Introduction
The global demographic transition toward aging populations poses a significant challenge for healthcare systems and societies [1, 2]. While life expectancy has steadily increased over the past century, the extension of the healthspan, the period of life free from chronic disease and functional decline, has remained unchanged. This widening gap between healthspan and lifespan contributes to morbidity, increased healthcare expenditure, and a reduced quality of life among older adults [3–6].
A key factor in this disparity is the difference between chronological age, a measure of elapsed lifetime, and biological age, which reflects the cumulative physiological experience of genetic, molecular, and environmental factors [7, 8]. Unlike chronological age, biological age (BA) can serve as a dynamic indicator of functional capacity, disease risk, and resilience reserve, which are much better predictors for all-cause mortality in the elderly [9–13]. However, to date, most aging studies have focused on single-omics datasets or narrowly targeted biomarker panels [14–27]. While these approaches have led to the identification of candidate biomarkers of aging, their correlations remain weak, and no standardized or universally validated biomarkers of aging currently exist - a persistent challenge in the field [28–30]. This limitation is partly due to the compartmentalized nature of earlier technologies, which were unable to comprehensively capture the multidimensional biological processes underlying aging. Advances in the quantification of BA, have opened new avenues for risk stratification and personalized interventions [31–34].
Recent technological advances have enabled the generation of high-quality, large-scale multi-omics datasets that encompass profiles of various biological samples across multiple domains, including genomics, transcriptomics, proteomics, metabolomics, lipidomics, and metagenomics [35–39]. This marks a critical inflection point in aging research, and for the first time, we possess the capability to interrogate aging as a systems-level process across multiple molecular and physiological levels in population-scale studies. Furthermore, the advent of artificial intelligence (AI) and large language model (LLM)-driven analytics provides unprecedented computational power to mine, interpret, and integrate these complex datasets [38, 40–44]. These tools enable the development of untargeted, data-driven pipelines for biomarker discovery, multidimensional modelling, and prediction of biological resilience in large cohorts. Cohort studies worldwide have begun employing these integrative multi-omics and AI-driven approaches to unravel the complexity of aging [45–48]. However, most existing datasets that use these analytical frameworks are predominantly derived from Western populations, leaving Asia, and particularly Indian populations, significantly underrepresented in global aging research [49, 50].
Interestingly, India presents a distinctive scientific opportunity in this regard, with an elderly population projected to reach 347 million by 2050, the country faces an urgent need to identify context-specific biomarkers of aging [51]. However, the country currently lacks comprehensive, high-quality reference datasets for aging and healthspan assessment. Clinical benchmarks and diagnostic cut-offs continue to rely heavily on data derived from Western cohorts, which may not accurately capture the physiological and molecular baselines of Indian populations [52, 53]. Moreover, the unique features of aging that may emerge from India’s extraordinary diversity in genetic ancestry, diet, lifestyle, environmental exposures, and rural-urban heterogeneity remain entirely unexplored [54–58]. These gaps highlight the critical need for a large-scale, integrative study to define India-specific signatures of biological aging and resilience, thereby enabling more precise, equitable, and population-relevant approaches to healthy aging.
To address these gaps, the Longevity India initiative was established to integrate cross-disciplinary expertise in aging biology, clinical research, and technology development [59]. Within this framework, the BHARAT Study (Biomarkers of Healthy Aging, Resilience, Adversity, and Transitions) has been designed as a large-scale, multi-modal, multi-parametric, and multi-omics cohort investigation. The study brings together researchers, clinicians, industry partners, and communities to generate high-quality data and address key questions related to healthy aging. By capturing the breadth of India’s demographic and environmental diversity, the BHARAT study aims to define biological age, resilience, and context-specific biomarkers that can inform predictive models and translational strategies for aging research.
Objectives
Primary objective
Develop and validate composite signatures of biological aging using cellular, molecular, clinical, and lifestyle metrics.
Secondary objectives
Identify resilience- and frailty-associated biomarkers.
Establish a validated frailty index tailored for Indian cohorts.
Build AI/ML-based predictive models for BA; disease risk and all-cause mortality.
Establish a scalable Indian biobank for aging research.
Establish baseline values for various biological features for India
Study team and collaborative framework
The BHARAT study is conducted under the framework of the Longevity India initiative, which comprises a multi-institutional, cross-disciplinary network of investigators from biological sciences, clinical medicine, engineering, and computational sciences.
The BHARAT Study is designed as a cross-sectional cohort study conceptualized between 2023-24, established partners, obtained ethical approvals in 2024 and initiated population sampling in 2025. The study employs a hub-and-spoke model in which IISc serves as the central hub while multiple clinical partners act as regional nodes for participant recruitment and primary data collection (Figure 1).
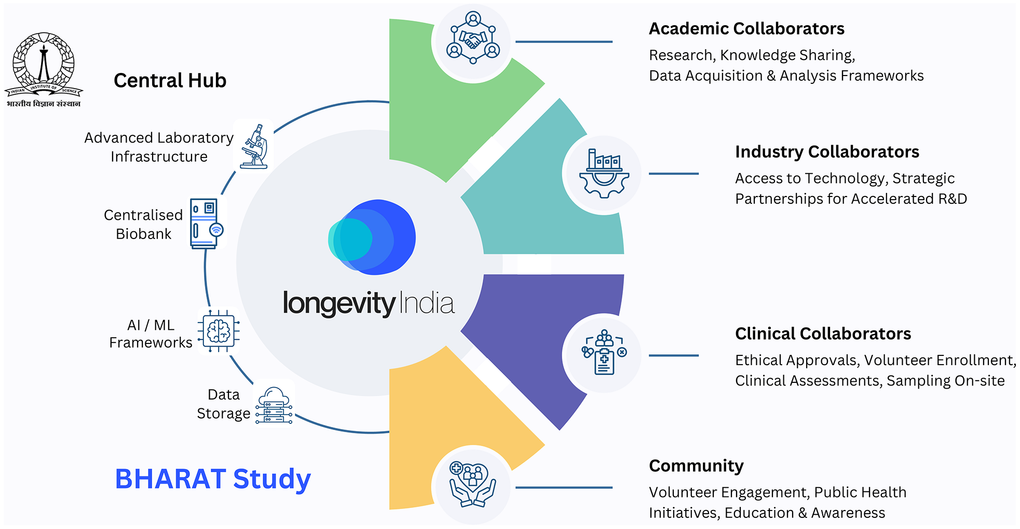
Figure 1. Study team and collaborative framework of the BHARAT study. Schematic representation of the hub-and-spoke collaborative model underpinning the BHARAT study, conducted under the Longevity India initiative. The Indian Institute of Science (IISc) functions as the central hub, providing advanced laboratory infrastructure, centralized biobanking, AI/ML and computational frameworks, and secure data storage and governance. Multiple regional clinical partners serve as spokes, facilitating participant recruitment, clinical phenotyping, and the collection of primary data and biospecimens.
Central hub (IISc)
The Indian Institute of Science serves as the central coordinating hub, hosting infrastructure and expertise for sampling, biobanking, data acquisition, mass-spectrometry based multi-omics data collection, immunophenotyping, development of data analytics and AI/ML pipelines. Dedicated computational infrastructure enables integration of high-resolution molecular data with clinical and lifestyle variables. The hub ensures centralized quality control, protocol standardization, and data governance across participating centres.
Partner institutions and organisations (Nodes)
The partner hospitals (MS Ramaiah Medical College, Bangalore Baptist Hospital, Sri Madhusudan Sai Institute of Medical Sciences and Research, Bangalore Medical College and Research Institute, and JSS Medical College) serve as regional nodes for participant recruitment and sampling. These include clinics and teaching hospitals equipped with trained medical personnel and infrastructure for screening, standardized sample and metadata collection, temporary storage, and transport of biological samples. Clinical teams are responsible for screening participants, administering questionnaires, conducting physical and cognitive assessments, and ensuring adherence to the inclusion-exclusion criteria. We have collaborated with several companies and communities (Vayah Vikas, Association of Healthcare Providers – India, Dementia India Alliance, Biopeak Wellness) across health and wellness, sports and fitness, senior care, and senior living to ensure diverse representation and recruitment of participants in the study, given that healthy individuals do not visit hospitals. These collaborators serve as additional nodes for identifying and enrolling participants in the study. Other integral nodes in this model include industry partners (TATA 1mg, Lal Path Labs, Decode Age, ARCA AI, Diagnoptics Inc., and others) that provide access to technologies and services, including metagenomics, epigenomics, and diagnostic tests for clinical benchmarking.
