Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), a member of β-Coronavirus (CoV) lineage B causing the outbreak of coronavirus disease 2019 (COVID-19), was first identified and reported in Wuhan, China in December, 2019 [1, 2]. SARS-CoV-2 has affected the world to varying degrees to become a major cause of significant morbidity and mortality globally, and the outbreak of COVID-19 has been declared as a global pandemic by WHO on March 11, 2020 [3]. As of Feb 25th, 2021, the total number of COVID-19 patients globally reported to WHO had reached 112,209,815 of confirmed cases and 2,490,776 deaths in 223 countries, areas or territories. The clinical and epidemiological features of patients with COVID-19 demonstrate that most first-episode patients are asymptomatic or experience mild to severe respiratory illness [4], and older men with comorbidities were more susceptible to infection and associated with higher disease severity and mortality that might due to their weaker immune functions [5, 6]. Besides, SARS-CoV-2 infection can cause multi-organ failure and systemic manifestations especially in critical illness [7].
The laboratory studies have shown that data of both lymphocyte subsets and cytokine profiles are crucial for diagnosis and treatment of patients with COVID-19. The majority of patients are reported to have lymphopenia, while the most available biomarkers of routine clinical tests including C-reactive protein (CRP), neutrophils and lymphocytes ratio (NLR), lactate dehydrogenase and D-dimer were associated with disease severity and could contribute in predicting clinical outcomes in COVID-19 patients [8, 9]. Consistently with SARS and MERS-CoV infections [10, 11], several studies have shown that pro-inflammatory cytokine profiles elevated in patients with COVID-19, and that cytokine storm are directly correlated with pulmonary inflammation, tissue injury and poor prognosis of severe cases [4, 12]. Higher levels of cytokine storm could aggravate the progression of illness in severe COVID-19 patients [7, 13]. Besides, previous studies demonstrated that interleukin-2R (IL-2R)/lymphocyte, interleukin-6 (IL-6) and interleukin-10 (IL-10) might be the potential biomarkers for early diagnosis and indicators for higher risk of disease deterioration when predicting the disease progression of COVID-19 [3, 4, 14]. As for lymphocyte subsets, circulating levels of lymphocyte subsets were suggested to be correlated with clinical features and treatment efficacy of SARS-CoV-2 infection which mainly involve T lymphocytes (particularly CD4+ and CD8+ T cells), and CD8+ T cells alone might be an independent biomarker for the disease severity and treatment efficacy [5, 12, 15]. Therefore, lymphocyte subsets and cytokine profiles seem to be promising for evaluating the disease severity of COVID-19 patients and providing reference for clinical intervention. However, evidence so far has shown inconsistency. A longitudinal analysis of lymphocyte subsets and cytokine profiles in 40 confirmed COVID-19 patients has showed that levels of cytokine profiles and counts of T cells in survivors of severe COVID-19 would return to normal levels comparable with mild patients at a later stage [16]. However, another study which included 25 confirmed COVID-19 patients who were admitted to ICU showed that circulating levels of IL-2, interleukin-4 (IL-4), tumor necrosis factor-α (TNF-α), interferon (IFN-γ) and CRP were not directly correlated with symptom severity of COVID-19 [17]. Thus, the pathophysiology in SARS-CoV-2 infection, especially the correlation of cytokine profiles and immunological features with different disease severity in patients with COVID-19, has not yet been completely understood, and available biomarkers for predicting the disease progression of COVID-19 patients are still limited.
In this study, we performed a comprehensive evaluation of features of 254 COVID-19 patients for the differences in laboratory biomarkers between severe or critical and non-severe patients, including inflammatory markers, cytokine profiles and immunological features. Besides, we developed two novel nomograms for the prediction of disease severity using inflammatory cytokine profiles or lymphocyte subsets data separately. These findings may help us extend our understanding of potential biomarkers associated with disease severity for early identification and guidance of treatment management.
Results
Demographic and baseline clinical characteristics of patients with disease severity in COVID-19
A total of 254 patients with confirmed SARS-Cov-2 infection were included in the study and classified into two groups by disease severity. 90 (35.4%) of them were severe or critical patients, and 164 (64.6%) were non-severe patients. Demographic and clinical characteristics of severe or critical and non-severe patients are shown in Table 1. The median age of severe or critical patients (70 years, IQR (64–77)) was significantly higher than non-severe patients (65 years, IQR (55–72)) (P < 0.001). Males in severe or critical patients were markedly more common than females (P < 0.05). 58.7% of severe or critical patients had at least one comorbidity, with hypertension and diabetes being the most common ones. Median hospitalization time duration for severe or critical patients was 17.5 days (IQR 6.8–29.2) and 23 days (IQR 15–28.8) for non-severe patients (Table 1).
Table 1. Differences of clinical characteristics and laboratory findings of patients with COVID-19 on admission.
| Clinical Characteristics | Normal Range | Median (IQR) or n/N (%) | P value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Non-severe (164) | Severe or Critical (90) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Age, y | 65 (55–72) | 70 (64–77) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sex, n (%) | 0.006 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Female | 91 (55.5%) | 33 (36.7%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Male | 73 (44.5%) | 57 (63.3%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Comorbidities, n/N (%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Any comorbidities | 21/38 (55.3%) | 37/63 (58.7%) | 0.836 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hypertension | 15/38 (39.5%) | 25/63 (39.7%) | 0.983 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diabetes | 4/38 (10.5%) | 16/63 (25.4%) | 0.078 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms, n/N (%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Fever | 33/38 (86.8%) | 53/63 (84.1%) | 0.780 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cough | 33/38 (86.8%) | 53/63 (84.1%) | 0.780 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Expectoration | 26/38 (68.4%) | 39/63 (61.9%) | 0.529 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Dyspnea | 25/38 (65.8%) | 41/63 (66.1%) | 0.972 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Myalgia or fatigue | 28/38 (73.7%) | 36/63 (57.1%) | 0.135 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chest tightness | 19/39 (48.7%) | 36/63 (57.1%) | 0.422 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Diarrhea | 18/38 (47.4%) | 28/63 (44.4%) | 0.838 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Nausea or vomiting | 17/38 (44.7%) | 13/63 (20.6%) | 0.014 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Headache | 21/38 (55.3%) | 21/63 (33.3%) | 0.038 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abdominal pain | 17/38 (44.7%) | 11/63 (17.5%) | 0.005 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Muscle soreness | 18/38 (47.4%) | 23/63 (36.5%) | 0.303 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sore throat | 13/38 (34.2%) | 17/63 (27.0%) | 0.503 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hemoptysis | 12/38 (30.8%) | 16/63 (25.4%) | 0.649 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Hospitalization period, d | 23 (15–28.8) | 17.5 (6.8–29.2) | 0.013 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Blood routine | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Leucocytes, × 109/L | 3.5–9.5 | 5.66 (4.51–6.93) | 9.26 (6.06–12.75) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased | 12 (7.3%) | 44 (48.9%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased | 9 (5.5%) | 5 (5.6%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Neutrophils, × 109/L | 1.8–6.3 | 3.84 (2.64–4.86) | 7.94 (4.83–11.70) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased | 20 (12.2%) | 55 (61.1%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased | 8 (4.9%) | 2 (2.2%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lymphocytes, × 109/L, | 1.1–3.2 | 1.16 (0.85–1.55) | 0.64 (0.44–0.87) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased | 75 (45.7%) | 76 (84.4%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NLR | 3.2 (2.0–5.2) | 11.5 (6.7–20.7) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased | 23 (14.0%) | 1 (1.1%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased | 34 (20.7%) | 72 (80.0%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monocyte, × 109/L | 0.1–0.6 | 0.51 (0.41–0.63) | 0.44 (0.28-0.64) | 0.024 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Platelets, mean (SD), × 109/L | 125–350 | 252 ± 95 | 181 ± 96 | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased | 27 (16.5%) | 4 (4.4%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased | 11 (6.7%) | 27 (30.0%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Infectious biomarkers | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PCT, ng/mL | 0.02–0.05 | 0.04 (0.03–0.08) | 0.16 (0.09–0.56) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ESR, mm/h | 0–20 mm/H | 41.0 (20.0–64.0) | 36 (17.0–64.2) | 0.610 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| cTnI, pg/mL | ≤15.6 | 4.0 (2.0–9.5) | 43.6 (6.6–305.6) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NT-proBNP, pg/mL | <738 | 174 (66–377) | 746 (229.5–2302) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CRP, mg/L | <1 | 10.6 (1.9–40.3) | 86.3 (43.6–174) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased | 130 (79.3%) | 86 (95.6%) | 0.007 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Organ Function | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ALT, U/mL | ≤ 33 | 23 (14–40) | 29.5 (18.0–44.2) | 0.011 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| AST, U/mL | ≤ 32 | 24 (18–33.8) | 42 (23.8–62.8) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TP, g/L | 64–83 | 69.9 (65.9–73.6) | 65.3 (59.5–70.4) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Albumin, g/L | 35–53 | 35.4 (32.4–38.7) | 30.8 (27.7–33.1) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Globulin, U/mL | 25–35 | 33.7 (30.5–37.8) | 34.8 (30.6–39.2) | 0.411 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Glu, mmol/L | 4.11–6.05 | 5.72 (5.13–6.94) | 7.37 (5.9–10.4) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TBIL, umol/L | ≤ 2 | 9.2 (7.0–13.2) | 13.4 (9.2–18.4) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DBIL, umol/L | ≤ 8 | 3.9 (3.0–5.3) | 6.2 (4.1–9.9) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IBIL, umol/L | ≤ 12.9 | 5.1 (3.9–7.8) | 6.2 (4.5–9.1) | 0.037 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ALP, U/L | 35–105 | 66.5 (54–86) | 79.5 (58–105) | 0.002 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GGT, U/L | 6–42 | 27.5 (18–46.8) | 36.5 (20.8–76.2) | 0.013 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TC, mmol/L | <5.18 | 3.8 (3.3–4.5) | 3.4 (2.9–4.3) | 0.006 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LDH, U/L | 135–214 | 258 (214–329.8) | 480 (325–652) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mb, ng/mL | ≤ 106 | 43.6 (29.8–81.8) | 117.1 (60.5–290.2) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CK-MB, ng/mL | ≤ 3.4 | 0.7 (0.4–1.2) | 1.8 (0.9–5.8) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creatinine, umol/L | 45–85 | 70 (59–84) | 85 (67–105.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Coagulation function | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PT, s | 11.5–14.5 | 13.7 (13.2–14.2) | 15.3 (14.2–16.7) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| INR | 0.8–1.2 | 1.04 (0.99–1.09) | 1.20 (1.10–1.34) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Fibrinogen, g/L | 2–4 | 5.05 (3.65–6.02) | 4.56 (3.10–6.13) | 0.175 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| APTT, s | 29–42 | 38.6 (36.3–42.3) | 40.1 (35.6–46.2) | 0.209 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| D-dimer, U/mL FEU | <0.5 | 1.05 (0.40–1.91) | 4.60 (1.44–21.00) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Urine routine | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Urine protein, n/N (%) | – | 61/149 (40.9%) | 62/76 (81.6%) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| USG, n, median (IQR) | 1.01–1.025 | 149, 1.015 (1.011–1.019) | 76, 1.02 (1.016–1.024) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| pH, n, median (IQR) | 4.5–8.0 | 149, 6.5 (6.0–7.0) | 76, 6.0 (5.5–6.5) | 0.002 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KET, n/N (%) | – | 11/149 (7.4%) | 26/76 (34.2%) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| URO, n/N (%) | – | 2/149 (1.3%) | 10/76 (13.2%) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Arterial Blood Gas | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| pH, n, median (IQR) | 7.35–7.45 | 40, 7.41 (7.39–7.46) | 37, 7.42 (7.38–7.46) | 0.984 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PaCO2, n, mmHg | 35–45 | 39, 40.9 (38.3–43.8) | 37, 36.1 (29.9–41.0) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PaO2, n, mmHg | 80–100 | 39, 142 (90.8–187) | 37, 84 (57.8–148.0) | 0.004 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SB, n, mmol/L | 21–25 | 38, 25.8 (24.8–27.9) | 37, 23.8 (21.2–25.4) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BEecf, n, mmol/L | –3–3 | 39, 2.1 (0.3–3.9) | 37, –0.9 (–4.2–1.3) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TCO2, n mmol/L | 24–32 | 38, 23.2 (21.7–24.6) | 37, 20.9 (16.5–23.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SaO2, n, median (IQR) | 91.9–99 % | 39, 99.2 (97.2–99.6) | 37, 96.2 (86.8–99.5) | 0.011 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviations: NLR: Neutrophil-to-lymphocyte ratio; PCT: Procalcitonin; ESR: Erythrocyte sedimentation rate; cTnI: Cardiac troponin I; NT-proBNP: N-terminal of the prohormone brain natriuretic peptide; CRP: C-reactive protein; ALT: Alanine aminotransferase; AST: Aspartate aminotransferase; TP: Total protein; Glu: Glucose; TBIL: Total bilirubin; DBIL: Direct bilirubin; IBIL: Indirect bilirubin; ALP: Alkaline phosphatase; GGT: γ-glutamyl transpeptidase; TC: Total cholesterol; LDH: Lactate dehydrogenase; Mb: Myoglobin; CK-MB: Creatine kinase-MB; PT: Prothrombin time; INR: International normalized ratio; APTT: Activated partial thromboplastin time; USG: Urine specific gravity; KET: Urine ketone body; URO: Urobilinogen; PaCO2: Partial pressure of carbon dioxide; PaO2: Arterial partial pressure of oxygen; SB: Standard bicarbonate; BEecf: Base excess of extracellular fluid; TCO2: Total carbon dioxide; SaO2: Oxygen saturation. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Differences of laboratory findings between severe or critical and non-severe COVID-19
Table 1 showed general data of indicators as the patients were divided into severe or critical group and non-severe group, which included indicators from blood and urine routine examinations and arterial blood gas analysis, and biomarkers related to organ function, coagulation function and infection. Severe or critical patients presented significantly higher leukocytes and neutrophils counts, elevated neutrophil-to-lymphocyte ratio (NLR) than non-severe patients (all P < 0.001), as well as decreased counts of lymphocytes, monocytes and platelets (P < 0.05). Compared with non-severe patients, the levels of several infection-related biomarkers were significantly higher in severe or critical patients, including procalcitonin, cardiac troponin I (cTnI), N-terminal prohormone of brain natriuretic peptide (NT-proBNP), C-reactive protein (CRP) (all P < 0.001). Also, severe or critical patients had markedly higher levels of serum biochemical indexes including alanine aminotransferase (ALT), aspartate aminotransferase (AST), glucose (Glu), total bilirubin (TBIL), alkaline phosphatase (ALP), γ-glutamyl transpeptidase (GGT), lactate dehydrogenase (LDH), myoglobin, creatine kinase-MB (CK-MB), and creatinine. And they had significantly lower levels of total protein (TP), albumin, total cholesterol (TC) (all P < 0.05). Levels of some coagulation factors were significantly elevated in severe or critical cases compared with non-severe patients, including prothrombin time (PT) and D-dimer (all P < 0.001).
