Introduction
Dopamine is primarily recognized as a crucial factor for neurodegenerative diseases such as Alzheimer’s disease and Parkinson’s disease, which are normally characterized by gradual cognitive decline and dysfunction [1–3]. Furthermore, dopamine, mainly located in the midbrain, performs through five distinct subtypes of dopamine receptors (namely D1, D2, D3, D4, and D5 receptors) [4]. The dopamine receptors, G-protein coupled receptors, have been a focus of attention in the arena of drug design and discovery, where it is a target for drugs which treat various disorders such as Parkinson’s disease and psychiatric disorders [5–9].
The dopamine receptor genes DRD1, DRD2, DRD3, DRD4, and DRD5 encode the D1, D2, D3, D4, and D5 subtype of the dopamine receptors, respectively. Genetic variants, in particular, single nucleotide polymorphisms (SNPs), have been described in the dopamine receptor gene research [10–12]. Literature suggests a connection between the dopamine receptor-related loci and neurodegenerative diseases, as well as between the dopamine receptor-related loci and cognitive functions in human beings [13]. For example, Tsang et al. [14] reported that the DRD1 gene was associated with inferior cognitive performance in the postmortem cohort of Caucasian samples with Alzheimer's disease. Furthermore, it has been demonstrated that the DRD2 gene was linked to better cognitive performance in verbal learning following traumatic brain injury [15] and may influence cognitive performance (such as number of categories achieved in the Wisconsin Card Sorting Test) in healthy individuals [16]. Mota et al. [17] also suggested that DRD2 co-regulates with three nearby genes (NCAM1, TTC12, and ANKK1) to form the NCAM1-TTC12-ANKK1-DRD2 locus which correlates with dopaminergic neurotransmission and neurogenesis. Moreover, Cordeddu et al. [18] found that DRD3 joins with its adjacent genes (LOC107986115, ZNF80, TIGIT, MIR568, and ZBTB20) to constitute the DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20 locus which contributes to Primrose syndrome, a genetic disorder with intellectual disability and learning difficulties. In addition, it has been indicated that the DRD4 gene exhibits a significant association in Alzheimer’s disease in the Taiwanese population [19] and is linked to a specific cognitive domain called perceptual speed performance in cognitively healthy individuals of European ancestry [20]. Finally, Hollingworth et al. [21] identified the SLC2A9 gene, which is neighboring to DRD5, as a susceptibility gene for psychosis and Alzheimer’s disease in populations of European ancestry.
In light of the aforementioned observations, it was presumed that the dopamine receptor-related loci, including the DRD1, NCAM1-TTC12-ANKK1-DRD2, DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20, DRD4, and DRD5-SLC2A9 loci, may play a key role in the pathogenesis of age-dependent cognitive decline and the development of cognitive aging. Here, we hypothesized that genetic variants such as SNPs in the dopamine receptor-related loci might be associated with cognitive aging in the population with Taiwanese ancestry. To the best of our knowledge, the effects of genetic variants in the dopamine receptor-related loci on cognitive aging are scant in regards to human studies. In addition, several studies [22–27] in the Taiwan Biobank have conducted association analyses on cognitive aging using Mini-Mental State Examination (MMSE) scores, a standard method provided in the Taiwan Biobank. Based on the aforementioned considerations, we conducted the first genetic association study between MMSE scores and SNPs in the dopamine receptor-related loci in the Taiwan Biobank. We detected associations between MMSE scores and genetic variants in three genes (including NCAM1 and TTC12 on chromosome 11 and ZBTB20 on chromosome 3) which have not been previously discovered. It has also been indicated that environmental factors such as physical activity are linked to cognitive aging as well [22–25]. Substantial evidence reveals that physical activity contributes to a lower risk of developing cognitive impairment and Alzheimer's disease [28]. Thus, we assessed the probable gene-physical activity interactions on cognitive aging and found potential gene-physical activity interactions with the NCAM1, TTC12, and ZBTB20 genes in influencing cognitive aging.
Results
The clinical and demographic characteristics of the study cohort
Table 1 illustrates the clinical and demographic characteristics of our study cohort, which consisted of 25,195 individuals from the Taiwan Biobank. The median MMSE score was 28 and the interquartile range was 26–29. Supplementary Table 1 presents the demographic characteristics and the relevant MMSE scores in our study cohort. In this study, we found that correlations between MMSE scores with age (P < 0.001), gender (P < 0.001), education (P < 0.001), physical activity (P < 0.001), smoking (P = 0.004), and chronic conditions (P < 0.001) were significant (Supplementary Table 1).