Expertise and governance
The collaborative network comprises clinicians, geriatricians, molecular biologists, engineers, computational biologists, and epidemiologists, supported by industry partners who contribute technological and analytical capabilities. The governance structure follows a centralized coordination model, where uniform operating procedures and data standards are maintained through regular inter-institutional meetings and audits.
The feasibility and scale of the study are enabled by a consortium of over 30 faculty members, 50 clinicians, and 100 trained research and technical staff spanning biology, medicine, and computational sciences. This integrated ecosystem allows seamless coordination between clinical and laboratory arms, ensuring consistency in sample processing, metadata capture, and analytical workflows.
Methods
Study design and setting
Currently, four centres across the state of Karnataka, India are operational, aiming to enrol 5,000 participants (based on approved ethical clearance), with each centre contributing approximately 1,000–2,000 individuals. The design is scalable, allowing expansion to new centres across India under the same standardized framework, with each additional site obtaining independent ethical approval through its community medicine department. This approach ensures both regional representation and long-term adaptability of the cohort.
Study participants
Inclusion criteria
Participants in the BHARAT Study are ‘apparently’ healthy volunteers aged 18 years or older, recruited from both hospital-linked and community-based settings. The inclusion criteria are designed to capture a representative cross-section of the healthy Indian population across sex, age, rural–urban strata, and, interestingly, for India, the vegetarian and meat-eating axes. Only individuals free from acute or chronic illnesses are enrolled to establish normative biological and physiological reference ranges across the lifespan.
Exclusion criteria
Participants with a known history, diagnosis, or clinical evidence of the following conditions are excluded from the study:
Chronic cardiac disorders: Present or past heart failure, cardiomyopathy (any type), or congenital valvular/structural heart disease (with or without treatment).
Chronic pulmonary disorders: Interstitial lung disease, severe chronic obstructive pulmonary disease (GOLD stage ≥3), or pulmonary hypertension of any etiology.
Chronic neurological disorders: Severe cerebrovascular events with residual neurological deficits, recurrent cerebrovascular accidents, neurodegenerative diseases (e.g., dementia, Parkinsonism, motor neuron disease), chronic demyelinating disorders, or progressive neuromuscular diseases (e.g., myasthenia, muscular dystrophy).
Chronic gastrointestinal disorders: Chronic liver disease (≥Stage 2), chronic pancreatitis, or inflammatory bowel disease.
Chronic kidney disease: Stage 3 or higher.
Autoimmune disorders: Any clinically diagnosed systemic or organ-specific autoimmune disease.
Transplant recipients: Individuals with prior solid-organ or bone marrow transplantation.
Active or prior malignancy (solid organ or haematological), including individuals currently undergoing or having completed cancer-directed therapy (chemotherapy, radiotherapy, immunotherapy, or hormonal therapy in the past or surgery in the last 1 year).
Recent infections or interventions: Fever or any active infection within the previous two weeks, or use of antibiotics during that period.
Recent alcohol intake: Consumption of alcohol within one-week preceding enrolment.
Recruitment and enrollment
Recruitment follows a dual-channel strategy involving (i) hospital-linked screening through participating clinical centres and (ii) community outreach to ensure demographic and geographic diversity. Potential volunteers are informed about the study objectives, procedures, and eligibility requirements prior to screening. Those meeting the inclusion criteria undergo clinical evaluation and questionnaire-based assessments before providing written informed consent (Figure 2).
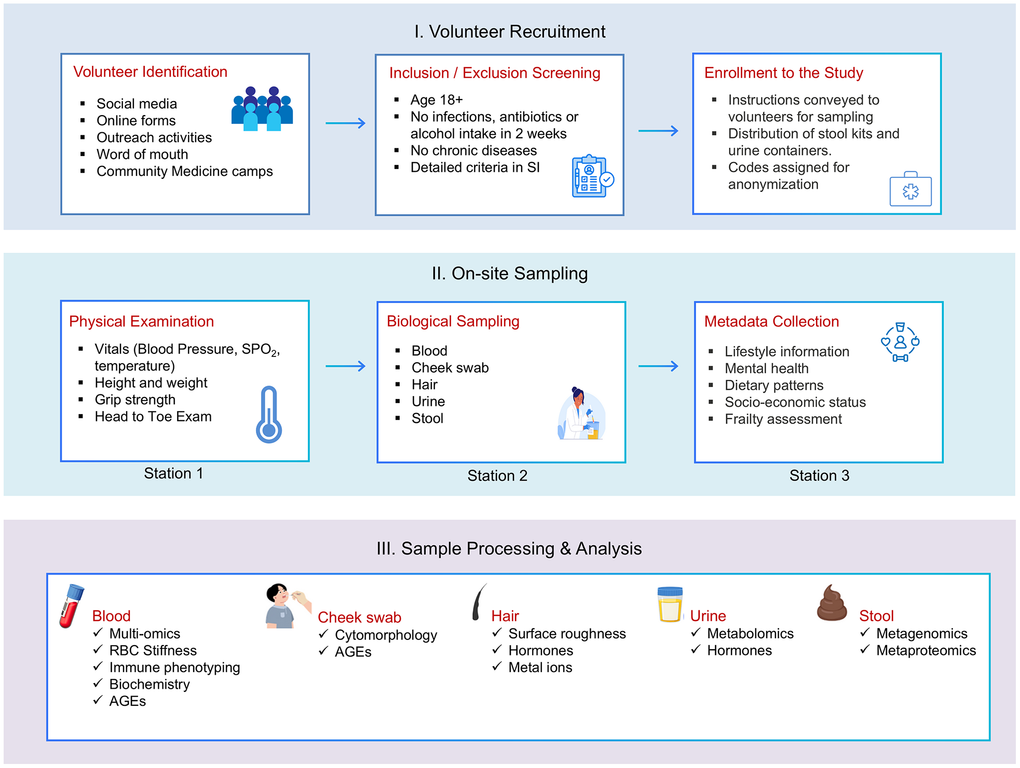
Figure 2. BHARAT study workflow for recruitment, sampling, and analysis. Schematic overview of the three-stage study pipeline comprising (I) volunteer recruitment and screening, (II) on-site clinical examination, biospecimen collection, and metadata acquisition, and (III) sample processing and multi-modal analysis. Blood, cheek swab, hair, urine, and stool samples are subjected to complementary molecular, cellular, biochemical, and omics-based assays to enable integrated phenotyping of biological aging across cohorts.
Each eligible participant is assigned a unique identification code enabling secure linkage of clinical, lifestyle, and omics data. This coding system also facilitates the de-identification of data across sites and ensures traceability within the biobanking framework.
Participant stratification
Participants are stratified into five age groups representing distinct life stages: 18–29, 30–44, 45–59, 60–74, and ≥75 years. Each age stratum maintains balanced representation by sex and rural–urban distribution, capturing India’s demographic diversity. The study design enables both cross-sectional analysis of age-associated biological and physiological variations and future longitudinal extensions through re-sampling and follow-up of select cohorts.
Metadata collection
All participants undergo a standardized, multi-modal assessment, including structured questionnaires that cover sociodemographic, dietary, physical activity, psychosocial, and quality-of-life information, as well as anthropometric measurements and a clinical vital assessment. The details of the metadata questionnaire are outlined in Supplementary Information.
Digital infrastructure
All participant data are collected at collaborating hospital and community sites using a secure, custom-designed electronic data capture platform developed in collaboration with ARCA AI. Data entered at participating sites are securely transmitted to the Indian Institute of Science (IISc), which serves as the centralized data repository and institutional custodian for the BHARAT Study. IISc hosts the synchronized database for all clinical, questionnaire, laboratory, and multi-omics data, under institutional oversight and role-based access control. Each participant is assigned a unique anonymized study identifier, enabling deterministic linkage across data modalities while ensuring strict separation of identifiable information from analytical datasets.