Compared with non-severe cases, the positive rates of urine protein, urine ketone body (KET) and urobilinogen (URO) in severe or critical cases were significantly higher (all P < 0.001). Severe or critical patients almost had significantly lower levels of all indicators in blood gas analysis, including partial pressure of carbon dioxide (PaCO2), arterial partial pressure of oxygen (PaO2), standard bicarbonate (SB), base excess of extracellular fluid (BE), total carbon dioxide (TCO2) and oxygen saturation (SaO2) (all P < 0.05) (Table 1).
Differences of immunologic features in disease severity of COVID-19
Table 2 showed inflammatory cytokine profiles and lymphocyte subsets in severe or critical patients and non-severe patients with COVID-19. We included interleukin-1β (IL-1β), interleukin-2R (IL-2R), interleukin-6 (IL-6), interleukin-8 (IL-8), interleukin-10 (IL-10) and tumor necrosis factor α (TNF-α) in cytokine profiles. Except for IL-1β which was undetectable (< 5 pg/mL), levels of all other cytokines were significantly increased in severe or critical patients compared with their normal ranges, and markedly higher than those in non-severe patients (all P < 0.001) (Table 2, Figure 1). Meanwhile, all lymphocyte subset counts including T, B, Th, Ts and NK cells were significantly decreased to be markedly lower in severe or critical cases (all P < 0.001) (Table 2, Figure 2). Nevertheless, there is no statistical significance in Th/Ts ratio between severe or critical and non-severe patients (P > 0.05).
Table 2. Differences of immune response in patients with COVID-19 on admission.
| Biomarker | Normal Range | N, Median (IQR) | P value | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Non-severe (164) | Severe or Critical (90) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Inflammatory cytokines | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IL-1β, n, pg/mL | <5 | 164, 5 (5–5) | 90, 5 (5–5) | 0.742 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IL-2R, n, U/mL | 223–710 | 164, 498 (318.5–750.8) | 90, 1111.5 (728.2–1656.0) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased, n (%) | 17 (10.4%) | 2 (2.2%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased, n (%) | 101 (61.6%) | 19 (21.1%) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IL-6, n, pg/mL | <7 | 164, 4.0 (1.5–13.0) | 90, 42.2 (12.9–96.3) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased, n (%) | 59 (36.0%) | 77 (85.6%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IL-8, n, pg/mL | <62 | 164, 7.8 (5.0–16.3) | 90, 22.9 (11.8–49.2) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased, n (%) | 2 (1.2%) | 18 (20.0%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IL-10, n, pg/mL | <9.1 | 164, 5.0 (5.0–5.0) | 90, 6.6 (5–13.4) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased, n (%) | 10 (6.1%) | 39 (43.3%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| TNF-α, n, pg/mL | <8.1 | 164, 6.6 (4.5–9.4) | 90, 9.8 (7.1–14.1) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased, n (%) | 44 (26.8%) | 49 (54.4%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lymphocyte Subsets | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| T cells + B cells + NK cells, n/μL | – – | 96, 1435 (1145.5–1718.8) | 41, 736 (332–1160.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| T cells (CD3+ CD19–), n/μL | 955–2860 | 96, 1005.5 (794–1231) | 41, 359 (198.5–804.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased, n/N (%) | 37/96 (38.5%) | 34/41 (82.9%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| B cells (CD3– CD19+), n/μL | 90–560 | 96, 177 (118.5–256) | 41, 84 (40–154) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased, n/N (%) | 17/96 (17.7%) | 22/41 (53.7%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Th cells (CD3+ CD4+), n/μL | 550–1440 | 96, 632 (509–809.5) | 41, 240 (102.5-552.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased, n/N (%) | 34/96 (35.4%) | 31/41 (75.6%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ts cells (CD3+ CD8+), n/μL | 320–1250 | 96, 314 (237–418) | 41, 128 (47–255.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased, n/N (%) | 49/96 (51.0%) | 36/41 (87.8%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NK cells (CD3–/CD16+ CD56+), n/μL | 150–1100 | 96, 190 (128–257) | 41, 60 (30.5–186.5) | <0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Decreased, n/N (%) | 36/96 (37.5%) | 28/41 (68.3%) | <0.001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Th/Ts ratio | 0.71–2.78 | 96, 2.04 (1.49–2.59) | 41, 2.29 (1.58–3.55) | 0.130 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Increased, n/N (%) | 16/96 (16.7%) | 13/41 (31.7%) | 0.137 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abbreviations: IL: Interleukin; TNF: Tumor necrosis factor; Th: helper T cells; Ts: suppressor T cells. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
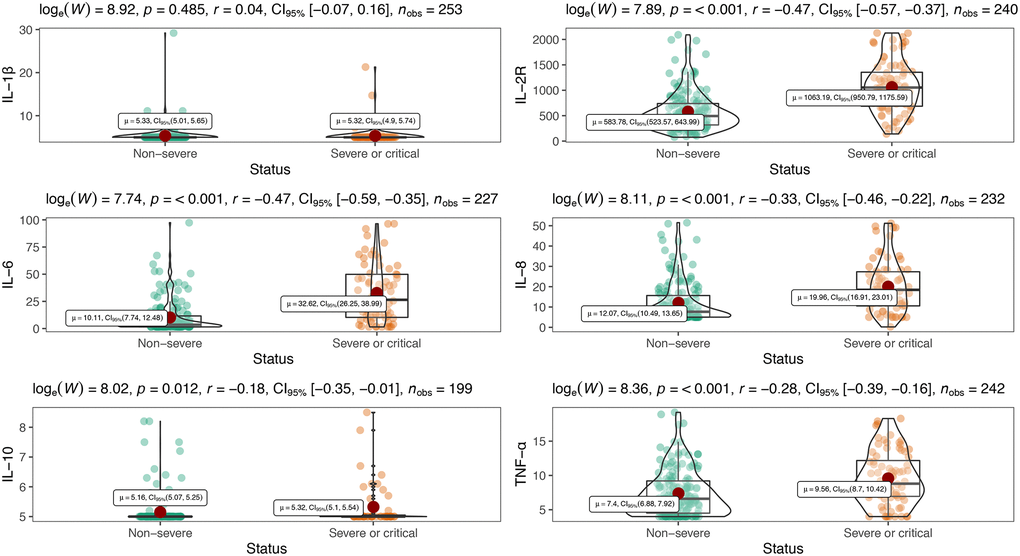
Figure 1. Distribution of peripheral inflammatory cytokines in severe and non-severe patients.