Table 1. Demographic and clinical characteristics of study subjects.
| Characteristic | Overall | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| No. of subjects, n | 25,195 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mean age ± SD, years | 64.3±3.2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Female, % | 61.2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Education level 1, % “1” for no formal education, “2” for homeschooling, “3” for elementary school, “4” for middle school, “5” for high school, "6" for college, and "7" for graduate school | 1: 8.9 2: 14.3 3: 12.8 4: 9.0 5: 23.4 6: 27.2 7: 4.4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical activity, % | 63.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Smoking, % | 5.6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alcohol drinking, % | 5.3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chronic conditions, % | 84.7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MMSE score, median (IQR) | 28 (26–29) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IQR, interquartile range; MMSE, Mini-Mental State Examination; SD, standard deviation. Data are presented as mean ± standard deviation. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1Education level is defined as the following seven levels: no formal education, homeschooling, elementary school, middle school, high school, college, and graduate school. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Physical activity and gene interaction analysis
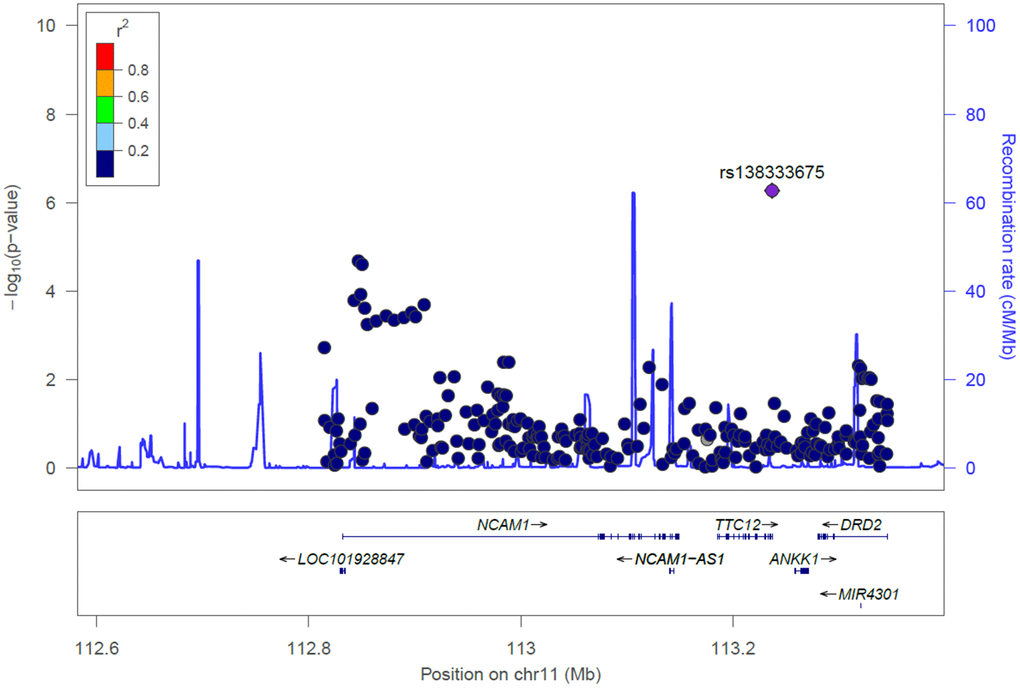
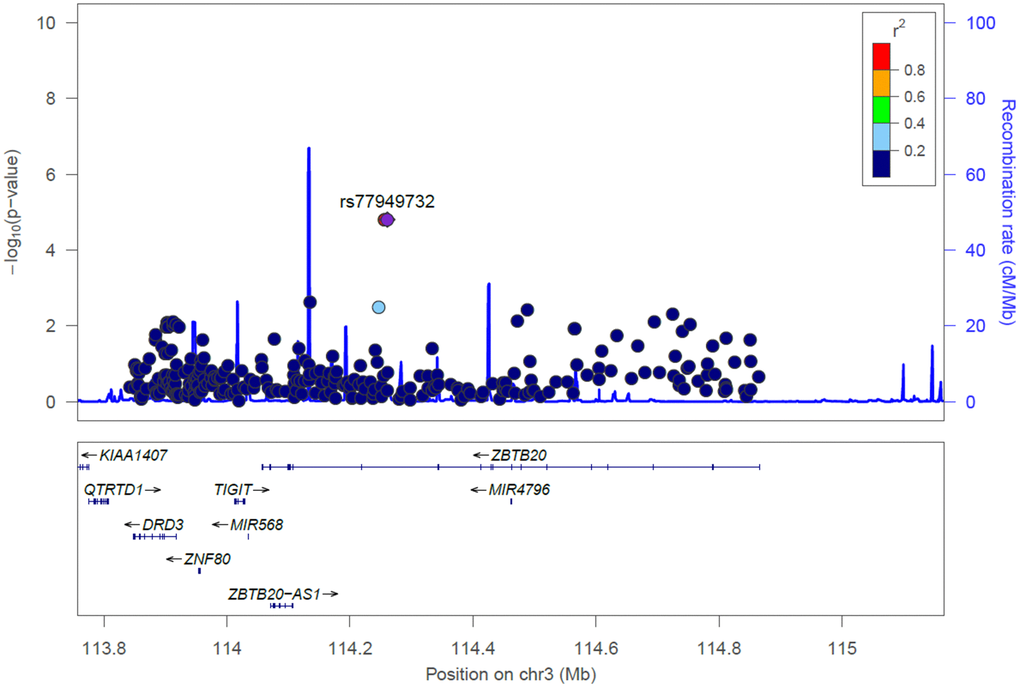
Next, we utilized categorized MMSE scores as an outcome (normal: MMSE score ≥ 24; cognitive impairment: MMSE score < 24) for physical activity and gene interaction analysis. The GMDR method was employed to estimate the effects of consolidation between physical activity and the three key SNPs (namely rs11214442 in NCAM1, rs138333675 in TTC12, and rs77949732 in ZBTB20) in cognitive aging by incorporating age, gender, and education as covariates. Here, we only selected one top SNP from each of NCAM1, TTC12, and ZBTB20. As illustrated in Table 3, there were significant two-way models concerning physical activity among rs11214442 in NCAM1, rs138333675 in TTC12, and rs77949732 in ZBTB20 (P < 0.001, < 0.001, and < 0.001, respectively), indicating potential physical activity and gene interactions between these genes and physical activity in regulating cognitive aging. The effect of these physical activity and gene interaction models remained significant after Bonferroni correction (P < 0.05/3 = 0.017).
