlated with the clinical features and predicted a poor prognosis. JAK1 expression was closely associated with the infiltrating level of immune cells; its low expression reduces the polarization of M1 macrophages. Our study revealed the potential role of JAK1 in tumor immunology and its prognostic value in LUAD. JAK1 might be used as a prognostic biomarker and therapeutic target for LUAD.
Materials and Methods
Differential expression analysis of JAK1 by oncomine and UALCAN database
Oncomine database (http://www.oncomine.org) is a web-based data open-mining platform for collecting, standardizing, analyzing the microarray data of different human cancers [42]. Besides, the UALCAN database (http://ualcan.path.uab.edu/analysis/) includes the mRNA sequencing data for 32 cancer types from TCGA [43]. In this study, we used the Oncomine and UALCAN database to study the difference of JAK1 expression between different tumors and adjacent normal samples. The threshold of Oncomine was set as follows: p-value =0.001, fold change =1.5, gene ranking =all, and the statistical significance of JAK1 expression was analyzed by Wilcoxon test. To analyze JAK1 expression in LUAD patients with different clinical features, the further UALCAN database analysis was conducted. Furthermore, the JAK1 protein expression data in LUAD were obtained from UALACN databases.
Differential expression of JAK1 methylation level by TCGA database
To explore the potential mechanism for the down-regulation of JAK1 expression, the methylation data of JAK1 in normal controls and LUAD patients were also obtained from the TCGA database, and the significance was determined by Student’s test. Additionally, the correlation between JAK1 gene expression and its methylation levels data was analyzed by the Spearman method. p-values <0.05 were considered statistically significant.
Survival analysis of JAK1 expression in LUAD based on Kaplan-Meier plotter database
The Kaplan-Meier plotter database (https://kmplot.com/analysis), contains survival information of 865 patients with LUAD. The prognostic value of JAK1 expression was assessed by overall survival (OS), progression-free survival (PFS), and post-progression survival (PPS) using the hazard ratio (HR), 95% confidence intervals (CI) and log-rank p-value [44]. All patient samples were divided into high expression groups and low expression groups based on the median expression levels of the JAK1. Besides, univariate Cox regression analysis was used to assess the prognostic value of JAK1. In this study, we used the JetSet scores to assess the representation of probe sets, and only the probe sets with the best JetSet scores for JAK1 were selected. The threshold of Cox p and log-rank p was set as 0.05.
Gene set enrichment analysis (GSEA) of JAK1 in LUAD
To further investigate the potential biological function of the JAK1 in LUAD, GSEA 4.0 was used to evaluate the associations between the high expression and low expression JAK1 groups and tumor-associated pathway in the TCGA lung adenocarcinoma dataset. The reference gene set C2 (c2.cp.kegg.v7.0.symbols.gmt) was selected and the JAK1 expression level was used as a phenotype label. The normalized enrichment score (NES) was acquired by analyzing with 1000 times permutations. The normal p-value <0.05 and the false discovery rate (FDR) <0.05 were set as the cut-off criteria.
Correlation analysis of JAK1 with immune infiltrating cells in TIMER and CIBERSORT databases
The TIMER database, which contains the infiltration levels of six types of immune cells (B cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, and dendritic cells) in various cancers, is an effective tool to analyze the abundance of tumor-infiltrating immune cell from the target gene expression data [45]. The associations between JAK1 expression levels and the above-mentioned immune cell infiltration were assessed by the TIMER database. CIBERSORT is a free trial database for evaluated immune cells composition of tissues based on their gene expression profiles [46]. After uploading the expression profiles from the TCGA-LUAD dataset with the standard gene annotations to the CIBERSORT portal, the deconvolution algorithm applying the LM22 signature and permutation =100 was used to calculated the immune cell infiltration. According to the median JAK1 expression values, the available data were split up into low and high JAK1 expression groups to evaluate the differences in the proportion of immune cells. The correlation of JAK1 expression and gene markers of M1 macrophages in LUAD was analyzed in the TIMER database. P-value <0.05 was set as the screening criterion.