Biological sampling
The following biological specimens are collected from all participants: blood, urine, stool, cheek swab, and hair. All biological specimens are collected under standardized conditions to minimize pre-analytical variability and ensure consistency across clinical sites. Participants fast overnight (8–10 hours), and sampling is conducted the next day between 7:30 and 9:30 a.m. by trained clinical staff, following aseptic procedures.
Blood
A total of 25 mL of venous blood is drawn using sterile, vacuum-sealed collection tubes. All vials are labelled with pre-assigned barcodes corresponding to participant IDs and immediately logged in the ODK-linked laboratory management system. EDTA-coated tubes are used for blood smears, whole blood, and plasma-based assays. Serum separator tubes (SST) are used for serum-based assays, and Fluoride-coated vials for blood biochemical assays. After collection, the EDTA tubes are gently inverted to prevent clotting. Peripheral blood smears for cytochemistry analysis are prepared and fixed with 100% methanol within 45 minutes of collection. Plasma separation is performed by centrifugation at 1200 × g for 10 minutes, followed by plasma extraction, aliquoting, and snap-freezing in liquid nitrogen. SST tubes are kept upright at room temperature to facilitate blood clotting for 10 minutes, followed by centrifugation at 1200 × g for 10 minutes. Serum is separated, aliquoted, and stored at 4° C for analysis or snap-frozen in liquid nitrogen for biobanking. Blood biochemistry is performed on fresh samples at a partner diagnostic laboratory. The complete list of tests is included in Supplementary Information.
Urine
Participants are provided with sterile urine collection containers and 5 mL cryovials. Midstream urine is collected in the morning, transferred aseptically into cryovials, snap-frozen in liquid nitrogen, and transported for biobanking.
Stool
Participants are given a stool collection kit (Decode Age) containing sterile containers with DNA stabilizer, spatulas, gloves, and illustrated instructions. Samples are collected within 24 hours of the sampling visit and stored temporarily at room temperature. The samples are collected the following day, transported and the aliquots are transferred to cryovials for storage at –20° C.
Cheek swab
Buccal epithelial cells are obtained using sterile cotton swabs. Each participant rinsed the mouth before collection. The swab is rotated along the inner cheek surface for 30 seconds on each side. Cells are transferred onto a marked region on a glass slide using a PAP pen, fixed in 10% neutral buffered formalin (NBF) for 45 minutes, and stored in phosphate-buffered saline (PBS) at 4° C until processing.
Hair
Approximately 10 strands of scalp hair are cut 0.5 cm close to the scalp from the posterior vertex using sterile scissors. The proximal 3 cm of each strand is retained, representing approximately three months of growth. Hair samples are stored at room temperature in a desiccated condition for subsequent analysis.
Transport and storage
All samples are transported to the central biobank at IISc using temperature-controlled containers (cool boxes with ice packs or liquid nitrogen dewars). Chain-of-custody logs are maintained for each sample batch. At IISc, samples are organized by specimen type and transferred into long-term storage under the following conditions:
Blood, serum, plasma and urine samples are stored at –150° C.
Cheek swabs and blood smears are stored at 4° C
Hair samples are stored at ambient temperature.
Each sample is assigned a unique digital code linking metadata, processing status, and future analytical use within the Longevity India biobank infrastructure.
Sample processing and multi-omics analysis
Following receipt at the IISc central biobanking facility, samples are processed within a controlled biospecimen handling pipeline in accordance with the BHARAT Study SOPs. The objective is to generate integrative molecular datasets spanning genetic, epigenetic, proteomic, metabolomic, lipidomic, immunophenotypic, and microbiome dimensions. Our multi-omics strategy integrates:
Epigenomics: DNA methylation profiling using Infinium Methylation Screening Array (270,000 CpG sites) and SNP genotyping using Axiom Asia Precision Medicine Array (750,000 markers tailored for Southeast Asian populations).
Proteomics: Untargeted plasma proteomics via nano-liquid chromatography-tandem mass spectrometry (nLC-MS/MS (EvoSEP Eno), Bruker timsTOF HT platform), complemented by targeted protein/peptide panels and cytokine panels (aptamer-based platforms).
Metabolomics and Lipidomics: Untargeted plasma and urine metabolomics via high-resolution LC-MS (Agilent UHPLC coupled to Bruker timsMetabo), employing modified Bligh-Dyer and Matyash extraction protocols.
Metagenomics: Shotgun metagenomic sequencing of stool samples via Oxford Nanopore Technology (ONT) platform for species- and function-level characterization of gut microbial communities.
Immune Phenotyping: Deep immunophenotyping using Beckman Coulter DuraClone™ dried antibody panels (T cell, B cell, granulocyte panels) via flow cytometry, assessing naïve, memory, senescent, and activated immune subsets.
Cellular Aging Markers: Advanced glycation end-product (AGE) quantification via immunocytochemistry on peripheral blood smears and buccal cells, red blood cell stiffness assessment via microfluidic devices, and hair surface roughness analysis via optical imaging.
Detailed protocols of sample processing and data collection are available in the Supplementary Information section.
Data management and integration
All multi-modal and multi-omics data are managed and stored at IISc, which maintains the common repository. Data architecture includes a local server (master node) with role-based access and routine backups, adhering to both HIPAA standards and Indian Data Protection and Privacy laws. We track samples using in-house laboratory information management software with data audit trails and instrument API integrations. We integrate all datasets (clinical metadata, laboratory measurements, multi-omics data) by linking each unique anonymized participant identifier to specimen and assay-level identifiers. We perform preprocessing, normalization, and multi-omics integration using standardized pipelines before downstream modelling of biological age, resilience, metabolic indices, and frailty using machine learning frameworks.
Discussion
The BHARAT Study represents India’s first large-scale, multi-omics investigation into biological aging and health trajectories, addressing a critical gap in global aging research. While landmark cohorts such as the UK Biobank (500,000 participants, predominantly European ancestry) [60], the All of Us Research Program (245,388 participants, 77% from underrepresented populations but primarily US-based) [61], and the Framingham Heart Study [62] have profoundly shaped our understanding of aging biology, Asian populations, which represent nearly 60% of the global population, remain conspicuously underrepresented within such studies (Figure 3). Among the limited Asian cohorts, the Singapore-based ABIOS study (420 participants, aged 21 and above, cross-sectional study design with epigenomics, proteomics, and metabolomics) [63] provides the closest methodological parallel to the BHARAT cohort, albeit at a substantially smaller scale (Figure 4). The Japanese Tohoku Medical Megabank (encompassing 150,000 participants with genomics and metabolomics) primarily focuses on disaster-affected populations and their health outcomes, although its tertiary assessments from 2021 have now begun to examine aging trajectories [64].
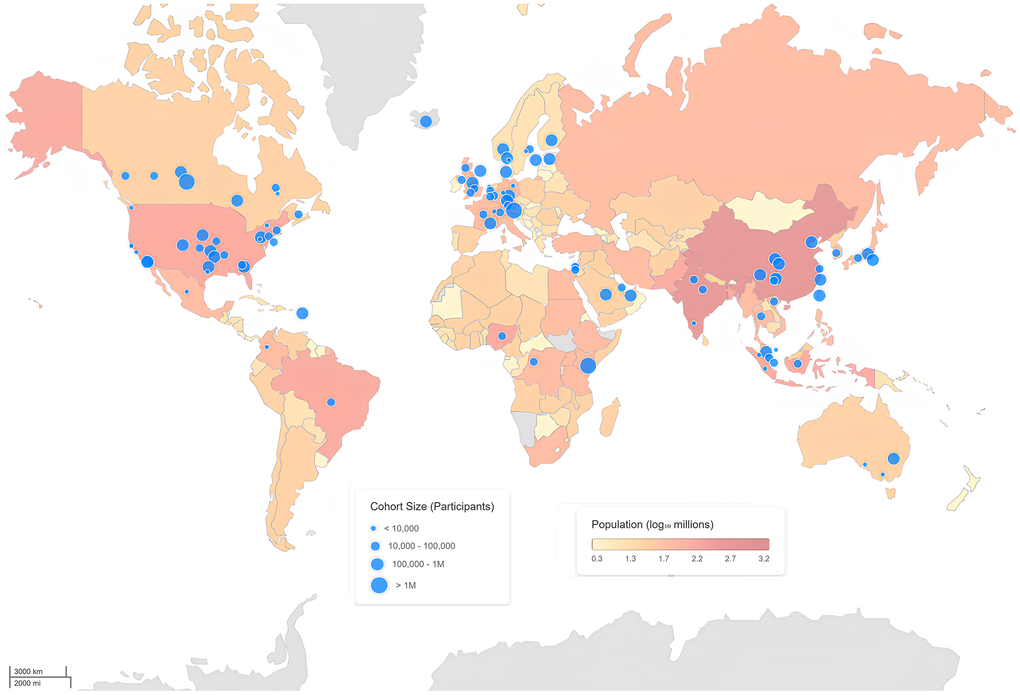
Figure 3. Global distribution of multi-omics cohorts studying healthy populations. World map depicting the geographic distribution of multi-omics studies conducted in healthy volunteers. Blue circles indicate the location of individual cohorts, with circle size proportional to cohort size (number of participants). Country-level population size (log scale, millions) is overlaid as a choropleth to provide demographic context. The map highlights the global concentration and regional disparities of existing multi-omics cohorts generating at least one omics dataset from biological samples (See Supplementary Information for details of datasets and analysis).