Table 3. Physical activity and gene interaction models identified by the GMDR method.
| 2-way interaction model | Testing accuracy (%) | P value | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical activity, rs11214442 in NCAM1 | 53.86 | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical activity, rs138333675 in TTC12 | 53.86 | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical activity, rs77949732 in ZBTB20 | 53.94 | < 0.001 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GMDR, generalized multifactor dimensionality reduction. P value was based on 1,000 permutations. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Analysis was obtained after adjustment for covariates including age, gender, education, smoking, alcohol drinking, and chronic conditions. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| P values of < 0.017 (Bonferroni correction: 0.05/3) are shown in bold. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Discussion
To our knowledge, this is the first study to date to determine whether the main impacts of SNPs in the dopamine receptor-related loci are significantly associated with the risk of cognitive aging on an individual-by-individual basis and by virtue of gene-physical activity interactions in elder subjects. Here, we observed for the first time that the dopamine receptor-related loci may play an essential role in influencing cognitive aging in elder Taiwanese individuals. Intriguingly, the significance persisted for the association of MMSE scores with six key SNPs after correcting for multiple testing, including rs11214442 in NCAM1, rs10891485 in NCAM1, and rs138333675 in TTC12, rs145272406 in ZBTB20, rs114295131 in ZBTB20, and rs77949732 in ZBTB20. Moreover, the findings indicated that gene-physical activity interactions in the NCAM1, TTC12, and ZBTB20 genes may modulate the etiology of cognitive aging.
Remarkably, the present study is the first to raise the possibility that three significant SNPs in the NCAM1-TTC12-ANKK1-DRD2 locus, such as rs11214442 in NCAM1, rs10891485 in NCAM1, and rs138333675 in TTC12, may be associated with cognitive aging. Furthermore, the significant association between these key SNPs and MMSE scores persisted even after applying Bonferroni correction. We also found that rs11214442 and rs10891485 in NCAM1 may play a functional role as eQTLs for the NCAM1 gene in artery/aorta tissues [29]. To our knowledge, no other studies have been conducted to pinpoint these three SNPs in NCAM1 and TTC12 with cognitive aging or age-related cognitive decline. Both the NCAM1 and TTC12 genes, located on chromosome 11q23.2, are good candidates for cognitive aging because they represent the NCAM1-TTC12-ANKK1-DRD2 locus which is linked to dopaminergic neurotransmission and neurogenesis [17]. It has also been indicated that NCAM1 may play a key role in impairment of spatial memory in epileptic rats [30] as well as in associative memory in nematodes and humans [31]. Moreover, it has been suggested that the NCAM1-TTC12-ANKK1-DRD2 locus should be considered as one single unit for investigating genetics in behavioral and psychiatric studies [17]. In addition, it has been reported that the NCAM1-TTC12-ANKK1-DRD2 locus is considered as a risk biomarker for alcohol dependence [32], nicotine dependence [33], heroin dependence [34], and comorbid alcohol and drug dependence [35].
Interestingly, the present study is the first to raise the possibility that three significant SNPs in the DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20 locus, such as rs145272406 in ZBTB20, rs114295131 in ZBTB20, and rs77949732 in ZBTB20, may be associated with cognitive aging. To our knowledge, no other studies have been conducted to pinpoint these three SNPs in ZBTB20 with cognitive aging or age-related cognitive decline. The ZBTB20 gene, located on chromosome 3q13.31, represents the DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20 locus. ZBTB20 is a good candidate for cognitive aging. For example, it has been reported that mutations in ZBTB20 are associated with Primrose syndrome, a genetic disorder characterized by intellectual disability, autism, and other behavioral concerns [18]. On another note, ZBTB20 has been previously implicated in influencing neurodevelopmental disorders in subjects of European ancestry [36]. Animal studies also revealed that the Zbtb20 gene is mainly expressed in brain and its encoded protein is implicated in hippocampal development and cerebellar granule cells [37, 38].
Intriguingly, we pinpointed the interplays between the dopamine receptor-related loci and physical activity in affecting cognitive aging, where the dopamine receptor-related loci encompass NCAM1, TTC12, and ZBTB20. This relationship could functionally manifest itself on the basis of epigenetic alterations [23]. Our findings are in line with other association studies in the Taiwanese population from the Taiwan Biobank, indicating that physical activity may modulate cognitive aging by means of likely complex gene-physical activity interplays with the interleukin-12 related genes (such as IL12A, IL12B, and IL12RB2) [23], the DNA repair gene EXO1 [24], circadian clock genes (such as RORA and RORB) [22], and Alzheimer's disease-associated genes (such as SLC24A4) [25]. Papenberg et al. [39] also suggested the detrimental effects of inflammation on cognitive aging in old adults who lack of physical activity. Furthermore, the dopamine receptor-related loci, such as DRD2 and DRD3, have been found to play a central role in inflammation, neuroinflammation, and neurodegeneration [40, 41]. Thus, it is plausible to hypothesize that the interplays found in this study may be linked to the negative effects of inflammation on cognitive aging in inactive old adults.
