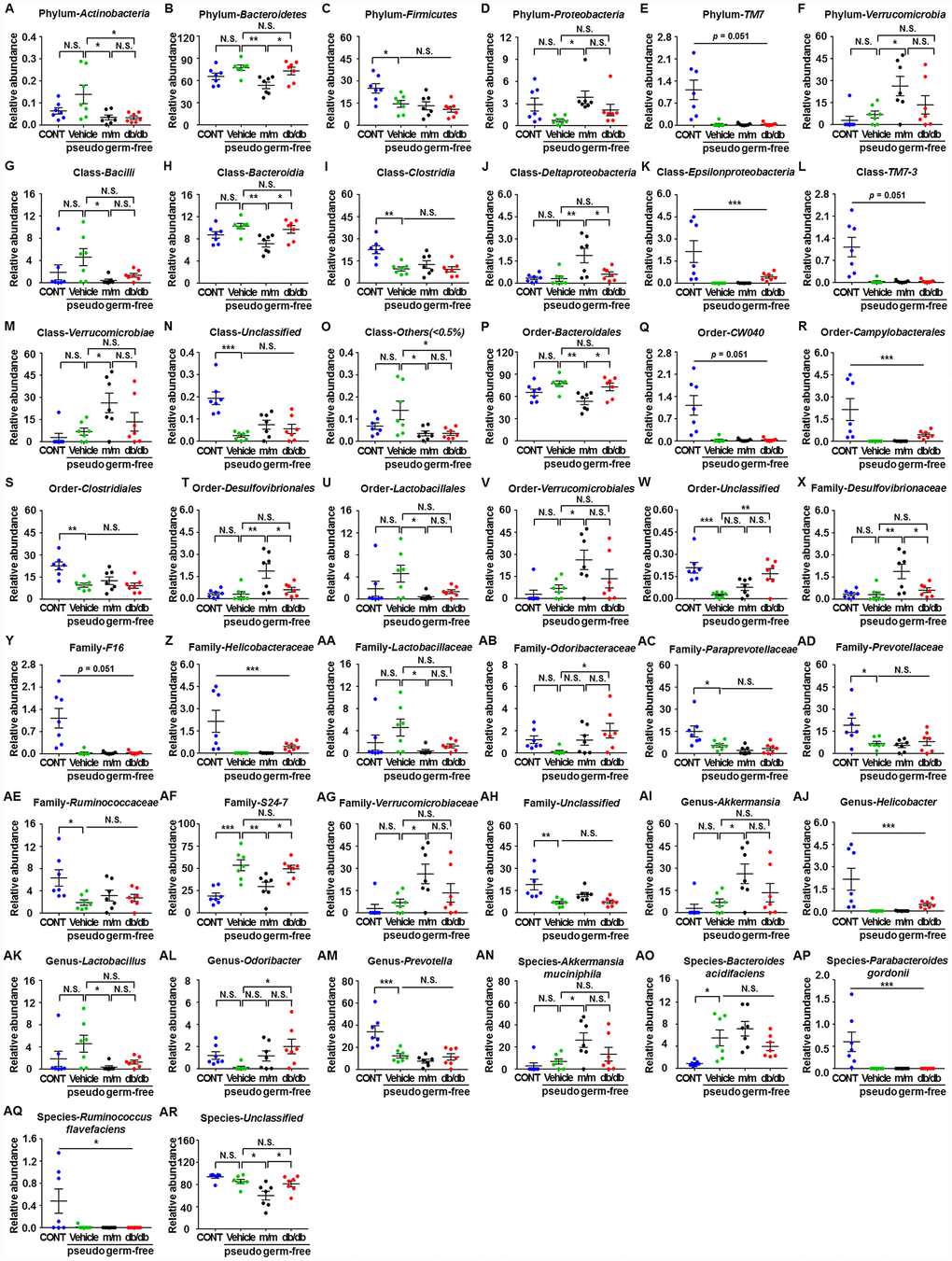
Figure 6.Differences in the relative abundances of gut microbiota composition among pseudo-germ-free mice after transplantation from db/db and m/m mice. (A) Phylum Actinobacteria (F3,24 = 4.607, P < 0.05); (B) Phylum Bacteroidetes (F3,24 = 5.458, P < 0.01); (C) Phylum Firmicutes (F3,24 = 5.936, P<0.01); (D) Phylum Proteobacteria (F3,24 = 3.204, P<0.05); (E) Phylum TM7 (Fisher’s exact test; P=0.051); (F) Phylum Verrucomicrobia (F3,24 = 4.348, P < 0.05); (G) Class Bacilli (F3,24 = 2.968, P = 0.0521); (H) Class Bacteroidia (F3,24 = 5.465, P<0.01); (I) Class Clostridia (F3,24 = 8.279, P < 0.001); (J) Class Deltaproteobacteria (F3,24 = 7.044, P < 0.01); (K) Class Epsilonproteobacteria (Fisher’s exact test; P<0.001); (L) Class TM7-3 (Fisher’s exact test; P = 0.051); (M) Class Verrucomicrobiae (F3,24 = 4.348, P < 0.05); (N) Class Unclassified (F3,24 = 12.87, P < 0.001); (O) Class Others (<0.5%) (F3,24 = 4.388, P < 0.05); (P) Order Bacteroidales (F3,24 = 5.465, P < 0.01); (Q) Order CW040 (F3,24 = 12.36, P < 0.001); (R) Order Campylobacterales (Fisher’s exact test; P < 0.001); (S) Order Clostridiales (F3,24 = 8.316, P < 0.001); (T) Order Desulfovibrionales (F3,24 = 7.044, P < 0.01); (U) Order Lactobacillales (F3,24 = 2.97, P=0.0520); (V) Order Verrucomicrobiales (F3,24 = 4.348, P < 0.05); (W) Order