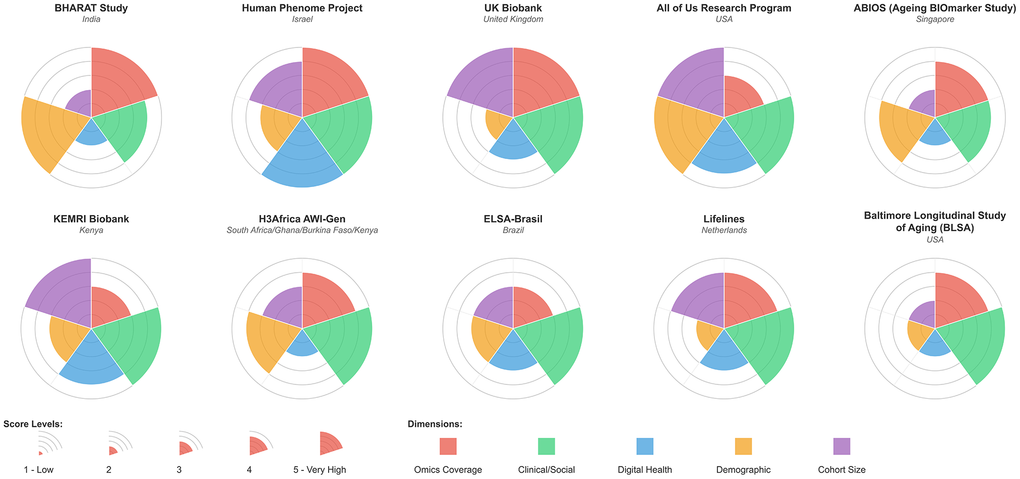
Figure 4. Radar plots of representative worldwide multi-omics cohorts scored on 5 different criteria: Omics coverage, clinical/social metadata collection, use of digital health tools, demographic coverage, and cohort size. Each category is scored on a scale of 1-5. The in-depth criteria and justification for scoring are given in Supplementary Information.
Despite these advances, the datasets that train most aging clocks, molecular risk predictors, and longevity biomarkers are derived almost exclusively from Western populations, limiting their applicability in Asian contexts. Population-level differences in genetic ancestry, infection burden, nutrition, environmental exposures, socioeconomic backgrounds and healthcare access have a significant influence on biological aging signatures [65–68]. The absence of Indian representation in such datasets, therefore, constrains both the calibration and interpretive power of aging models. There is a pressing need to establish population-specific, multi-dimensional datasets that reflect the unique biological and socio-environmental heterogeneity of India. The BHARAT Study aims to directly address this gap by establishing an integrated, multi-omics study cohort of Indian adults across diverse geographic, cultural, and socio-economic backgrounds. Its sampling framework is geared to balance representation across sex, age, income, and rural-urban strata, thereby creating a robust reference landscape for understanding variability in physiological and molecular parameters. Current clinical benchmarks in India are largely extrapolated from Western reference intervals, with few longitudinal or cross-sectional datasets defining normative ranges for healthy Indian adults, such as those from Phenome India [69]. By systematically characterizing healthy individuals across strata, this study provides the first dataset to recalibrate biomarker thresholds, reference intervals, and health trajectories specifically within the Indian context. Longitudinal studies in health trajectories and disease burden assessment have been done in India by two major focused studies, The Longitudinal Ageing Study in India (LASI) [70] and Indian Study of Healthy Ageing (ISHA) [71]. Both studies collect high-quality anthropomorphic data and basic molecular features but lack in-depth untargeted omics profiling, which is the unique aspect of the BHARAT.
This study builds upon methodological foundations established by pioneering aging cohorts while addressing critical gaps in population representation. The Stanford 1000 Immunomes Project (n=1,001, ages 8-96, 12-year longitudinal follow-up), demonstrated that deep immune phenotyping combined with machine learning can identify inflammatory biomarkers, most notably the iAge clock centered on CXCL9 that predicts multimorbidity, frailty, and cardiovascular aging independent of chronological age [46]. This work established inflammaging as a quantifiable, mechanistically actionable hallmark of biological aging and pioneered the use of deep autoencoders for high-dimensional immunological data, methods we aim to adapt for our own multi-omics integration. The InCHIANTI study (Italy, n=1,200, ages 65+) provided proteomic and metabolomic aging signatures validating that circulating biomarkers track with physical function, frailty, and mortality across European populations [72]. In India, the Council of Scientific and Industrial Research (CSIR) Phenome India initiative announced a large-scale multi-omics cohort across 10,000 CSIR employees, pensioners, and their spouses for cardiometabolic disease risk prediction from diverse geographic regions [69]. Phenome India exclusively recruits CSIR employees and their families, a relatively homogenous, urban, educated cohort with stable employment, limiting generalizability to India’s broader socioeconomic and rural-urban diversity. BHARAT complements this effort by targeting community-based recruitment spanning rural and urban sites across diverse socioeconomic strata, with explicit focus on aging biology and developing population-specific molecular aging features validated against health trajectories.
A distinguishing strength of the BHARAT study lies in its multi-omics multi-feature data collection design. While pioneering global aging cohorts such as the Stanford 1000 Immunomes Project, Baltimore Longitudinal Study of Aging [73], and Multi-Ethnic Study of Atherosclerosis [74] established foundational frameworks for immune aging, biological clocks, and phenotypic trajectories, they predominantly employed targeted proteomic platforms such as Olink or SomaLogic. These aptamer-based approaches, though highly reproducible and scalable, are limited to pre-selected analytes and cannot discover novel biomarkers outside their fixed panels [63, 75]. BHARAT incorporates an unbiased, discovery-driven LC-MS/MS pipeline for proteomics, metabolomics, lipidomics, and molecular profiling, enabling hypothesis-free detection of novel metabolites, post-translational modifications, and population-specific signatures absent from Western reference databases. These layers are complemented by targeted panels for hormones, metabolites, cytokines, and clinical biochemistry, creating a comprehensive, clinically relevant, and usable biochemical framework for integrative analyses. Further, deep immune phenotyping adds granularity to cellular and functional heterogeneity within the Indian context, a critical aspect in a tropical country with high exposure to infectious diseases and chronic inflammatory states [76]. Together, these datasets enable the identification of systemic, multi-modal biomarkers that capture health, resilience, transitions through various health phases in life and quality of health decline across the aging spectrum.
Furthermore, the microbiome analysis included in the study extends beyond conventional 16S rRNA-based approaches and employs nanopore-based whole-metagenome shotgun sequencing, enabling species- and function-level characterization of microbial communities and their derived metabolites. Given India’s extraordinary dietary diversity, fermentation practices, infectious disease burden and sanitation gradients, such data are important for decoding microbiome-mediated influences on metabolic and immune aging. Standardized protocols for sample collection, stabilization, and sequencing depth have been optimized for reproducibility, ensuring technical comparability with other global cohorts while retaining contextual specificity.