The JAK1 expression and proportions of immune cells in LUAD cell lines with SARS-CoV-2 infection
GEO (https://www.ncbi.nlm.nih.gov/geo/) is an open-access database that contains various microarrays and high-throughput functional genomics data. To further analyzed the expression levels of JAK1 in LUAD after SARS-CoV-2 infection, the expression profile of GSE147507 and GSE148729 was finally obtained in the GEO database. The GSE147507 database which contained 6 SARS-CoV-2 infected A549 cell samples and 6 control A549 cell samples were based on the GPL18573 platform (Illumina NextSeq 500). Besides, the GSE148729 database which included 6 SARS-CoV-2 infected Calu3 cell samples and 6 control Calu3 cell samples were based on the GPL18573 platform (Illumina NextSeq 500). The selection criteria of samples in GSE147507 and GSE148729 were set as follows: LUAD cell lines [sample type], SARS-CoV-2 [virus infection], RNA-seq data [data type], and p-value was set as 0.05.
Statistical analysis
A two-tailed Student’s test was used to analyze the expression data in different clinicopathological groups and SARS-CoV-2 infected groups. Overall survival (OS), progression-free survival (PFS), and post-progression survival (PPS) were determined using Kaplan-Meier analysis and a log-rank test. Spearman’s correlation analysis was used to evaluate the linear relationship between the JAK1 mRNA expression levels and the immune cell infiltration level. All statistical tests and charts in this study were conducted using SPSS software (version 22.0; IBM Corp.). p-values <0.05 were considered statistically significant. All graphics were integrated and performed using GraphPad Prism 8.
Supplementary Materials
Author Contributions
JLC and YZ contributed to the study conception and design. HTL, LLL, HD, and LXY contributed to data extraction, collation, and analysis. JLC and YZ contributed to the draft writing, and YZ contributed to quality assessment and manuscript revision. All authors have read and approved the manuscript for publication.
Acknowledgments
All the work of this study was supported by the National Natural Science Foundation of China (No. 81860284, NO. 31960146). We thank the staffs of the databases for the development of tumor molecular biology and cytobiology.
Conflicts of Interest
The authors declare no conflicts of interest.
Funding
This study was supported by the National Natural Science Foundation of China (No. 81860284, NO. 31960146).
References
- 1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492 [PubMed]
- 2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018; 68:7–30. https://doi.org/10.3322/caac.21442 [PubMed]
- 3. Ma KY, Schonnesen AA, Brock A, Van Den Berg C, Eckhardt SG, Liu Z, Jiang N. Single-cell RNA sequencing of lung adenocarcinoma reveals heterogeneity of immune response-related genes. JCI Insight. 2019; 4:e121387. https://doi.org/10.1172/jci.insight.121387 [PubMed]
- 4. Coate LE, Shepherd FA. Maintenance therapy in advanced non-small cell lung cancer: evolution, tolerability and outcomes. Ther Adv Med Oncol. 2011; 3:139–57. https://doi.org/10.1177/1758834011399306 [PubMed]