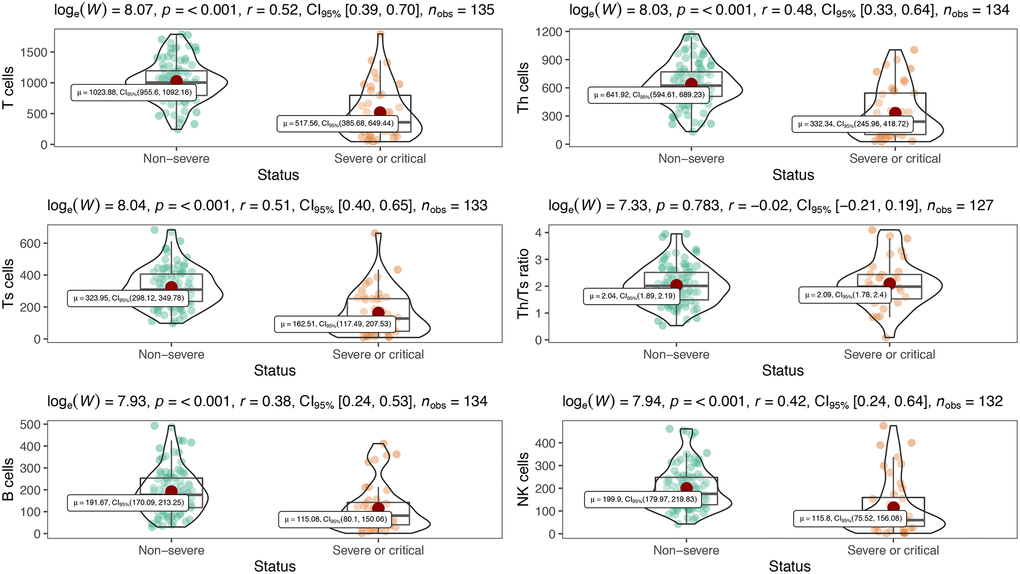
Figure 2. Peripheral lymphocyte subsets between severe and non-severe.
Nomograms development and validation for predicting the disease severity of COVID-19
Considering that there was the strong multicollinearity between inflammatory cytokine profiles and lymphocyte subsets, we established two logistic regression models based on inflammatory cytokine profiles and lymphocyte subsets separately. Multivariable analysis models showed that age, IL-2R, IL-6, IL-8, IL-10 and T cells could be the independent risk factors for disease severity (Supplementary Tables 1 and 2). Therefore, we also established two nomograms for predicting the disease severity of COVID-19 based upon the factors screened by the multivariable analyses (Figures 4 and 5), which were used to estimate the probability of higher disease severity by projecting the total score to the lower risk scale.
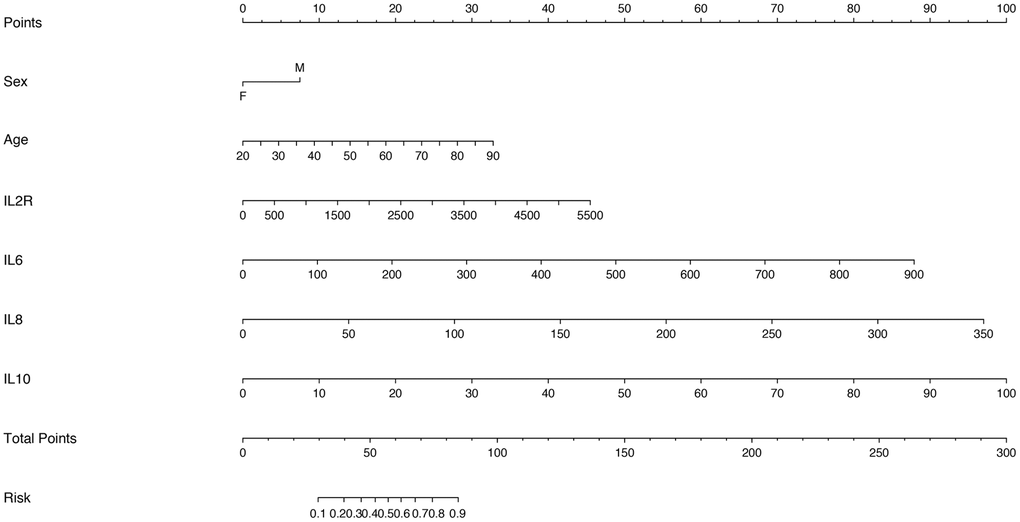
Figure 4. Nomogram constructed by using inflammatory cytokines for prediction of disease severity of COVID-19.
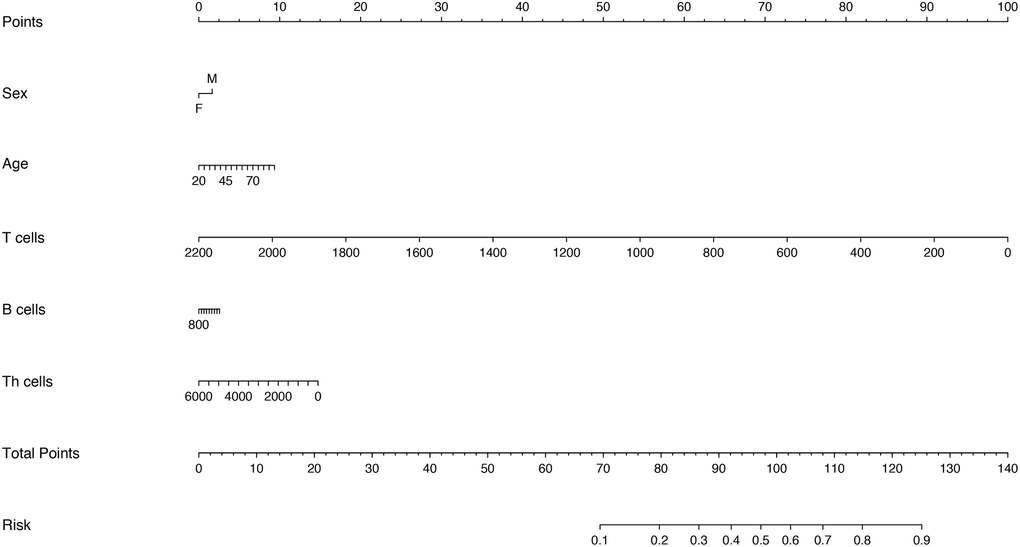
Figure 5. Nomogram constructed by using lymphocyte subsets for prediction of disease severity of COVID-19.
ROC curves showed that AUROCs for the two constructed diagnostic nomograms were 0.834 (95% CI: 0.779–0.888) and 0.841 (95% CI: 0.756–0.925) (Figure 6A, Figure 7A), and the nomograms both exhibited robust discrimination ability. The bootstrapped-concordance indexes of two nomograms were 0.834 and 0.841. The calibration curves of two nomograms (Figure 6B and Figure 7B) demonstrated that they could lead to comparable prediction results for the probability of disease severity that were also quite similar to the actual probability. Besides, the two models were well calibrated with Hosmer-Lemeshow goodness-of-fit results of 5.49 (p = 0.703) and 14.69 (p = 0.065). Decision curve analyses (DCA) results showed that the two nomograms provide more benefits for practical application (Figure 6C and Figure 7C). In the validation set, the bootstrapped-concordance indexes of two nomograms were 0.860 (0.802–0.919) and 0.852 (0.757–0.946). The calibration curves of two nomograms produced similar results between the predicted probability of disease severity and the actual probability, and DCA results validated the clinical usefulness of the nomograms (Figure 6D–6F and Figure 7D–7F).
