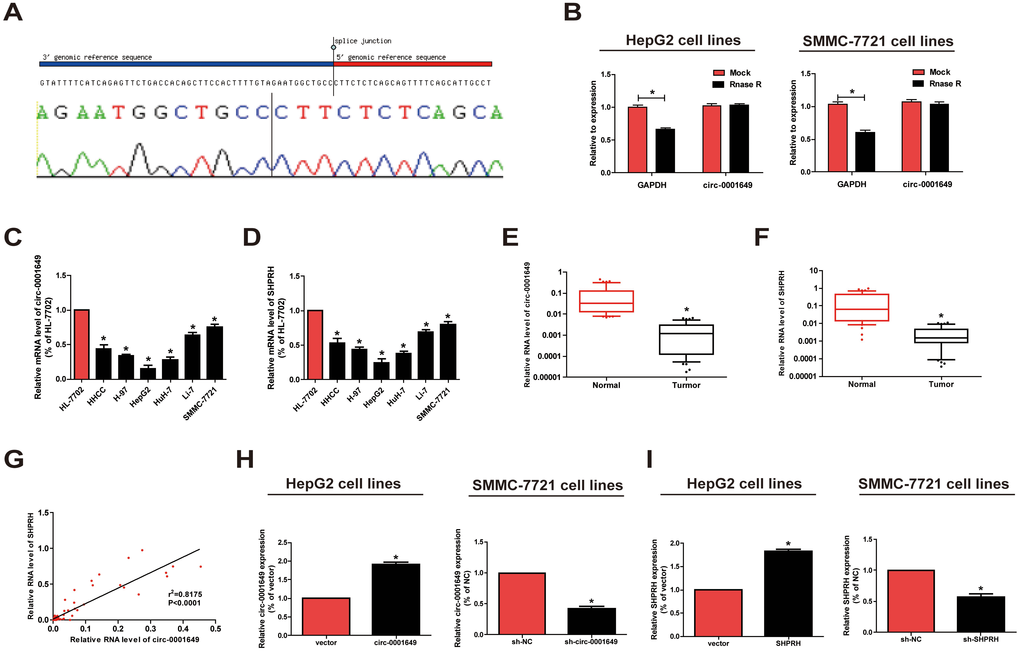
Figure 1.Characteristics and expression of circ-0001649 in HCC. (A) The sequence of circ-0001649 in circBase (upper panel) was consistent with the result of Sanger sequencing (lower panel). (B) Circ-0001649 was resistant to RNaseR digestion in HCC cell lines. (C) Circ-0001649 expression was verified by qRT-PCR in HHCC, H-97, HepG2, HuH-7, Li-7, SMMC-7721 cell lines and normal cell line HL-7702. Compared with HL-7702 cells, the expression of circ-0001649 in liver cancer cells was significantly reduced. Its expression was the lowest in HepG2 cells and the highest in SMMC-7721 cells. (D) SHPRH expression was verified by qRT-PCR in HCC cell lines and normal cell line HL-7702. (E) qRT-PCR detection of the relative expression of circ-0001649 in paired HCC tumor and normal tissues (n=84). (F) qRT-PCR detection of the relative expression of SHPRH in paired HCC tumor and normal tissues (n=84). (G) Correlation between circ-0001649 and SHPRH in HCC samples. (H) Expression of circ-0001649 was upregulated in circ-0001649 overexpression group and downregulated in the sh-circ-0001649 group. (I) Expression of SHPRH was upregulated in the SHPRH overexpression group and downregulated in the sh-SHPRH group. *P<0.05, data represent the mean ± SD.