In our analysis, there was also evidence of associations (P < 0.05) between MMSE scores and other dopamine receptor-related loci, such as DRD1, ARL2BPP6-DRD1, DRD1-SFXN1, DRD2, NCAM1-LOC105369498, DRD3, DRD3-LOC107986115, ZNF80-TIGIT, DRD4-DEAF1, and SLC2A9. The DRD1, ARL2BPP6-DRD1, and DRD1-SFXN1 loci are located on chromosome 5q35.2. The DRD2 and NCAM1-LOC105369498 loci, located on chromosome 11q23.2, represent the NCAM1-TTC12-ANKK1-DRD2 locus. The DRD3, DRD3-LOC107986115, and ZNF80-TIGIT loci, located on chromosome 3q13.31, represents the DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20 locus. The DRD4-DEAF1 locus is located on chromosome 11p15.5. The SLC2A9 gene, located on chromosome 4p16.1, represents the DRD5-SLC2A9 locus. In accordance with our results, it has been reported that DRD1 [14], DRD2 [15, 16], DRD3 [18], DRD4 [19, 20], and SLC2A9 [21] are associated with cognitive performance or cognitive impairment.
This study had some limitations. First, the age effect was limited due to a homogeneous cohort in terms of age (i.e., the mean age of 64.3 years and the standard deviation of 3.2 years). Second, studies on cognitive aging require repetitive cognitive assessments of each individual over years and our study is limited to a single cognitive assessment of each individual. Furthermore, in our statistical analyses, adjustments were not made for other risk factors, namely brain injury and exposure to pesticides/toxins, due to a lack of such data. Future studies making use of multi-omics data are also warranted to provide the molecular mechanisms underlying cognitive aging.
In conclusion, the present study accomplished a thorough investigation of the associations of cognitive aging with the dopamine receptor-related loci in old adults in the Taiwanese population. Moreover, the present study revealed the impacts of gene-physical activity interactions in the dopamine receptor-related loci with regard to cognitive aging. In particular, if the current findings are reproduced in statistically well-powered distinct samples, the present study pinpoints the effects of the dopamine receptor-related loci on the risk of cognitive aging on an individual-by-individual basis as well as via sophisticated gene-physical activity interplays. The present study implicates that dopamine receptor-mediated signaling should be the focus of future studies on pathogenesis of age-dependent cognitive decline and probable drug targets for drug design and discovery. Distinct studies with greater replication datasets will potentially create further insights into the role of the dopamine receptor-related loci demonstrated in the present study.
Materials and Methods
Study population
Our study cohort composed of 25,195 subjects for subsequent analyses. This study involved Taiwanese individuals from the Taiwan Biobank, which collected specimens and relevant information from participants in recruitment centers across Taiwan [22, 25–27, 42, 43]. There were the following two inclusion criteria: (1) participants who were 60 years old or over; and (2) participants who were self-reported as being of Taiwanese ancestry [43]. The exclusion criterion was participants with a history of cancer [43]. Ethical approval for the study was granted by the Institutional Review Board of the Taiwan Biobank before performing the study (approval number: 201506095RINC). The approved informed consent form was signed by each subject. All experiments were achieved by means of proper regulations and guidelines.
A subject’s education level includes the following seven levels: no formal education, homeschooling, elementary school, middle school, high school, college, and graduate school [22–27]. A subject who had exercised for over 30 minutes each time and over three times each week was defined as a measure of physical activity [22–25]. A subject with current smoking for more than 6 months was defined as a current smoker [22–25]. A subject with a volume of 150mL of alcohol intake per week for more than 6 months was defined as a current alcohol drinker [22–25]. Status of chronic conditions was defined as whether a subject or a subject's family member (i.e., family history) has had the following chronic conditions: Parkinson's disease, heart disease, stroke, and/or diabetes.
Cognitive assessment
We utilized the 30-point MMSE, a cognitive impairment screening tool in the Taiwan Biobank, as previously described [22–27]. In brief, we assessed global cognitive performance by employing the 30-point MMSE, which encompasses questions in accordance with the five areas of recall, registration, language, attention and calculation, and orientation [27]. We assessed the MMSE score both as a continuous phenotype and as a binary phenotype in accordance with the following previously-established MMSE thresholds [44]: MMSE score ≥ 24 (normal) and MMSE score < 24 (cognitive impairment). The cognitive assessment was carried out in the local languages (such as Taiwanese and/or Taiwanese Mandarin). The cognitive cut-off score of 24 was originated from previous studies [44] and was based on a Taiwanese version of MMSE.
Laboratory assessments: genotyping
DNA was extracted from blood samples by employing QIAamp DNA blood kits following the manufacturer’s instructions (Qiagen, Valencia, CA, USA). The quality of the isolated genomic DNA was carried out by utilizing agarose gel electrophoresis, and the quantity was completed by spectrophotometry [45]. SNP genotyping was evaluated by employing the custom Taiwan Biobank chips, which were accomplished by using the Axiom Genome-Wide Array Plate System (Affymetrix, Santa Clara, CA, USA). The custom Taiwan Biobank chips were created to collect genetic profiles in Taiwanese subjects by utilizing SNPs on the Axiom Genome-Wide CHB 1 Array (Affymetrix, Santa Clara, CA, USA) and the Human Exome BeadChip (Illumina, Inc., San Diego, CA, USA) [43].