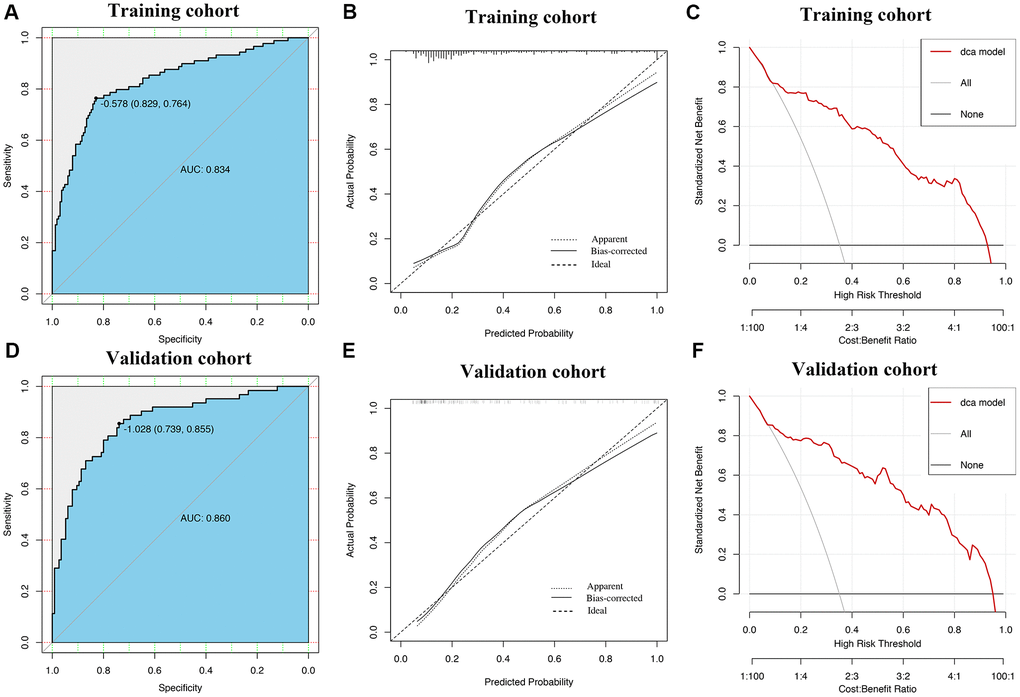
Figure 6. Construction and validation of the nomogram by using inflammatory cytokines. The area under receiver operating characteristic curve values, the calibration curve and decision curve analysis in training set (A–C) and in validation set (D–F) are shown.
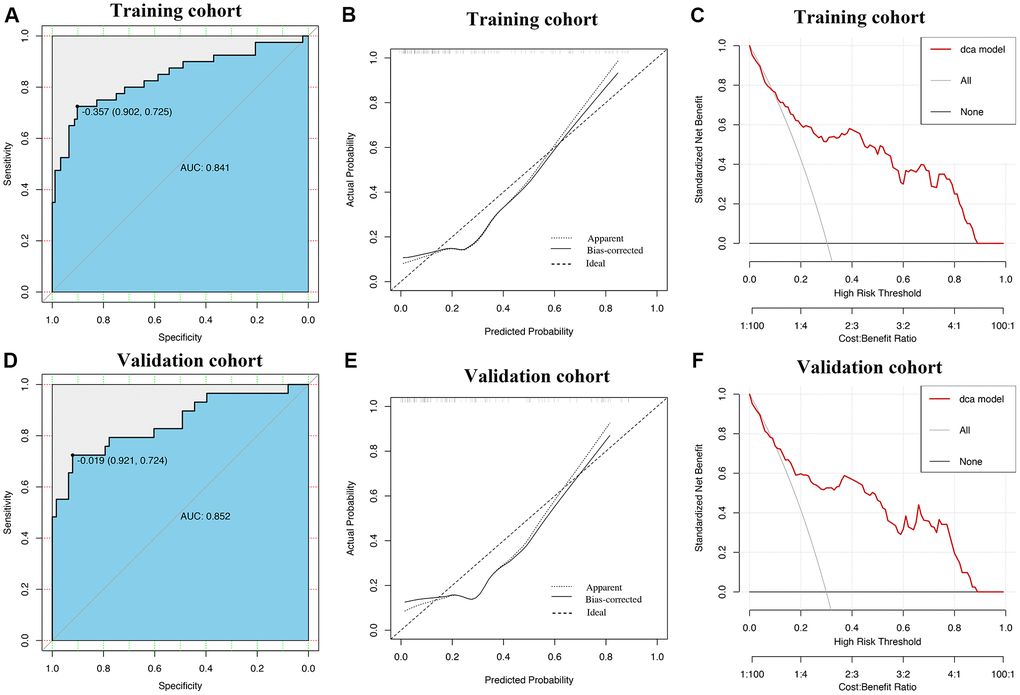
Figure 7. Construction and validation of the nomogram by using lymphocyte subsets. The area under receiver operating characteristic curve values, the calibration curve and decision curve analysis in training set (A–C) and in validation set (D–F) are shown.
Discussion
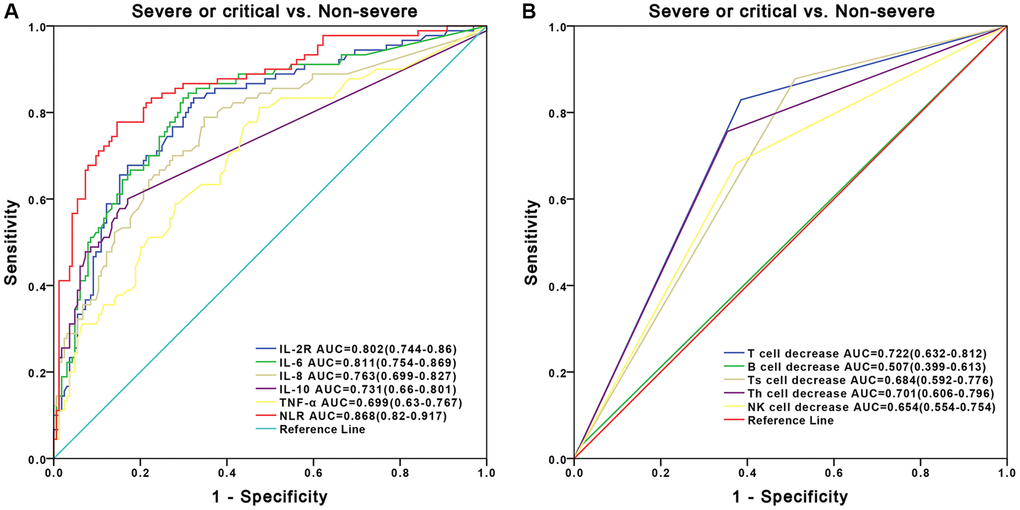
Both clinical and immunologic features of patients with COVID-19 have been reported recently [18–21], but how we can make use of these data in differential diagnosis for severe or critical COVID-19 patients still remains a challenge. Besides, there is still insufficient knowledge regarding the immunological indicators related to the mechanism and disease severity of COVID-19. Here, our study on COVID-19 patients at severe or critical and non-severe stages have mainly demonstrated the following findings. First, older males are more susceptible to SARS-CoV-2 infection than females, which was consistent with previous studies [6, 22–25]. Second, the levels of inflammatory cytokines were significantly higher in severe or critical patients than those in non-severe patients while the levels of lymphocyte subsets were significantly lower. Finally, two newly-developed diagnostic nomograms based on inflammatory cytokine profiles and lymphocyte subsets were separately constructed with AUROCs of 0.834 and 0.841. Both of them exhibited robust discriminative ability, which were well improved compared to using blood marker alone (Figure 3). The constructed nomograms could potentially be applied in clinic to differentially diagnose severe or critical and non-severe COVID-19 patients.