In addition to deep molecular profiling described above, the study’s metadata and phenotypic framework provide a rich contextual foundation for multi-omics interpretation. Comprehensive data on clinical history, comorbidities, anthropometry, diet, sleep, physical activity, mental health (DASS-21), cognitive performance (MMSE), quality of life (WHO-QOL) and frailty are systematically collected through standardized instruments and approaches by the clinical teams. These have been translated and culturally adapted into regional languages to ensure conceptual equivalence and reduce respondent bias.
Collectively, the BHARAT Study is positioned to deliver multiple scientific outputs. These include (i) validated biomarkers and reference datasets for healthy aging in Indian adults, (ii) integrative pipelines for multi-omics data analysis and biological age or physiologically relevant wellness state estimation, (iii) retraining and calibration of biological clocks for non-Western populations, and (iv) generation of high-quality data for machine learning models and diagnostic development. In the broader context of global geroscience, this study establishes a scientifically rigorous framework for understanding biological aging in a highly diverse population. By generating interoperable, high-resolution data suited for mechanistic modelling and machine learning, BHARAT contributes a resource of global relevance that would be capable of refining universal models of aging biology while revealing novel, population-specific pathways that inform prevention and intervention strategies. Integrating these models with clinical and lifestyle data will facilitate the development of personalized risk and overall mortality prediction tools and diagnostic panels tailored to Indian populations.
Beyond biomarker discovery, BHARAT’s cross-layered datasets can uncover mechanistic nodes that link metabolic, immune, and microbial alterations to aging phenotypes, unlocking targets for therapeutic interventions. These insights can qualify and enrich the design of dietary, pharmacological, and lifestyle interventions aimed at promoting healthy aging. In parallel, the study’s biobanking infrastructure and standardized analytical pipelines will enable partnerships with national and international consortia, ensuring that BHARAT contributes both to global knowledge and to the development of contextually relevant strategies for extending healthspan.
Supplementary Materials
Author Contributions
Conceptualization and Study Design: Deepak Kumar Saini, Harshavardhan Rao B, Ramray Bhat, Srimonta Gayen, Shantanu Shukla, Siddharth Jhunjhunwala, Prosenjit Sen, Prabhdeep Kaur, Suramya Asthana, Amruth Deepak Bhat, Swati Negi, and Seema Yadav; Sampling: Gayathri Mahadevan, Lakshmi Kothegala, Anupama Sudhakaran, Annwesha Roy, Meet Makwana, Rahul Patel, Caroline Elizabeth George, Harshavardhan Rao B.; Manuscript Preparation: Suramya Asthana, Amruth Deepak Bhat, Swati Negi, Seema Yadav, Srimonta Gayen, Shantanu Shukla, Siddharth Jhunjhunwala, Ramray Bhat and Deepak Kumar Saini.
Study contributors
Longevity India Team: Ambarish Ghosh, Ananya Sarkar, Anchita Gopikrishnan, Annapoorni Rangarajan, Anuva R, Arvind Ramanathan, Ayushi Verma, Bishal Singh, Chiranjib Bhattacharya, Inbasekar C, Jalak Mehta, Jayant Sehgal, Kaushik Chatterjee, Mohit Kumar Jolly, Narendra Dixit, Nikhil Gandasi, Nischitha Baliga, Sagarika Prabhakar, Sanjeeva Srivastava, Sanjiv Sambandan, Shovamayee Maharana, Shweta Thakar, Subba Rao GS, Swaminathan S, Tavpritesh Sethi, Vinod Bhat, Vishal Rao US, and Yogesh.
Clinical Partners: Maddipati Tatarao, Suman Sarah Varghese, Manjula KS, S Praveena, Sharon Rimai (Bangalore Baptist Hospital); Shreya Thadigotla, Prateek Reddy, Drashti Godhani, Anagha Pritvi Raj, Bhoomika Y., Harshit V. (MS Ramaiah Hospital); Sathish Babu K.N., Phaneendra Mallekavu (SMSIMSR); Anil S. Bilimale, Sheeba B, Divya K L, Hemanth R, Aishwarya N (JSS Medical College); Shreya Gudi, Raghvendra BM (BMCRI); Amit Bahuguna, D Navya, Devyani Sapra, Ashwin P Nair, Vishal Gautam (Command Hospital Bengaluru).
Industry Partners: Jagan L, Aditi Muglikar, Darshit Patel (Decode Age); John Mulder (Diagnoptics Inc.); Tushar Asthana, Shibu V Ignatious, Ramya Satish Kumar, Karthik Iyer, Rohit Rajan (ARCA AI); Gaurav Agarwal, Shanmugam RK, Prashant Nag, Arpita Roy Dam, Vinisha Nahata, Swati Gupta (TATA 1mg Labs); Vandana Lal, Alok Sharma, Archana Grover (Lal Path Labs).
Acknowledgements
We acknowledge the collective efforts and commitment of all collaborators and contributors who have been integral to the conception, execution, and ongoing conduct of this study. This work is supported by financial and institutional support from the Indian Council of Medical Research, Prashanth Prakash Family Foundation, Kris Gopalakrishnan, Kiran Mazumdar-Shaw, and several other donors and supporters whose contributions have been critical to enabling large-scale, population-based aging research in India. We extend our sincere appreciation to the Longevity India Advisory Board for their strategic guidance. We also acknowledge the Division of Interdisciplinary Sciences (IDS) at the Indian Institute of Science, and in particular the leadership and institutional support provided by Prof. Navakanta Bhat, which has been instrumental in fostering an interdisciplinary environment conducive to this research. Finally, we express our deepest gratitude to the community volunteers and study participants who generously contributed their time, biological samples, and data. We also thank the field teams, clinical staff, and student volunteers for their dedication, which ensured rigorous data collection, ethical conduct, and operational excellence across study sites.
Conflicts of Interest
DKS is a scientific advisor for Biopeak Wellness and the Founder of Rejuvome Therapeutics. These companies have no role in the current study design. None of the industry partners has any role in the study design or interpretation. Other academic authors have no conflicts to declare.
Ethical Statement and Consent
The overall research protocol for the BHARAT study has been reviewed and approved by the Institutional Human Ethics Committee of the Indian Institute of Science (IISc), Bengaluru (IHEC No. 14/21.03.2024). Ethical approvals for participant recruitment, clinical assessments, and biological sample collection have been obtained independently by the respective clinical collaborating institutions in accordance with regulatory requirements. These include M.S. Ramaiah Medical College (Approval No. MSRMC/EC/AP-13/11-2023), Bangalore Baptist Hospital (Approval No. BBH/IRB/2025/03), Sri Madhusudan Sai Institute of Medical Sciences and Research (Approval No. IEC/Certificate/34/2024), Bangalore Medical College and Research Institute (Approval No. BMCRI/EC/20/23-24), and JSS Medical College (Approval No. JSSMC/IEC/24102025/23NCT/2025-26). All study procedures are conducted in compliance with national ethical guidelines.
Funding
Funding for this work has been provided by the Indian Council of Medical Research (ICMR grant no. EMDR/15/CAR-2024-01-000071) and philanthropic support from Prashanth Prakash Family Foundation, Kris Gopalakrishnan and Dr. Kiran Mazumdar-Shaw.