We searched for variants in the DRD1, NCAM1-TTC12-ANKK1-DRD2, DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20, DRD4, and DRD5-SLC2A9 loci by referring to the complete list of SNPs available in the custom Taiwan Biobank chips. In addition, we performed quality control procedures for subsequent analysis [46, 47]. The quality control procedure excluded troublesome SNPs that were not in Hardy-Weinberg equilibrium (HWE) (with a P-value less than 0.05) or had a genotyping call rate less than 95% or minor allele frequency (MAF) < 1%. We evaluated MAFs, genotyping call rates, P values for HWE using PLINK [48]. After the quality control procedure, the SNP panel consisted of 792 SNPs in the DRD1, NCAM1-TTC12-ANKK1-DRD2, DRD3-LOC107986115-ZNF80-TIGIT-MIR568-ZBTB20, DRD4, and DRD5-SLC2A9 loci.
Statistical analysis
The Student’s t test was performed to measure the difference in the means of two continuous variables [23]. We conducted the chi-square test for categorical data. The criterion for significance was set at P < 0.05 for all tests. Data are presented as the mean ± standard deviation.
In this study, linear regression analysis was carried out to evaluate the relationship between MMSE scores and our variables of interest such as age, gender, education, physical activity, smoking, alcohol drinking, and chronic conditions. In addition, we assessed the association of the investigated SNP with MMSE scores by a general linear model using age, gender, education, physical activity, smoking, alcohol drinking, and chronic conditions as covariates [49]. The genotype frequencies were performed for Hardy-Weinberg equilibrium to detect genotyping errors [50] by employing a χ2 goodness-of-fit test with one degree of freedom (that is, the number of genotypes minus the number of alleles). Adjustments for multiple testing were conducted by employing the Bonferroni correction. The criterion for significance was defined as P < 0.05 for all tests. Data were shown by the mean ± standard deviation.
In order to examine gene-physical activity interplays, we utilized the generalized multifactor dimensionality reduction (GMDR) method [51]. We scrutinized two-way interactions by employing 10-fold cross-validation. The GMDR method provided various output parameters, such as empirical P values and the testing accuracy, to evaluate each interaction. Furthermore, covariates such as age, gender, education, smoking, alcohol drinking, and chronic conditions were utilized for gene-physical activity interaction analysis in our interaction models. We completed the empirical P value of the testing accuracy for each interaction by employing permutation testing (based on 1,000 shuffles). Finally, we utilized the Bonferroni correction to correct for multiple testing.
In addition, we employed HaploReg (http://compbio.mit.edu/HaploReg) [52] to test if there is a functional role as eQTLs for the SNPs in the specific genes.
Author Contributions
Eugene Lin: study conception and design, analysis and interpretation of data, draft manuscript. Po-Hsiu Kuo: acquisition of data. Wan-Yu Lin: acquisition of data. Yu-Li Liu: acquisition of data. Albert C. Yang: acquisition of data. Shih-Jen Tsai: study conception and design.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Funding
This work was supported by grant MOST 109-2634-F-075-001 from Taiwan Ministry of Science and Technology, and grant V108D44-001-MY3-1 from the Taipei Veterans General Hospital.
References
- 1. Martorana A, Koch G. “Is dopamine involved in Alzheimer’s disease?”. Front Aging Neurosci. 2014; 6:252. https://doi.org/10.3389/fnagi.2014.00252 [PubMed]
- 2. Pan X, Kaminga AC, Wen SW, Wu X, Acheampong K, Liu A. Dopamine and Dopamine Receptors in Alzheimer’s Disease: A Systematic Review and Network Meta-Analysis. Front Aging Neurosci. 2019; 11:175. https://doi.org/10.3389/fnagi.2019.00175 [PubMed]
- 3. Rubinstein TC, Giladi N, Hausdorff JM. The power of cueing to circumvent dopamine deficits: a review of physical therapy treatment of gait disturbances in Parkinson’s disease. Mov Disord. 2002; 17:1148–60. https://doi.org/10.1002/mds.10259 [PubMed]