In terms of laboratory findings, lymphopenia and increased levels of NLR and infection-related biomarkers were the most common features in severe or critical patients compared to non-severe patients, which was similar to the results in SARS and MERS infections and consistent with previous studies as well [5]. Besides, we found levels of ALT, AST, Glu, TBIL, ALP, GGT, LDH, myoglobin, CK-MB, creatinine and coagulation factors were significantly higher in severe or critical patients than in non-severe patients. Significantly lower levels of TP, albumin, TC and blood gas indicators found in severe or critical patients were also consistent with previous studies [14, 18, 26, 27]. The phenomenon can be explained by the finding of previous study that cytokine storm induced by SARS-CoV-2 infection would cause hepatotoxicity and subsequently critical hypoalbuminemia, which are correlated with inflammatory responses and result in high disease severity and mortality of COVID-19 [26]. In terms of urine routine, the positive rates of urine protein, urine ketone body and urobilinogen in severe or critical cases were significantly higher than non-severe patients. Moreover, we observed that CRP and NLR were significantly positive correlated with Glu, CK, CK-MB, AST and LDH. CRP and NLR have been regarded as predictors of systemic inflammation, bacterial or viral infections and tissue injuries [12, 28, 29]. Recently, several studies have demonstrated that they might be useful potential biomarkers for advanced disease progression of COVID-19 while NLR can reflect the degree of imbalance in systemic inflammation and immune response [12, 16, 29]. In longitudinal analysis, NLR and neutrophil-to-CD8+ T cell ratio (N8R) have been indicated as important prognostic factors for severe COVID-19 [16]. It was suggested that monitoring laboratory markers and cytokines could be helpful in predicting the progression, improving treatment efficacy and reducing mortality of COVID-19 [3, 21, 29].
We noted that SARS-CoV-2 infection would induce significant rises in peripheral cytokine profiles and reductions in lymphocytes subsets. Levels of cytokine profiles were significantly higher in severe or critical patients who also presented markedly lower of lymphocyte subsets counts compared with non-severe patients, which were consistent with previous studies and similar to the findings in SARS and MERS [5, 15, 30–32]. Specifically, we also analyzed the correlations between these indexes and found that cytokine profiles were all significantly negatively correlated with lymphocytes subsets. Serum biochemical indexes such as Glu, CK, CK-MB, AST, and LDH as well as CRP and NLR were all significantly positively correlated with cytokine profiles but significantly negatively correlated with T, B, Th, Ts and NK cells. ALT was positively correlated with IL-2R, IL-6, NLR and CRP, while it had no correlations with IL-8, TNF-α or lymphocyte subsets. No correlations were observed between Glu and AST with Th/Ts ratio. Additionally, we found TC was positively associated with lymphocyte subsets but negatively associated with cytokine profiles, CRP, NLR and Th/Ts ratio.
It is believed that SARS-CoV-2 infection would induce cytokine storm and dysregulation of immune response, which is similar with SARS infection. SARS-CoV repaid replication and dysregulated type I interferon would cause elevated levels of cytokines and suboptimal T cell responses in SARS-CoV infected mice [33]. Several studies have established theories that IL-2R/lymphocyte, IL-6 and IL-10 can be used as predictors for early identification of disease and prediction of the risk of disease deterioration [3, 4, 34]. These cytokines can be produced at multiple sites of tissue inflammation and released into the circulation by various types of cells during sepsis and acute organ injuries [34, 35]. They may also play important roles in immune response during viral infections as higher levels of cytokine storm is reported to be correlated with more rapid disease progression, resulting in high incidence of immune disorders and mortality [15, 36, 37]. We found that SARS-CoV-2 infection induced evident lymphopenia and lowered the levels of lymphocyte subsets, which affects the maintenance of immune response and associated with disease severity, and these findings were in consistence with results of previous studies [15, 21, 38, 39]. SARS-CoV-2 infection mainly affects T lymphocytes especially CD4+ and CD8+ T cells, which were significantly decreased and associated with disease severity in COVID-19. CD8+ T cell was the lymphocyte subset with the most obvious decline in longitudinal analysis, which might contribute to inhibiting overactive innate immune responses [16, 18, 39]. However, the functions of B and NK cells are often ignored despite the finding that levels of them both decreased in patients. Previous studies suggested that B and NK cells participated in controlling SARS-CoV-2 infection and were associated with disease severity [15].
Besides, based upon the multivariate models using inflammatory cytokines and lymphocyte subsets, we developed two novel nomograms for differentially diagnosis of severe or critical and non-severe COVID-19 patients. Our results showed that the two models improved the diagnostic power compared with the power of models using each blood indicator alone, and AUROCs for the nomograms were greater than those of blood marker alone in the differentially diagnosis of disease severity (Figure 3). Furthermore, the two novel nomograms showed robust discriminative ability. Given the above findings, our study demonstrated that systemic inflammation responses, cytokine storms and dysregulated immune responses occurred and exerted combined effects on the clinical progression in COVID-19, which makes monitoring these biomarkers helpful for predicting the disease severity and reducing mortality of COVID-19.
There were still several limitations in our study. Firstly, a single-center, retrospective study using data from the designated hospital, which mainly receives severe patients in Wuhan, China might cause several potential biases. Secondly, there was insufficient information of possible temporal changes of cytokine profiles and immunological features in SARS-CoV-2 infected patients. Studies of larger sample sizes may better solve the problem. Thirdly, bacterial coinfection might exit in patients with COVID-19 and thus interference the manifestations of their immune responses. Fourth, the nomograms were constructed by retrospective analysis based on data from one single hospital. Despite there were internal validation, we still lacked data for external validation. Though the nomograms demonstrated good performance, they are preferred to be validated by performing a multiple-center prospective study.
Conclusions
In summary, blood inflammatory cytokines and lymphocyte subsets may be potential biomarkers for early screening, diagnosis and treatment of severe or critical COVID-19 patients. We proposed two novel nomograms based on inflammatory cytokines and lymphocyte subsets separately that can accurately predict disease severity of COVID-19. We hope that our novel nomograms may facilitate the identification of severe or critical patients and assist physicians in making optimized treatment suggestions.
Materials and Methods
Study design and participants
The study was approved by the Ethical Committee of Tongji Hospital of Huazhong University of Science and Technology (No.TJ-IRB20200364) and China-Japan Union Hospital of Jilin University (No.2020032607). All 254 patients with COVID-19 were enrolled in the study and admitted to Tongji Hospital of Huazhong University of Science and Technology from Jan 10, 2020 to Mar 11, 2020. Diagnosis, clinical classifications and complication definitions of patients with COVID-19 were made according to China’s Novel Coronavirus Pneumonia Diagnosis and Treatment Guidance (Seventh version) [40]. Patients with SARS-CoV-2 infection were confirmed by qRT-PCR test on swabs or sputum specimens on admission. Patients younger than 18-year or those with infection of HBV, HCV and HIV were excluded in the study.
Data collection of clinical and laboratory analysis
Demographic characteristics, clinical symptoms and laboratory data of the patients including inflammatory cytokine profiles and lymphocyte subsets analyses on admission were collected and extracted from their electronic medical records. All data were extracted with a unified data collection form and checked by two independent physicians.
Statistical analysis
SPSS 22.0 and R 3.6.1 software were used for statistical analysis. We presented count data as percentages, and measurement data were described as means (SD) or medians (IQR). Statistical differences were analyzed by Chi-square test or the Fisher exact test for count data, and Mann-Whitney U-test or unpaired t-test for continuous data. Spearman rank coefficient analyses were performed for determining correlations between the immunologic biomarkers and laboratory-related indexes. The diagnostic values of peripheral inflammatory cytokines and lymphocyte subsets for predicting the disease severity of COVID-19 were evaluated using area under the receiver operating characteristic (AUROC) curves. The potential risk factors of disease severity in COVID-19 patients were examined by multivariable logistic regression model. Before entering the models, the variables were screened by variance inflation factor (VIF) using ‘vif’ function of ‘car’ package, and only variables with VIFs < 5 entered the models. All data were used as training set for the development of the prediction nomograms, and the discrimination power of nomograms were evaluated by the concordance index, ROC curves and calibration plots with 1000 bootstrap resampling. Besides, the calibration of models was assessed by Hosmer-Lemeshow goodness-of-fit test with ‘ResourceSelection’ package. 70% of the original data were randomly selected as the internal validation set for evaluating the application efficacy of the prediction nomograms. P < 0.05 indicated statistical significance in all analysis.