References
- 1. Chang AY, Bolongaita S, Cao B, Castro MC, Karlsson O, Mao W, Norheim OF, Ogbuoji O, Jamison DT. Epidemiological and demographic trends and projections in global health from 1970 to 2050: a descriptive analysis from the third Lancet Commission on Investing in Health, Global Health 2050. Lancet. 2025; 406:940–9. https://doi.org/10.1016/S0140-6736(25)00902-X [PubMed]
- 2. GBD 2021 Demographics Collaborators. Global age-sex-specific mortality, life expectancy, and population estimates in 204 countries and territories and 811 subnational locations, 1950-2021, and the impact of the COVID-19 pandemic: a comprehensive demographic analysis for the Global Burden of Disease Study 2021. Lancet. 2024; 403:1989–2056. https://doi.org/10.1016/S0140-6736(24)00476-8 [PubMed]
- 3. Garmany A, Terzic A. Healthspan-lifespan gap differs in magnitude and disease contribution across world regions. Commun Med (Lond). 2025; 5:381. https://doi.org/10.1038/s43856-025-01111-2 [PubMed]
- 4. Garmany A, Yamada S, Terzic A. Longevity leap: mind the healthspan gap. NPJ Regen Med. 2021; 6:57. https://doi.org/10.1038/s41536-021-00169-5 [PubMed]
- 5. Crimmins EM. Lifespan and Healthspan: Past, Present, and Promise. Gerontologist. 2015; 55:901–11. https://doi.org/10.1093/geront/gnv130 [PubMed]
- 6. Burch JB, Augustine AD, Frieden LA, Hadley E, Howcroft TK, Johnson R, Khalsa PS, Kohanski RA, Li XL, Macchiarini F, Niederehe G, Oh YS, Pawlyk AC, et al. Advances in geroscience: impact on healthspan and chronic disease. J Gerontol A Biol Sci Med Sci. 2014; 69:S1–3. https://doi.org/10.1093/gerona/glu041 [PubMed]
- 7. Levine ME. Modeling the rate of senescence: can estimated biological age predict mortality more accurately than chronological age? J Gerontol A Biol Sci Med Sci. 2013; 68:667–74. https://doi.org/10.1093/gerona/gls233 [PubMed]
- 8. Mathur A, Taurin S, Alshammary S. New insights into methods to measure biological age: a literature review. Front Aging. 2024; 5:1395649. https://doi.org/10.3389/fragi.2024.1395649 [PubMed]
- 9. Bortz J, Guariglia A, Klaric L, Tang D, Ward P, Geer M, Chadeau-Hyam M, Vuckovic D, Joshi PK. Biological age estimation using circulating blood biomarkers. Commun Biol. 2023; 6:1089. https://doi.org/10.1038/s42003-023-05456-z [PubMed]
- 10. Brinkley TE, Justice JN, Basu S, Bauer SR, Loh KP, Mukli P, Ng TK, Turney IC, Ferrucci L, Cummings SR, Kritchevsky SB. Research priorities for measuring biologic age: summary and future directions from the Research Centers Collaborative Network Workshop. Geroscience. 2022; 44:2573–83. https://doi.org/10.1007/s11357-022-00661-w [PubMed]
- 11. Drewelies J, Hueluer G, Duezel S, Vetter VM, Pawelec G, Steinhagen-Thiessen E, Wagner GG, Lindenberger U, Lill CM, Bertram L, Gerstorf D, Demuth I. Using blood test parameters to define biological age among older adults: association with morbidity and mortality independent of chronological age validated in two separate birth cohorts. Geroscience. 2022; 44:2685–99. https://doi.org/10.1007/s11357-022-00662-9 [PubMed]
- 12. Wu JW, Yaqub A, Ma Y, Koudstaal W, Hofman A, Ikram MA, Ghanbari M, Goudsmit J. Biological age in healthy elderly predicts aging-related diseases including dementia. Sci Rep. 2021; 11:15929. https://doi.org/10.1038/s41598-021-95425-5 [PubMed]
- 13. Belsky DW, Caspi A, Houts R, Cohen HJ, Corcoran DL, Danese A, Harrington H, Israel S, Levine ME, Schaefer JD, Sugden K, Williams B, Yashin AI, et al. Quantification of biological aging in young adults. Proc Natl Acad Sci USA. 2015; 112:E4104–10. https://doi.org/10.1073/pnas.1506264112 [PubMed]
- 14. Sehgal R, Markov Y, Qin C, Meer M, Hadley C, Shadyab AH, Casanova R, Manson JE, Bhatti P, Moore AZ, Crimmins EM, Hagg S, Assimes TL, et al. Systems Age: a single blood methylation test to quantify aging heterogeneity across 11 physiological systems. Nat Aging. 2025; 5:1880–96. https://doi.org/10.1038/s43587-025-00958-3 [PubMed]
- 15. Argentieri MA, Xiao S, Bennett D, Winchester L, Nevado-Holgado AJ, Ghose U, Albukhari A, Yao P, Mazidi M, Lv J, Millwood I, Fry H, Rodosthenous RS, et al. Proteomic aging clock predicts mortality and risk of common age-related diseases in diverse populations. Nat Med. 2024; 30:2450–60. https://doi.org/10.1038/s41591-024-03164-7 [PubMed]
- 16. Liu Z, Kuo PL, Horvath S, Crimmins E, Ferrucci L, Levine M. A new aging measure captures morbidity and mortality risk across diverse subpopulations from NHANES IV: A cohort study. PLoS Med. 2018; 15:e1002718. https://doi.org/10.1371/journal.pmed.1002718 [PubMed]
- 17. Fong S, Pabis K, Latumalea D, Dugersuren N, Unfried M, Tolwinski N, Kennedy B, Gruber J. Principal component-based clinical aging clocks identify signatures of healthy aging and targets for clinical intervention. Nat Aging. 2024; 4:1137–52. https://doi.org/10.1038/s43587-024-00646-8 [PubMed]
- 18. Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, Klotzle B, Bibikova M, Fan JB, Gao Y, Deconde R, Chen M, Rajapakse I, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell. 2013; 49:359–67. https://doi.org/10.1016/j.molcel.2012.10.016 [PubMed]
- 19. Bocklandt S, Lin W, Sehl ME, Sánchez FJ, Sinsheimer JS, Horvath S, Vilain E. Epigenetic predictor of age. PLoS One. 2011; 6:e14821. https://doi.org/10.1371/journal.pone.0014821 [PubMed]
- 20. Chen W, Qian W, Wu G, Chen W, Xian B, Chen X, Cao Y, Green CD, Zhao F, Tang K, Han JD. Three-dimensional human facial morphologies as robust aging markers. Cell Res. 2015; 25:574–87. https://doi.org/10.1038/cr.2015.36 [PubMed]
- 21. Cole JH, Poudel RP, Tsagkrasoulis D, Caan MW, Steves C, Spector TD, Montana G. Predicting brain age with deep learning from raw imaging data results in a reliable and heritable biomarker. Neuroimage. 2017; 163:115–24. https://doi.org/10.1016/j.neuroimage.2017.07.059 [PubMed]
- 22. Mutz J, Iniesta R, Lewis CM. Metabolomic age (MileAge) predicts health and life span: A comparison of multiple machine learning algorithms. Sci Adv. 2024; 10:eadp3743. https://doi.org/10.1126/sciadv.adp3743 [PubMed]
- 23. Krištić J, Vučković F, Menni C, Klarić L, Keser T, Beceheli I, Pučić-Baković M, Novokmet M, Mangino M, Thaqi K, Rudan P, Novokmet N, Sarac J, et al. Glycans are a novel biomarker of chronological and biological ages. J Gerontol A Biol Sci Med Sci. 2014; 69:779–89. https://doi.org/10.1093/gerona/glt190 [PubMed]
- 24. Enroth S, Enroth SB, Johansson Å, Gyllensten U. Protein profiling reveals consequences of lifestyle choices on predicted biological aging. Sci Rep. 2015; 5:17282. https://doi.org/10.1038/srep17282 [PubMed]
- 25. Lehallier B, Shokhirev MN, Wyss-Coray T, Johnson AA. Data mining of human plasma proteins generates a multitude of highly predictive aging clocks that reflect different aspects of aging. Aging Cell. 2020; 19:e13256. https://doi.org/10.1111/acel.13256 [PubMed]
- 26. Alpert A, Pickman Y, Leipold M, Rosenberg-Hasson Y, Ji X, Gaujoux R, Rabani H, Starosvetsky E, Kveler K, Schaffert S, Furman D, Caspi O, Rosenschein U, et al. A clinically meaningful metric of immune age derived from high-dimensional longitudinal monitoring. Nat Med. 2019; 25:487–95. https://doi.org/10.1038/s41591-019-0381-y [PubMed]