- 5. Somasundaram A, Burns TF. The next generation of immunotherapy: keeping lung cancer in check. J Hematol Oncol. 2017; 10:87. https://doi.org/10.1186/s13045-017-0456-5 [PubMed]
- 6. Assi HI, Kamphorst AO, Moukalled NM, Ramalingam SS. Immune checkpoint inhibitors in advanced non-small cell lung cancer. Cancer. 2018; 124:248–61. https://doi.org/10.1002/cncr.31105 [PubMed]
- 7. Lynch TJ, Bondarenko I, Luft A, Serwatowski P, Barlesi F, Chacko R, Sebastian M, Neal J, Lu H, Cuillerot JM, Reck M. Ipilimumab in combination with paclitaxel and carboplatin as first-line treatment in stage IIIB/IV non-small-cell lung cancer: results from a randomized, double-blind, multicenter phase II study. J Clin Oncol. 2012; 30:2046–54. https://doi.org/10.1200/JCO.2011.38.4032 [PubMed]
- 8. Naylor EC, Desani JK, Chung PK. Targeted therapy and immunotherapy for lung cancer. Surg Oncol Clin N Am. 2016; 25:601–09. https://doi.org/10.1016/j.soc.2016.02.011 [PubMed]
- 9. Zhan J, Wang P, Li S, Song J, He H, Wang Y, Liu Z, Wang F, Bai H, Fang W, Du Q, Ye M, Chang Z, et al. HOXB13 networking with ABCG1/EZH2/slug mediates metastasis and confers resistance to cisplatin in lung adenocarcinoma patients. Theranostics. 2019; 9:2084–99. https://doi.org/10.7150/thno.29463 [PubMed]
- 10. Schwartz DM, Bonelli M, Gadina M, O’Shea JJ. Type I/II cytokines, JAKs, and new strategies for treating autoimmune diseases. Nat Rev Rheumatol. 2016; 12:25–36. https://doi.org/10.1038/nrrheum.2015.167 [PubMed]
- 11. Ghoreschi K, Laurence A, O’Shea JJ. Janus kinases in immune cell signaling. Immunol Rev. 2009; 228:273–87. https://doi.org/10.1111/j.1600-065X.2008.00754.x [PubMed]
- 12. Liu D, Huang Y, Zeng J, Chen B, Huang N, Guo N, Liu L, Xu H, Mo X, Li W. Down-regulation of JAK1 by RNA interference inhibits growth of the lung cancer cell line A549 and interferes with the PI3K/mTOR pathway. J Cancer Res Clin Oncol. 2011; 137:1629–40. https://doi.org/10.1007/s00432-011-1037-6 [PubMed]
- 13. Wehde BL, Rädler PD, Shrestha H, Johnson SJ, Triplett AA, Wagner KU. Janus kinase 1 plays a critical role in mammary cancer progression. Cell Rep. 2018; 25:2192–207.e5. https://doi.org/10.1016/j.celrep.2018.10.063 [PubMed]
- 14. Sexl V, Kovacic B, Piekorz R, Moriggl R, Stoiber D, Hoffmeyer A, Liebminger R, Kudlacek O, Weisz E, Rothammer K, Ihle JN. Jak1 deficiency leads to enhanced abelson-induced B-cell tumor formation. Blood. 2003; 101:4937–43. https://doi.org/10.1182/blood-2001-11-0142 [PubMed]
- 15. Xie HJ, Bae HJ, Noh JH, Eun JW, Kim JK, Jung KH, Ryu JC, Ahn YM, Kim SY, Lee SH, Yoo NJ, Lee JY, Park WS, Nam SW. Mutational analysis of JAK1 gene in human hepatocellular carcinoma. Neoplasma. 2009; 56:136–40. https://doi.org/10.4149/neo_2009_02_136 [PubMed]
- 16. Jeong EG, Kim MS, Nam HK, Min CK, Lee S, Chung YJ, Yoo NJ, Lee SH. Somatic mutations of JAK1 and JAK3 in acute leukemias and solid cancers. Clin Cancer Res. 2008; 14:3716–21. https://doi.org/10.1158/1078-0432.CCR-07-4839 [PubMed]
- 17. Owen KL, Brockwell NK, Parker BS. JAK-STAT signaling: a double-edged sword of immune regulation and cancer progression. Cancers (Basel). 2019; 11:2002. https://doi.org/10.3390/cancers11122002 [PubMed]