Author Contributions
ZL, NJ, TL and QY designed the article. ZL, BY, MJ, YS, YH, YL, and YW collected and analyzed the data. ZL, NJ, XL and BY drafted of the manuscript. XL, DS, PM, JZ, TL and QY discussed and revised the manuscript. All authors reviewed and approved the final manuscript.
Acknowledgments
The authors would like to express our appreciation to all physicians for their effort for fighting COVID-19.
Conflicts of Interest
The authors declare no conflicts of interest related to this study.
Funding
No grants were received from any agencies from public, commercial or other sectors with respect to the research work.
References
- 1. Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, et al, and China Novel Coronavirus Investigating and Research Team. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020; 382:727–33. https://doi.org/10.1056/NEJMoa2001017 [PubMed]
- 2. Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020; 395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5 [PubMed]
- 3. Hou H, Zhang B, Huang H, Luo Y, Wu S, Tang G, Liu W, Mao L, Mao L, Wang F, Sun Z. Using IL-2R/lymphocytes for predicting the clinical progression of patients with COVID-19. Clin Exp Immunol. 2020; 201:76–84. https://doi.org/10.1111/cei.13450 [PubMed]
- 4. Han H, Ma Q, Li C, Liu R, Zhao L, Wang W, Zhang P, Liu X, Gao G, Liu F, Jiang Y, Cheng X, Zhu C, Xia Y. Profiling serum cytokines in COVID-19 patients reveals IL-6 and IL-10 are disease severity predictors. Emerg Microbes Infect. 2020; 9:1123–30. https://doi.org/10.1080/22221751.2020.1770129 [PubMed]
- 5. Chen G, Wu D, Guo W, Cao Y, Huang D, Wang H, Wang T, Zhang X, Chen H, Yu H, Zhang X, Zhang M, Wu S, et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J Clin Invest. 2020; 130:2620–29. https://doi.org/10.1172/JCI137244 [PubMed]
- 6. Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020; 395:507–13. https://doi.org/10.1016/S0140-6736(20)30211-7 [PubMed]
- 7. Yang X, Yu Y, Xu J, Shu H, Xia J, Liu H, Wu Y, Zhang L, Yu Z, Fang M, Yu T, Wang Y, Pan S, et al. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir Med. 2020; 8:475–81. https://doi.org/10.1016/s2213-2600(20)30079-5 [PubMed]
- 8. Guan WJ, Ni ZY, Hu Y, Liang WH, Ou CQ, He JX, Liu L, Shan H, Lei CL, Hui DS, Du B, Li LJ, Zeng G, et al, and China Medical Treatment Expert Group for Covid-19. Clinical Characteristics of Coronavirus Disease 2019 in China. N Engl J Med. 2020; 382:1708–20. https://doi.org/10.1056/NEJMoa2002032 [PubMed]
- 9. Cai J, Li H, Zhang C, Chen Z, Liu H, Lei F, Qin JJ, Liu YM, Zhou F, Song X, Zhou J, Zhao YC, Wu B, et al. The Neutrophil-to-Lymphocyte Ratio Determines Clinical Efficacy of Corticosteroid Therapy in Patients with COVID-19. Cell Metab. 2021; 33:258–69.e3. https://doi.org/10.1016/j.cmet.2021.01.002 [PubMed]
- 10. Wong CK, Lam CWK, Wu AKL, Ip WK, Lee NLS, Chan IHS, Lit LCW, Hui DSC, Chan MHM, Chung SSC, Sung JJY. Plasma inflammatory cytokines and chemokines in severe acute respiratory syndrome. Clin Exp Immunol. 2004; 136:95–103. https://doi.org/10.1111/j.1365-2249.2004.02415.x [PubMed]
- 11. Mahallawi WH, Khabour OF, Zhang Q, Makhdoum HM, Suliman BA. MERS-CoV infection in humans is associated with a pro-inflammatory Th1 and Th17 cytokine profile. Cytokine. 2018; 104:8–13. https://doi.org/10.1016/j.cyto.2018.01.025 [PubMed]
- 12. Qin C, Zhou L, Hu Z, Zhang S, Yang S, Tao Y, Xie C, Ma K, Shang K, Wang W, Tian DS. Dysregulation of immune response in patients with COVID-19 in Wuhan, China. Clin Infect Dis. 2020; 71:762–68. https://doi.org/10.1093/cid/ciaa248 [PubMed]
- 13. Catanzaro M, Fagiani F, Racchi M, Corsini E, Govoni S, Lanni C. Immune response in COVID-19: addressing a pharmacological challenge by targeting pathways triggered by SARS-CoV-2. Signal Transduct Target Ther. 2020; 5:84. https://doi.org/10.1038/s41392-020-0191-1 [PubMed]
- 14. Giamarellos-Bourboulis EJ, Netea MG, Rovina N, Akinosoglou K, Antoniadou A, Antonakos N, Damoraki G, Gkavogianni T, Adami ME, Katsaounou P, Ntaganou M, Kyriakopoulou M, Dimopoulos G, et al. Complex Immune Dysregulation in COVID-19 Patients with Severe Respiratory Failure. Cell Host Microbe. 2020; 27:992–1000.e3. https://doi.org/10.1016/j.chom.2020.04.009 [PubMed]
- 15. Wang F, Nie J, Wang H, Zhao Q, Xiong Y, Deng L, Song S, Ma Z, Mo P, Zhang Y. Characteristics of Peripheral Lymphocyte Subset Alteration in COVID-19 Pneumonia. J Infect Dis. 2020; 221:1762–69. https://doi.org/10.1093/infdis/jiaa150 [PubMed]
- 16. Liu J, Li S, Liu J, Liang B, Wang X, Wang H, Li W, Tong Q, Yi J, Zhao L, Xiong L, Guo C, Tian J, et al. Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS-CoV-2 infected patients. EBioMedicine. 2020; 55:102763. https://doi.org/10.1016/j.ebiom.2020.102763 [PubMed]
- 17. Luo W, Zhang JW, Zhang W, Lin YL, Wang Q. Circulating levels of IL-2, IL-4, TNF-α, IFN-γ, and C-reactive protein are not associated with severity of COVID-19 symptoms. J Med Virol. 2021; 93:89–91. https://doi.org/10.1002/jmv.26156 [PubMed]
- 18. Sun Y, Dong Y, Wang L, Xie H, Li B, Chang C, Wang FS. Characteristics and prognostic factors of disease severity in patients with COVID-19: The Beijing experience. J Autoimmun. 2020; 112:102473. https://doi.org/10.1016/j.jaut.2020.102473 [PubMed]
- 19. Chen D, Li X, Song Q, Hu C, Su F, Dai J, Ye Y, Huang J, Zhang X. Assessment of Hypokalemia and Clinical Characteristics in Patients With Coronavirus Disease 2019 in Wenzhou, China. JAMA Netw Open. 2020; 3:e2011122. https://doi.org/10.1001/jamanetworkopen.2020.11122 [PubMed]