- 4. Kumar U, Patel SC. Immunohistochemical localization of dopamine receptor subtypes (D1R-D5R) in Alzheimer’s disease brain. Brain Res. 2007; 1131:187–96. https://doi.org/10.1016/j.brainres.2006.10.049 [PubMed]
- 5. Jatana N, Thukral L, Latha N. Structural signatures of DRD4 mutants revealed using molecular dynamics simulations: Implications for drug targeting. J Mol Model. 2016; 22:14. https://doi.org/10.1007/s00894-015-2868-x [PubMed]
- 6. Lindsley CW, Hopkins CR. Return of D4 Dopamine Receptor Antagonists in Drug Discovery. J Med Chem. 2017; 60:7233–43. https://doi.org/10.1021/acs.jmedchem.7b00151 [PubMed]
- 7. Oak JN, Oldenhof J, Van Tol HH. The dopamine D(4) receptor: one decade of research. Eur J Pharmacol. 2000; 405:303–27. https://doi.org/10.1016/s0014-2999(00)00562-8 [PubMed]
- 8. Wong AH, Van Tol HH. The dopamine D4 receptors and mechanisms of antipsychotic atypicality. Prog Neuropsychopharmacol Biol Psychiatry. 2003; 27:1091–99. https://doi.org/10.1016/j.pnpbp.2003.09.005 [PubMed]
- 9. Beaulieu JM, Gainetdinov RR. The physiology, signaling, and pharmacology of dopamine receptors. Pharmacol Rev. 2011; 63:182–217. https://doi.org/10.1124/pr.110.002642 [PubMed]
- 10. Bookman EB, Taylor RE, Adams-Campbell L, Kittles RA. DRD4 promoter SNPs and gender effects on Extraversion in African Americans. Mol Psychiatry. 2002; 7:786–89. https://doi.org/10.1038/sj.mp.4001075 [PubMed]
- 11. Simpson J, Vetuz G, Wilson M, Brookes KJ, Kent L. The DRD4 receptor Exon 3 VNTR and 5' SNP variants and mRNA expression in human post-mortem brain tissue. Am J Med Genet B Neuropsychiatr Genet. 2010; 153:1228–33. https://doi.org/10.1002/ajmg.b.31084 [PubMed]
- 12. Wang E, Ding YC, Flodman P, Kidd JR, Kidd KK, Grady DL, Ryder OA, Spence MA, Swanson JM, Moyzis RK. The genetic architecture of selection at the human dopamine receptor D4 (DRD4) gene locus. Am J Hum Genet. 2004; 74:931–44. https://doi.org/10.1086/420854 [PubMed]
- 13. Savitz J, Solms M, Ramesar R. The molecular genetics of cognition: dopamine, COMT and BDNF. Genes Brain Behav. 2006; 5:311–28. https://doi.org/10.1111/j.1601-183X.2005.00163.x [PubMed]
- 14. Tsang J, Fullard JF, Giakoumaki SG, Katsel P, Katsel P, Karagiorga VE, Greenwood TA, Braff DL, Siever LJ, Bitsios P, Haroutunian V, Roussos P. The relationship between dopamine receptor D1 and cognitive performance. NPJ Schizophr. 2015; 1:14002. https://doi.org/10.1038/npjschz.2014.2 [PubMed]
- 15. Yue JK, Winkler EA, Rick JW, Burke JF, McAllister TW, Oh SS, Burchard EG, Hu D, Rosand J, Temkin NR, Korley FK, Sorani MD, Ferguson AR, et al, and TRACK-TBI Investigators. DRD2 C957T polymorphism is associated with improved 6-month verbal learning following traumatic brain injury. Neurogenetics. 2017; 18:29–38. https://doi.org/10.1007/s10048-016-0500-6 [PubMed]
- 16. Rodriguez-Jimenez R, Hoenicka J, Jimenez-Arriero MA, Ponce G, Bagney A, Aragues M, Palomo T. Performance in the Wisconsin Card Sorting Test and the C957T polymorphism of the DRD2 gene in healthy volunteers. Neuropsychobiology. 2006; 54:166–70. https://doi.org/10.1159/000098652 [PubMed]
- 17. Mota NR, Araujo-Jnr EV, Paixão-Côrtes VR, Bortolini MC, Bau CH. Linking dopamine neurotransmission and neurogenesis: The evolutionary history of the NTAD (NCAM1-TTC12-ANKK1-DRD2) gene cluster. Genet Mol Biol. 2012; 35:912–18. https://doi.org/10.1590/s1415-47572012000600004 [PubMed]
- 18. Cordeddu V, Redeker B, Stellacci E, Jongejan A, Fragale A, Bradley TE, Anselmi M, Ciolfi A, Cecchetti S, Muto V, Bernardini L, Azage M, Carvalho DR, et al. Mutations in ZBTB20 cause Primrose syndrome. Nat Genet. 2014; 46:815–17. https://doi.org/10.1038/ng.3035 [PubMed]
- 19. Lin WY, Wu BT, Lee CC, Sheu JJ, Liu SH, Wang WF, Tsai CH, Liu HP, Tsai FJ. Association analysis of dopaminergic gene variants (Comt, Drd4 And Dat1) with Alzheimer s disease. J Biol Regul Homeost Agents. 2012; 26:401–10. [PubMed]
- 20. Barral S, Habeck C, Gazes E, De Jager PL, Bennett DA, Stern Y. A Dopamine Receptor genetic variant enhances perceptual speed in cognitive healthy subjects. Alzheimers Dement (N Y). 2017; 3:254–61. https://doi.org/10.1016/j.trci.2017.03.004 [PubMed]