Unclassified (F3,24 = 9.449, P<0.001); (X) Family Desulfovibrionaceae (F3,24 = 7.044, P < 0.01); (Y) Family F16 (F3,24 = 12.36, P < 0.001); (Z) Family Helicobacteraceae (Fisher’s exact test; P < 0.001); (AA) Family Lactobacillaceae (F3,24 = 3.018, P<0.05); (AB) Family Odoribacteraceae (F3,24 = 3.087, P<0.05); (AC) Family Paraprevotellaceae (F3,24 = 7.505, P < 0.01); (AD) Family Prevotellaceae (F3,24 = 4.557, P < 0.05); (AE) Family Ruminococcaceae (F3,24 = 4.012, P < 0.05); (AF) Family S24-7 (F3,24 = 11.79, P < 0.001); (AG) Family Verrucomicrobiaceae (F3,24 = 4.348, P < 0.05); (AH) Family Unclassified (F3,24 = 7.609, P < 0.001); (AI) Genus Akkermansia (F3,24 = 4.348, P < 0.05); (AJ) Genus Helicobacter (Fisher’s exact test; P < 0.001); (AK) Genus Lactobacillus (F3,24 = 3.018, P<0.05); (AL) Genus Odoribacter (F3,24 = 3.087, P < 0.05); (AM) Genus Prevotella (F3,24 = 12.57, P < 0.001); (AN) Species Akkermansia muciniphila (F3,24 = 4.348, P < 0.05); (AO) Species Bacteroides acidifaciens (F3,24 = 6.558, P < 0.01); (AP) Species Parabacteroides gordonii (Fisher’s exact test; P < 0.001); (AQ) Species Ruminococcus flavefaciens (Fisher’s exact test; P < 0.05); (AR) Species Unclassified (F3,24 = 7.505, P < 0.01). Data are shown as mean ± SEM values (n=7). *P<0.05, **P<0.01, ***P<0.001. CONT, control; NS, not significant; SEM, standard error of the mean.