- 20. Suleyman G, Fadel RA, Malette KM, Hammond C, Abdulla H, Entz A, Demertzis Z, Hanna Z, Failla A, Dagher C, Chaudhry Z, Vahia A, Abreu Lanfranco O, et al. Clinical Characteristics and Morbidity Associated With Coronavirus Disease 2019 in a Series of Patients in Metropolitan Detroit. JAMA Netw Open. 2020; 3:e2012270. https://doi.org/10.1001/jamanetworkopen.2020.12270 [PubMed]
- 21. Azkur AK, Akdis M, Azkur D, Sokolowska M, van de Veen W, Brüggen MC, O’Mahony L, Gao Y, Nadeau K, Akdis CA. Immune response to SARS-CoV-2 and mechanisms of immunopathological changes in COVID-19. Allergy. 2020; 75:1564–81. https://doi.org/10.1111/all.14364 [PubMed]
- 22. Zhang G, Hu C, Luo L, Fang F, Chen Y, Li J, Peng Z, Pan H. Clinical features and short-term outcomes of 221 patients with COVID-19 in Wuhan, China. J Clin Virol. 2020; 127:104364. https://doi.org/10.1016/j.jcv.2020.104364 [PubMed]
- 23. Young BE, Ong SW, Kalimuddin S, Low JG, Tan SY, Loh J, Ng OT, Marimuthu K, Ang LW, Mak TM, Lau SK, Anderson DE, Chan KS, et al, and Singapore 2019 Novel Coronavirus Outbreak Research Team. Epidemiologic Features and Clinical Course of Patients Infected With SARS-CoV-2 in Singapore. JAMA. 2020; 323:1488–94. https://doi.org/10.1001/jama.2020.3204 [PubMed]
- 24. Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus–Infected Pneumonia in Wuhan, China. JAMA. 2020; 323:1061–69. https://doi.org/10.1001/jama.2020.1585 [PubMed]
- 25. Su W, Qiu Z, Zhou L, Hou J, Wang Y, Huang F, Zhang Y, Jia Y, Zhou J, Liu D, Xia Z, Xia ZY, Lei S. Sex differences in clinical characteristics and risk factors for mortality among severe patients with COVID-19: a retrospective study. Aging (Albany NY). 2020; 12:18833–43. https://doi.org/10.18632/aging.103793 [PubMed]
- 26. Huang W, Li C, Wang Z, Wang H, Zhou N, Jiang J, Ni L, Zhang XA, Wang DW. Decreased serum albumin level indicates poor prognosis of COVID-19 patients: hepatic injury analysis from 2,623 hospitalized cases. Sci China Life Sci. 2020; 63:1678–87. https://doi.org/10.1007/s11427-020-1733-4 [PubMed]
- 27. Bode B, Garrett V, Messler J, McFarland R, Crowe J, Booth R, Klonoff DC. Glycemic Characteristics and Clinical Outcomes of COVID-19 Patients Hospitalized in the United States. J Diabetes Sci Technol. 2020; 14:813–21. https://doi.org/10.1177/1932296820924469 [PubMed]
- 28. Berhane M, Melku M, Amsalu A, Enawgaw B, Getaneh Z, Asrie F. The Role of Neutrophil to Lymphocyte Count Ratio in the Differential Diagnosis of Pulmonary Tuberculosis and Bacterial Community-Acquired Pneumonia: a Cross-Sectional Study at Ayder and Mekelle Hospitals, Ethiopia. Clin Lab. 2019; 65. https://doi.org/10.7754/Clin.Lab.2018.180833 [PubMed]
- 29. Liu F, Li L, Xu M, Wu J, Luo D, Zhu Y, Li B, Song X, Zhou X. Prognostic value of interleukin-6, C-reactive protein, and procalcitonin in patients with COVID-19. J Clin Virol. 2020; 127:104370. https://doi.org/10.1016/j.jcv.2020.104370 [PubMed]
- 30. Chien JY, Hsueh PR, Cheng WC, Yu CJ, Yang PC. Temporal changes in cytokine/chemokine profiles and pulmonary involvement in severe acute respiratory syndrome. Respirology. 2006; 11:715–22. https://doi.org/10.1111/j.1440-1843.2006.00942.x [PubMed]
- 31. Jiang Y, Xu J, Zhou C, Wu Z, Zhong S, Liu J, Luo W, Chen T, Qin Q, Deng P. Characterization of cytokine/chemokine profiles of severe acute respiratory syndrome. Am J Respir Crit Care Med. 2005; 171:850–57. https://doi.org/10.1164/rccm.200407-857OC [PubMed]
- 32. Chu H, Zhou J, Wong BH, Li C, Cheng ZS, Lin X, Poon VK, Sun T, Lau CC, Chan JF, To KK, Chan KH, Lu L, et al. Productive replication of Middle East respiratory syndrome coronavirus in monocyte-derived dendritic cells modulates innate immune response. Virology. 2014; 454–455:197–205. https://doi.org/10.1016/j.virol.2014.02.018 [PubMed]
- 33. Channappanavar R, Fehr AR, Vijay R, Mack M, Zhao J, Meyerholz DK, Perlman S. Dysregulated Type I Interferon and Inflammatory Monocyte-Macrophage Responses Cause Lethal Pneumonia in SARS-CoV-Infected Mice. Cell Host Microbe. 2016; 19:181–93. https://doi.org/10.1016/j.chom.2016.01.007 [PubMed]
- 34. Fu Y, Cheng Y, Wu Y. Understanding SARS-CoV-2-Mediated Inflammatory Responses: From Mechanisms to Potential Therapeutic Tools. Virol Sin. 2020; 35:266–71. https://doi.org/10.1007/s12250-020-00207-4 [PubMed]
- 35. Gadient RA, Patterson PH. Leukemia inhibitory factor, Interleukin 6, and other cytokines using the GP130 transducing receptor: roles in inflammation and injury. Stem Cells. 1999; 17:127–37. https://doi.org/10.1002/stem.170127 [PubMed]
- 36. Channappanavar R, Perlman S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin Immunopathol. 2017; 39:529–39. https://doi.org/10.1007/s00281-017-0629-x [PubMed]
- 37. Liu Y, Zhang C, Huang F, Yang Y, Wang F, Yuan J, Zhang Z, Qin Y, Li X, Zhao D, Li S, Tan S, Wang Z, et al. Elevated plasma level of selective cytokines in COVID-19 patients reflect viral load and lung injury. Natl Sci Rev. 2020; 7:1003–11. https://doi.org/10.1093/nsr/nwaa037
- 38. Grifoni A, Weiskopf D, Ramirez SI, Mateus J, Dan JM, Moderbacher CR, Rawlings SA, Sutherland A, Premkumar L, Jadi RS, Marrama D, de Silva AM, Frazier A, et al. Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell. 2020; 181:1489–501.e15. https://doi.org/10.1016/j.cell.2020.05.015 [PubMed]
- 39. Tan L, Wang Q, Zhang D, Ding J, Huang Q, Tang YQ, Wang Q, Miao H. Lymphopenia predicts disease severity of COVID-19: a descriptive and predictive study. Signal Transduct Target Ther. 2020; 5:33. https://doi.org/10.1038/s41392-020-0148-4 [PubMed]
- 40. National Health Commission of the People’s Republic of China. The Seventh Revised Trial Version of the Novel Coronavirus Pneumonia Diagnosis and Treatment Guidance. http://www.nhc.gov.cn/yzygj/s7652m/202003/a31191442e29474b98bfed5579d5af95.shtml.