- 18. Zhang X, Zeng Y, Qu Q, Zhu J, Liu Z, Ning W, Zeng H, Zhang N, Du W, Chen C, Huang JA. PD-L1 induced by IFN-γ from tumor-associated macrophages via the JAK/STAT3 and PI3K/AKT signaling pathways promoted progression of lung cancer. Int J Clin Oncol. 2017; 22:1026–33. https://doi.org/10.1007/s10147-017-1161-7 [PubMed]
- 19. Lu C, Talukder A, Savage NM, Singh N, Liu K. JAK-STAT-mediated chronic inflammation impairs cytotoxic T lymphocyte activation to decrease anti-PD-1 immunotherapy efficacy in pancreatic cancer. Oncoimmunology. 2017; 6:e1291106. https://doi.org/10.1080/2162402X.2017.1291106 [PubMed]
- 20. Chen B, Lai J, Dai D, Chen R, Li X, Liao N. JAK1 as a prognostic marker and its correlation with immune infiltrates in breast cancer. Aging (Albany NY). 2019; 11:11124–35. https://doi.org/10.18632/aging.102514 [PubMed]
- 21. Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y, Ren R, Leung KS, Lau EH, Wong JY, Xing X, Xiang N, Wu Y, et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N Engl J Med. 2020; 382:1199–207. https://doi.org/10.1056/NEJMoa2001316 [PubMed]
- 22. Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020; 395:565–74. https://doi.org/10.1016/S0140-6736(20)30251-8 [PubMed]
- 23. Burki TK. Coronavirus in China. Lancet Respir Med. 2020; 8:238. https://doi.org/10.1016/S2213-2600(20)30056-4 [PubMed]
- 24. COVID-19 Coronavirus pandemic. [cited 2020 JUNE 11]: https://www.worldometers.info/coronavirus/.
- 25. Liang W, Guan W, Chen R, Wang W, Li J, Xu K, Li C, Ai Q, Lu W, Liang H, Li S, He J. Cancer patients in SARS-CoV-2 infection: a nationwide analysis in China. Lancet Oncol. 2020; 21:335–37. https://doi.org/10.1016/S1470-2045(20)30096-6 [PubMed]
- 26. Rogado J, Pangua C, Serrano-Montero G, Obispo B, Marino AM, Pérez-Pérez M, López-Alfonso A, Gullón P, Lara MÁ. Covid-19 and lung cancer: a greater fatality rate? Lung Cancer. 2020; 146:19–22. https://doi.org/10.1016/j.lungcan.2020.05.034 [PubMed]
- 27. Yang L, Xue H, Sun Y, Zhang L, Xue F, Ge R. CircularRNA-9119 protects hepatocellular carcinoma cells from apoptosis by intercepting miR-26a/JAK1/STAT3 signaling. Cell Death Dis. 2020; 11:605. https://doi.org/10.1038/s41419-020-02807-0 [PubMed]
- 28. Simonian M, Mosallayi M, Mirzaei H. Circulating miR-21 as novel biomarker in gastric cancer: diagnostic and prognostic biomarker. J Cancer Res Ther. 2018; 14:475. https://doi.org/10.4103/0973-1482.175428 [PubMed]
- 29. Robertson KD. DNA methylation and human disease. Nat Rev Genet. 2005; 6:597–610. https://doi.org/10.1038/nrg1655 [PubMed]
- 30. Sansone P, Bromberg J. Targeting the interleukin-6/Jak/stat pathway in human Malignancies. J Clin Oncol. 2012; 30:1005–14. https://doi.org/10.1200/JCO.2010.31.8907 [PubMed]
- 31. Xiong H, Zhang ZG, Tian XQ, Sun DF, Liang QC, Zhang YJ, Lu R, Chen YX, Fang JY. Inhibition of JAK1, 2/STAT3 signaling induces apoptosis, cell cycle arrest, and reduces tumor cell invasion in colorectal cancer cells. Neoplasia. 2008; 10:287–97. https://doi.org/10.1593/neo.07971 [PubMed]
- 32. Yu L, Chen W, Tang Q, Ji KY. Micheliolide inhibits liver cancer cell growth via inducing apoptosis and perturbing actin cytoskeleton. Cancer Manag Res. 2019; 11:9203–12. https://doi.org/10.2147/CMAR.S216870 [PubMed]