- 21. Hollingworth P, Sweet R, Sims R, Harold D, Russo G, Abraham R, Stretton A, Jones N, Gerrish A, Chapman J, Ivanov D, Moskvina V, Lovestone S, et al, and GERAD Consortium, and National Institute on Aging Late-Onset Alzheimer’s Disease Family Study Group. Genome-wide association study of Alzheimer’s disease with psychotic symptoms. Mol Psychiatry. 2012; 17:1316–27. https://doi.org/10.1038/mp.2011.125 [PubMed]
- 22. Lin E, Kuo PH, Liu YL, Yang AC, Kao CF, Tsai SJ. Effects of circadian clock genes and environmental factors on cognitive aging in old adults in a Taiwanese population. Oncotarget. 2017; 8:24088–98. https://doi.org/10.18632/oncotarget.15493 [PubMed]
- 23. Lin E, Kuo PH, Liu YL, Yang AC, Tsai SJ. Association and Interaction Effects of Interleukin-12 Related Genes and Physical Activity on Cognitive Aging in Old Adults in the Taiwanese Population. Front Neurol. 2019; 10:1065. https://doi.org/10.3389/fneur.2019.01065 [PubMed]
- 24. Lin E, Kuo PH, Liu YL, Yang AC, Tsai SJ. Polymorphisms of the DNA repair gene EXO1 modulate cognitive aging in old adults in a Taiwanese population. DNA Repair (Amst). 2019; 78:1–6. https://doi.org/10.1016/j.dnarep.2019.03.013 [PubMed]
- 25. Lin E, Tsai SJ, Kuo PH, Liu YL, Yang AC, Kao CF. Association and interaction effects of Alzheimer’s disease-associated genes and lifestyle on cognitive aging in older adults in a Taiwanese population. Oncotarget. 2017; 8:24077–87. https://doi.org/10.18632/oncotarget.15269 [PubMed]
- 26. Lin E, Tsai SJ, Kuo PH, Liu YL, Yang AC, Kao CF, Yang CH. The rs1277306 Variant of the REST Gene Confers Susceptibility to Cognitive Aging in an Elderly Taiwanese Population. Dement Geriatr Cogn Disord. 2017; 43:119–27. https://doi.org/10.1159/000455833 [PubMed]
- 27. Lin E, Tsai SJ, Kuo PH, Liu YL, Yang AC, Kao CF, Yang CH. The ADAMTS9 gene is associated with cognitive aging in the elderly in a Taiwanese population. PLoS One. 2017; 12:e0172440. https://doi.org/10.1371/journal.pone.0172440 [PubMed]
- 28. Erickson KI, Hillman C, Stillman CM, Ballard RM, Bloodgood B, Conroy DE, Macko R, Marquez DX, Petruzzello SJ, Powell KE, and FOR 2018 PHYSICAL ACTIVITY GUIDELINES ADVISORY COMMITTEE*. Physical Activity, Cognition, and Brain Outcomes: A Review of the 2018 Physical Activity Guidelines. Med Sci Sports Exerc. 2019; 51:1242–51. https://doi.org/10.1249/MSS.0000000000001936 [PubMed]
- 29. GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015; 348:648–60. https://doi.org/10.1126/science.1262110 [PubMed]
- 30. Kong Q, Min X, Sun R, Gao J, Liang R, Li L, Chu X. Effects of pharmacological treatments on hippocampal NCAM1 and ERK2 expression in epileptic rats with cognitive dysfunction. Oncol Lett. 2016; 12:1783–91. https://doi.org/10.3892/ol.2016.4882 [PubMed]
- 31. Vukojevic V, Mastrandreas P, Arnold A, Peter F, Kolassa IT, Wilker S, Elbert T, de Quervain DJ, Papassotiropoulos A, Stetak A. Evolutionary conserved role of neural cell adhesion molecule-1 in memory. Transl Psychiatry. 2020; 10:217. https://doi.org/10.1038/s41398-020-00899-y [PubMed]
- 32. Yang BZ, Kranzler HR, Zhao H, Gruen JR, Luo X, Gelernter J. Association of haplotypic variants in DRD2, ANKK1, TTC12 and NCAM1 to alcohol dependence in independent case control and family samples. Hum Mol Genet. 2007; 16:2844–53. https://doi.org/10.1093/hmg/ddm240 [PubMed]
- 33. Gelernter J, Yu Y, Weiss R, Brady K, Panhuysen C, Yang BZ, Kranzler HR, Farrer L. Haplotype spanning TTC12 and ANKK1, flanked by the DRD2 and NCAM1 loci, is strongly associated to nicotine dependence in two distinct American populations. Hum Mol Genet. 2006; 15:3498–507. https://doi.org/10.1093/hmg/ddl426 [PubMed]
- 34. Nelson EC, Lynskey MT, Heath AC, Wray N, Agrawal A, Shand FL, Henders AK, Wallace L, Todorov AA, Schrage AJ, Saccone NL, Madden PA, Degenhardt L, et al. ANKK1, TTC12, and NCAM1 polymorphisms and heroin dependence: importance of considering drug exposure. JAMA Psychiatry. 2013; 70:325–33. https://doi.org/10.1001/jamapsychiatry.2013.282 [PubMed]
- 35. Yang BZ, Kranzler HR, Zhao H, Gruen JR, Luo X, Gelernter J. Haplotypic variants in DRD2, ANKK1, TTC12, and NCAM1 are associated with comorbid alcohol and drug dependence. Alcohol Clin Exp Res. 2008; 32:2117–27. https://doi.org/10.1111/j.1530-0277.2008.00800.x [PubMed]
- 36. Rasmussen MB, Nielsen JV, Lourenço CM, Melo JB, Halgren C, Geraldi CV, Marques W