- 27. Meyer DH, Schumacher B. BiT age: A transcriptome-based aging clock near the theoretical limit of accuracy. Aging Cell. 2021; 20:e13320. https://doi.org/10.1111/acel.13320 [PubMed]
- 28. Veytsman B, Cui T, Baranova A. Practical Detection of Biological Age: Why It Is not a Trivial Task. Springer, Cham; 2019: 7–21. https://doi.org/10.1007/978-3-030-24970-0_2
- 29. Herzog CM, Goeminne LJ, Poganik JR, Barzilai N, Belsky DW, Betts-LaCroix J, Chen BH, Chen M, Cohen AA, Cummings SR, Fedichev PO, Ferrucci L, Fleming A, et al., and Biomarkers of Aging Consortium. Challenges and recommendations for the translation of biomarkers of aging. Nat Aging. 2024; 4:1372–83. https://doi.org/10.1038/s43587-024-00683-3 [PubMed]
- 30. Perri G, French C, Agostinis-Sobrinho C, Anand A, Antarianto RD, Arai Y, Baur JA, Cauli O, Clivaz-Duc M, Colloca G, Demetriades C, de Lucia C, Di Gessa G, et al. An Expert Consensus Statement on Biomarkers of Aging for Use in Intervention Studies. J Gerontol A Biol Sci Med Sci. 2025; 80:glae297. https://doi.org/10.1093/gerona/glae297 [PubMed]
- 31. Wu L, Xie X, Liang T, Ma J, Yang L, Yang J, Li L, Xi Y, Li H, Zhang J, Chen X, Ding Y, Wu Q. Integrated Multi-Omics for Novel Aging Biomarkers and Antiaging Targets. Biomolecules. 2021; 12:39. https://doi.org/10.3390/biom12010039 [PubMed]
- 32. Tao X, Zhu Z, Wang L, Li C, Sun L, Wang W, Gong W. Biomarkers of Aging and Relevant Evaluation Techniques: A Comprehensive Review. Aging Dis. 2024; 15:977–1005. https://doi.org/10.14336/AD.2023.00808-1 [PubMed]
- 33. Ji Q, Jiang X, Wang M, Xin Z, Zhang W, Qu J, Liu GH. Multimodal Omics Approaches to Aging and Age-Related Diseases. Phenomics. 2024; 4:56–71. https://doi.org/10.1007/s43657-023-00125-x [PubMed]
- 34. Silva N, Rajado AT, Esteves F, Brito D, Apolónio J, Roberto VP, Binnie A, Araújo I, Nóbrega C, Bragança J, Castelo-Branco P, and ALFAScore Consortium. Measuring healthy ageing: current and future tools. Biogerontology. 2023; 24:845–66. https://doi.org/10.1007/s10522-023-10041-2 [PubMed]
- 35. Babu M, Snyder M. Multi-Omics Profiling for Health. Mol Cell Proteomics. 2023; 22:100561. https://doi.org/10.1016/j.mcpro.2023.100561 [PubMed]
- 36. Misra BB, Langefeld C, Olivier M, Cox LA. Integrated omics: tools, advances and future approaches. J Mol Endocrinol. 2019; 62:R21–45. https://doi.org/10.1530/JME-18-0055 [PubMed]
- 37. Luo Y, Zhao C, Chen F. Multiomics Research: Principles and Challenges in Integrated Analysis. Biodes Res. 2024; 6:0059. https://doi.org/10.34133/bdr.0059 [PubMed]
- 38. Kioroglou D, Gil-Redondo R, Embade N, Bizkarguenaga M, Conde R, Millet O, Mato JM, Marigorta UM. Multi-omic integration sets the path for early prevention strategies on healthy individuals. NPJ Genom Med. 2025; 10:35. https://doi.org/10.1038/s41525-025-00491-7 [PubMed]
- 39. Subramanian I, Verma S, Kumar S, Jere A, Anamika K. Multi-omics Data Integration, Interpretation, and Its Application. Bioinform Biol Insights. 2020; 14:1177932219899051. https://doi.org/10.1177/1177932219899051 [PubMed]
- 40. Li Y, Huang Q, Jiang J, Du X, Xiang W, Zhang S, Pan Z, Zhao L, Cui Y, Ke L, Yin B, Liu L, Feng G, et al. Large language model-based biological age prediction in large-scale populations. Nat Med. 2025; 31:2977–90. https://doi.org/10.1038/s41591-025-03856-8 [PubMed]
- 41. Ahmed Z, Wan S, Zhang F, Zhong W. Artificial intelligence for omics data analysis. BioMed Central. 2024; 1:4. https://doi.org/10.1186/s44330-024-00004-5
- 42. Wu Y, Xie L. AI-driven multi-omics integration for multi-scale predictive modeling of genotype-environment-phenotype relationships. Comput Struct Biotechnol J. 2025; 27:265–77. https://doi.org/10.1016/j.csbj.2024.12.030 [PubMed]
- 43. Nam Y, Kim J, Jung SH, Woerner J, Suh EH, Lee DG, Shivakumar M, Lee ME, Kim D. Harnessing Artificial Intelligence in Multimodal Omics Data Integration: Paving the Path for the Next Frontier in Precision Medicine. Annu Rev Biomed Data Sci. 2024; 7:225–50. https://doi.org/10.1146/annurev-biodatasci-102523-103801 [PubMed]
- 44. Yetgin A. Revolutionizing multi-omics analysis with artificial intelligence and data processing. Quant Biol. 2025; 13:e70002. https://doi.org/10.1002/qub2.70002 [PubMed]
- 45. Bürkle A, Moreno-Villanueva M, Bernhard J, Blasco M, Zondag G, Hoeijmakers JH, Toussaint O, Grubeck-Loebenstein B, Mocchegiani E, Collino S, Gonos ES, Sikora E, Gradinaru D, et al. MARK-AGE biomarkers of ageing. Mech Ageing Dev. 2015; 151:2–12. https://doi.org/10.1016/j.mad.2015.03.006 [PubMed]
- 46. Sayed N, Huang Y, Nguyen K, Krejciova-Rajaniemi Z, Grawe AP, Gao T, Tibshirani R, Hastie T, Alpert A, Cui L, Kuznetsova T, Rosenberg-Hasson Y, Ostan R, et al. An inflammatory aging clock (iAge) based on deep learning tracks multimorbidity, immunosenescence, frailty and cardiovascular aging. Nat Aging. 2021; 1:598–615. https://doi.org/10.1038/s43587-021-00082-y [PubMed]. Erratum in: Nat Aging. 2021;1:748. DOI: 10.1038/s43587-021-00102-x PMID:37117770.
- 47. Wang Y, Xiao S, Liu B, Jiang R, Liu Y, Hang Y, Chen L, Chen R, Vitiello MV, Bennett D, Wang B, Lv J, Yu C, et al. Organ-specific proteomic aging clocks predict disease and longevity across diverse populations. Nat Aging. 2026; 6:162–80. https://doi.org/10.1038/s43587-025-01016-8 [PubMed]
- 48. Garg M, Karpinski M, Matelska D, Middleton L, Burren OS, Hu F, Wheeler E, Smith KR, Fabre MA, Mitchell J, O'Neill A, Ashley EA, Harper AR, et al. Disease prediction with multi-omics and biomarkers empowers case-control genetic discoveries in the UK Biobank. Nat Genet. 2024; 56:1821–31. https://doi.org/10.1038/s41588-024-01898-1 [PubMed]
- 49. Gibbs SF, Pilbrow AP, Poppe KK, Earle NJ, Jones GT, Faatoese AF. What's counted counts: the implications of underrepresentation for the application of epigenetic clocks in diverse populations. Clin Epigenetics. 2025; 17:161. https://doi.org/10.1186/s13148-025-01954-5 [PubMed]
- 50. Dokuru DR, Horwitz TB, Freis SM, Stallings MC, Ehringer MA. South Asia: The Missing Diverse in Diversity. Behav Genet. 2024; 54:51–62. https://doi.org/10.1007/s10519-023-10161-y [PubMed]
- 51. India Ageing Report 2023: Key Insights And Recommendations For Elderly Care. IMPRI Impact And Policy Research Institute. 2026. https://www.impriindia.com/insights/policy-update/india-ageing-report-2023-key-insights-and-recommendations-for-elderly-care/.
- 52. Malati T. Whether western normative laboratory values used for clinical diagnosis are applicable to Indian population? An overview on reference interval. Indian J Clin Biochem. 2009; 24:111–22. https://doi.org/10.1007/s12291-009-0022-1 [PubMed]
- 53. EOI: ICMR’s Taskforce on Establishment of Reference Intervals in Indian Population (TERIIP Phase II): A Multicentric Observational Cross-Sectional Study among Child and Adolescent Population (<18 years). ICMR. 2026. https://www.icmr.gov.in/post/eoi-icmrs-taskforce-on-establishment-of-reference-intervals-in-indian-population-teriip-phase-ii-a-multicentric-observational-cross-sectional-study-among-child-and-adolescent-population-18-years-last-date-december-15-2025.