- 33. Yamaguchi H, Condeelis J. Regulation of the actin cytoskeleton in cancer cell migration and invasion. Biochim Biophys Acta. 2007; 1773:642–52. https://doi.org/10.1016/j.bbamcr.2006.07.001 [PubMed]
- 34. Sica A, Mantovani A. Macrophage plasticity and polarization: in vivo veritas. J Clin Invest. 2012; 122:787–95. https://doi.org/10.1172/JCI59643 [PubMed]
- 35. Ye Y, Xu Y, Lai Y, He W, Li Y, Wang R, Luo X, Chen R, Chen T. Long non-coding RNA cox-2 prevents immune evasion and metastasis of hepatocellular carcinoma by altering M1/M2 macrophage polarization. J Cell Biochem. 2018; 119:2951–63. https://doi.org/10.1002/jcb.26509 [PubMed]
- 36. Ma J, Liu L, Che G, Yu N, Dai F, You Z. The M1 form of tumor-associated macrophages in non-small cell lung cancer is positively associated with survival time. BMC Cancer. 2010; 10:112. https://doi.org/10.1186/1471-2407-10-112 [PubMed]
- 37. Chen G, Wu D, Guo W, Cao Y, Huang D, Wang H, Wang T, Zhang X, Chen H, Yu H, Zhang X, Zhang M, Wu S, et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J Clin Invest. 2020; 130:2620–29. https://doi.org/10.1172/JCI137244 [PubMed]
- 38. Farrar MA, Schreiber RD. The molecular cell biology of interferon-gamma and its receptor. Annu Rev Immunol. 1993; 11:571–611. https://doi.org/10.1146/annurev.iy.11.040193.003035 [PubMed]
- 39. Shi H, Wang W, Yin J, Ouyang Y, Pang L, Feng Y, Qiao L, Guo X, Shi H, Jin R, Chen D. The inhibition of IL-2/IL-2R gives rise to CD8+ T cell and lymphocyte decrease through JAK1-STAT5 in critical patients with COVID-19 pneumonia. Cell Death Dis. 2020; 11:429. https://doi.org/10.1038/s41419-020-2636-4 [PubMed]
- 40. Fu X, Xu M, Song Y, Li Y, Zhang H, Zhang J, Zhang C. Enhanced interaction between SEC2 mutant and TCR vβ induces MHC II-independent activation of T cells via PKCθ/NF-κB and IL-2R/STAT5 signaling pathways. J Biol Chem. 2018; 293:19771–84. https://doi.org/10.1074/jbc.RA118.003668 [PubMed]
- 41. Wan SX, Yi QJ, Fan SB, Lv JL, Zhang XX, Guo L, Lang CH, Xiao Q, Xiao KH, Yi ZJ, Qiang M, Xiang JL, Zhang BS, Chen YP. Characteristics of lymphocyte subsets and cytokines in peripheral blood of 123 hospitalized patients with 2019 novel coronavirus pneumonia (NCP). medRxiv. 2020. [Epub ahead of print]. https://doi.org/10.1101/2020.02.10.20021832
- 42. Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ, Kincead-Beal C, Kulkarni P, Varambally S, Ghosh D, Chinnaiyan AM. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007; 9:166–80. https://doi.org/10.1593/neo.07112 [PubMed]
- 43. Chandrashekar DS, Bashel B, Balasubramanya SA, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BV, Varambally S. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017; 19:649–58. https://doi.org/10.1016/j.neo.2017.05.002 [PubMed]
- 44. Nagy Á, Lánczky A, Menyhárt O, Győrffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018; 8:9227. https://doi.org/10.1038/s41598-018-27521-y [PubMed]
- 45. Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q, Li B, Liu XS. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020; 48:W509–14. https://doi.org/10.1093/nar/gkaa407 [PubMed]
- 46. Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015; 12:453–57. https://doi.org/10.1038/nmeth.3337 [PubMed]