Jr , Rodrigues GR, Thomassen M, Bak M, Hansen C, Ferreira SI, Venâncio M, et al. Neurodevelopmental disorders associated with dosage imbalance of ZBTB20 correlate with the morbidity spectrum of ZBTB20 candidate target genes. J Med Genet. 2014; 51:605–13. https://doi.org/10.1136/jmedgenet-2014-102535 [PubMed] - 37. Mitchelmore C, Kjaerulff KM, Pedersen HC, Nielsen JV, Rasmussen TE, Fisker MF, Finsen B, Pedersen KM, Jensen NA. Characterization of two novel nuclear BTB/POZ domain zinc finger isoforms. Association with differentiation of hippocampal neurons, cerebellar granule cells, and macroglia. J Biol Chem. 2002; 277:7598–609. https://doi.org/10.1074/jbc.M110023200 [PubMed]
- 38. Xie Z, Ma X, Ji W, Zhou G, Lu Y, Xiang Z, Wang YX, Zhang L, Hu Y, Ding YQ, Zhang WJ. Zbtb20 is essential for the specification of CA1 field identity in the developing hippocampus. Proc Natl Acad Sci USA. 2010; 107:6510–15. https://doi.org/10.1073/pnas.0912315107 [PubMed]
- 39. Papenberg G, Ferencz B, Mangialasche F, Mecocci P, Cecchetti R, Kalpouzos G, Fratiglioni L, Bäckman L. Physical activity and inflammation: effects on gray-matter volume and cognitive decline in aging. Hum Brain Mapp. 2016; 37:3462–73. https://doi.org/10.1002/hbm.23252 [PubMed]
- 40. Montoya A, Elgueta D, Campos J, Chovar O, Falcón P, Matus S, Alfaro I, Bono MR, Pacheco R. Dopamine receptor D3 signalling in astrocytes promotes neuroinflammation. J Neuroinflammation. 2019; 16:258. https://doi.org/10.1186/s12974-019-1652-8 [PubMed]
- 41. Shao W, Zhang SZ, Tang M, Zhang XH, Zhou Z, Yin YQ, Zhou QB, Huang YY, Liu YJ, Wawrousek E, Chen T, Li SB, Xu M, et al. Suppression of neuroinflammation by astrocytic dopamine D2 receptors via αB-crystallin. Nature. 2013; 494:90–94. https://doi.org/10.1038/nature11748 [PubMed]
- 42. Lin E, Yang AC, Tsai SJ. Association between metabolic syndrome and cognitive function in old adults in a Taiwanese population. Taiwanese Journal of Psychiatry. 2017; 31:232–40.
- 43. Chen CH, Yang JH, Chiang CW, Hsiung CN, Wu PE, Chang LC, Chu HW, Chang J, Song IW, Yang SL, Chen YT, Liu FT, Shen CY. Population structure of Han Chinese in the modern Taiwanese population based on 10,000 participants in the Taiwan Biobank project. Hum Mol Genet. 2016; 25:5321–31. https://doi.org/10.1093/hmg/ddw346 [PubMed]
- 44. Sheehan B. Assessment scales in dementia. Ther Adv Neurol Disord. 2012; 5:349–58. https://doi.org/10.1177/1756285612455733 [PubMed]
- 45. Lin E, Kuo PH, Liu YL, Yang AC, Tsai SJ. Transforming growth factor-β signaling pathway-associated genes SMAD2 and TGFBR2 are implicated in metabolic syndrome in a Taiwanese population. Sci Rep. 2017; 7:13589. https://doi.org/10.1038/s41598-017-14025-4 [PubMed]
- 46. Lin E, Kuo PH, Liu YL, Yang AC, Kao CF, Tsai SJ. Association and interaction of APOA5, BUD13, CETP, LIPA and health-related behavior with metabolic syndrome in a Taiwanese population. Sci Rep. 2016; 6:36830. https://doi.org/10.1038/srep36830 [PubMed]
- 47. Hou SJ, Tsai SJ, Kuo PH, Liu YL, Yang AC, Lin E, Lan TH. An association study in the Taiwan Biobank reveals RORA as a novel locus for sleep duration in the Taiwanese Population. Sleep Med. 2020; 73:70–75. https://doi.org/10.1016/j.sleep.2020.04.008 [PubMed]
- 48. Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007; 81:559–75. https://doi.org/10.1086/519795 [PubMed]
- 49. Lin E, Kuo PH, Liu YL, Yang AC, Kao CF, Tsai SJ. Effects of circadian clock genes and health-related behavior on metabolic syndrome in a Taiwanese population: Evidence from association and interaction analysis. PLoS One. 2017; 12:e0173861. https://doi.org/10.1371/journal.pone.0173861 [PubMed]
- 50. Hosking L, Lumsden S, Lewis K, Yeo A, McCarthy L, Bansal A, Riley J, Purvis I, Xu CF. Detection of genotyping errors by Hardy-Weinberg equilibrium testing. Eur J Hum Genet. 2004; 12:395–99. https://doi.org/10.1038/sj.ejhg.5201164 [PubMed]
- 51. Lou XY, Chen GB, Yan L, Ma JZ, Zhu J, Elston RC, Li MD. A generalized combinatorial approach for detecting gene-by-gene and gene-by-environment interactions with application to nicotine dependence. Am J Hum Genet. 2007; 80:1125–37. https://doi.org/10.1086/518312 [PubMed]
- 52. Ward LD, Kellis M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016; 44:D877–81. https://doi.org/10.1093/nar/gkv1340 [PubMed]