- 54. Das A, Dhillon P. Understanding healthy ageing in India: insights from multivariate regression trees. Aging Clin Exp Res. 2024; 36:158. https://doi.org/10.1007/s40520-024-02815-6 [PubMed]
- 55. Bhattacharyya C, Subramanian K, Uppili B, Biswas NK, Ramdas S, Tallapaka KB, Arvind P, Rupanagudi KV, Maitra A, Nagabandi T, De T, Singh K, Sharma P, et al., and GenomeIndia Consortium. Mapping genetic diversity with the GenomeIndia project. Nat Genet. 2025; 57:767–73. https://doi.org/10.1038/s41588-025-02153-x [PubMed]
- 56. Pengpid S, Peltzer K. Rural-urban health differences among aging adults in India. Heliyon. 2023; 10:e23397. https://doi.org/10.1016/j.heliyon.2023.e23397 [PubMed]
- 57. Prabhakaran D, Sieber NL, Jaganathan S, Mandal S, Prabhakaran P, Walia GK, Menon JS, Rajput P, Gupta T, Mohan S, Kondal D, Rajiva A, Dutta A, et al. Health effects of selected environmental Exposomes Across the Life courSe in Indian populations using longitudinal cohort studies: GEOHealth HEALS Study protocol. BMJ Open. 2024; 14:e087445. https://doi.org/10.1136/bmjopen-2024-087445 [PubMed]
- 58. Anjana RM, Sudha V, Abirami K, Gayathri R, Manasa VS, Deepa M, Pradeepa R, Unnikrishnan R, Joshi S, Saboo B, Gupta A, Joshi PP, Adhikari P, et al. Dietary profiles and associated metabolic risk factors in India from the ICMR-INDIAB survey-21. Nat Med. 2025; 31:3813–24. https://doi.org/10.1038/s41591-025-03949-4 [PubMed]
- 59. Longevity India. 2026. https://www.longevity.iisc.ac.in/.
- 60. Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, Downey P, Elliott P, Green J, Landray M, Liu B, Matthews P, Ong G, et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015; 12:e1001779. https://doi.org/10.1371/journal.pmed.1001779 [PubMed]
- 61. Denny JC, Rutter JL, Goldstein DB, Philippakis A, Smoller JW, Jenkins G, Dishman E, and All of Us Research Program Investigators. The "All of Us" Research Program. N Engl J Med. 2019; 381:668–76. https://doi.org/10.1056/NEJMsr1809937 [PubMed]
- 62. Dawber TR, Meadors GF, MOORE FE
Jr . Epidemiological approaches to heart disease: the Framingham Study. Am J Public Health Nations Health. 1951; 41:279–81. https://doi.org/10.2105/ajph.41.3.279 [PubMed] - 63. Lu JK, Wang W, Soh J, Sandalova E, Lim ZM, Seetharaman SK, Han JJ, Teo DB, Kennedy BK, Goh J, Maier AB. Characterizing biomarkers of ageing in Singaporeans: the ABIOS observational study protocol. Geroscience. 2025; 47:3997–4008. https://doi.org/10.1007/s11357-025-01511-1 [PubMed]
- 64. Ogishima S, Nagaie S, Mizuno S, Ishiwata R, Iida K, Shimokawa K, Takai-Igarashi T, Nakamura N, Nagase S, Nakamura T, Tsuchiya N, Nakaya N, Murakami K, et al., and Tohoku Medical Megabank Project Study Group. dbTMM: an integrated database of large-scale cohort, genome and clinical data for the Tohoku Medical Megabank Project. Hum Genome Var. 2021; 8:44. https://doi.org/10.1038/s41439-021-00175-5 [PubMed]
- 65. WHO Expert Consultation. Appropriate body-mass index for Asian populations and its implications for policy and intervention strategies. Lancet. 2004; 363:157–63. https://doi.org/10.1016/S0140-6736(03)15268-3 [PubMed]
- 66. van Niel J, Geelhoed-Duijvestijn PH, Numans ME, Kharagjitsing AV, Vos RC. Type 2 diabetes in South Asians compared to Europeans: Higher risk and earlier development of major cardiovascular events irrespective of the presence and degree of retinopathy. Results from The HinDu The Hague Diabetes Study. Endocrinol Diabetes Metab. 2021; 4:e00242. https://doi.org/10.1002/edm2.242 [PubMed]
- 67. Hammer T, Langholz E. The epidemiology of inflammatory bowel disease: balance between East and West? A narrative review. Dig Med Res. 2020; 3:48–8. https://doi.org/10.21037/dmr-20-149
- 68. Dikid T, Jain SK, Sharma A, Kumar A, Narain JP. Emerging and re-emerging infections in India: an overview. Indian J Med Res. 2013; 138:19–31. [PubMed]
- 69. Sengupta S, and Phenome India Consortium. Study research protocol for Phenome India-CSIR Health Cohort Knowledgebase: A prospective multi-modal follow-up study on a nationwide employee cohort. Biol Methods Protoc. 2025; 10:bpaf061. https://doi.org/10.1093/biomethods/bpaf061 [PubMed]
- 70. Perianayagam A, Bloom D, Lee J, Parasuraman S, Sekher TV, Mohanty SK, Chattopadhyay A, Govil D, Pedgaonkar S, Gupta S, Agarwal A, Posture A, Weerman A, Pramanik S. Cohort Profile: The Longitudinal Ageing Study in India (LASI). Int J Epidemiol. 2022; 51:e167–76. https://doi.org/10.1093/ije/dyab266 [PubMed]
- 71. Mhatre SS, Bragg F, Panse N, Judge PK, Manjrekar A, Burrett JA, Patil S, Davey Smith G, Kotkar L, Relton CL, Doibale P, Gadhave B, Chaturvedi P, et al. Cohort Profile: Indian Study of Healthy Ageing (ISHA-Barshi). Int J Epidemiol. 2024; 53:dyae079. https://doi.org/10.1093/ije/dyae079 [PubMed]
- 72. Albites-Sanabria J, Palumbo P, Bandinelli S, D'Ascanio I, Mellone S, Paraschiv-Ionescu A, Küderle A, Cereatti A, Del Din S, Kluge F, Gazit E, Jansen CP, Delgado-Ortiz L, et al. Walking into aging: real-world mobility patterns and digital benchmarks from the InCHIANTI Study. NPJ Aging. 2025; 11:60. https://doi.org/10.1038/s41514-025-00245-w [PubMed]
- 73. Kuo PL, Schrack JA, Shardell MD, Levine M, Moore AZ, An Y, Elango P, Karikkineth A, Tanaka T, de Cabo R, Zukley LM, AlGhatrif M, Chia CW, et al. A roadmap to build a phenotypic metric of ageing: insights from the Baltimore Longitudinal Study of Aging. J Intern Med. 2020; 287:373–94. https://doi.org/10.1111/joim.13024 [PubMed]
- 74. Blaha MJ, DeFilippis AP. Multi-Ethnic Study of Atherosclerosis (MESA): JACC Focus Seminar 5/8. J Am Coll Cardiol. 2021; 77:3195–216. https://doi.org/10.1016/j.jacc.2021.05.006 [PubMed]
- 75. Kliuchnikova AA, Ilgisonis EV, Archakov AI, Ponomarenko EA, Moskalev AA. Proteomic Markers of Aging and Longevity: A Systematic Review. Int J Mol Sci. 2024; 25:12634. https://doi.org/10.3390/ijms252312634 [PubMed]
- 76. Agarwal D, Paul S, Lele P, Piprode V, Kawade A, Hajela N, Bavdekar A, Parulekar V, Ginde M, Paranjape G, Matsuda K, Hori T, Juvekar S, Lal G. Changes in immunological parameters by ageing in rural healthy Indian adults and their associations with sex and lifestyle. Sci Rep. 2022; 12:15012. https://doi.org/10.1038/s41598-022-19227-z [PubMed]

