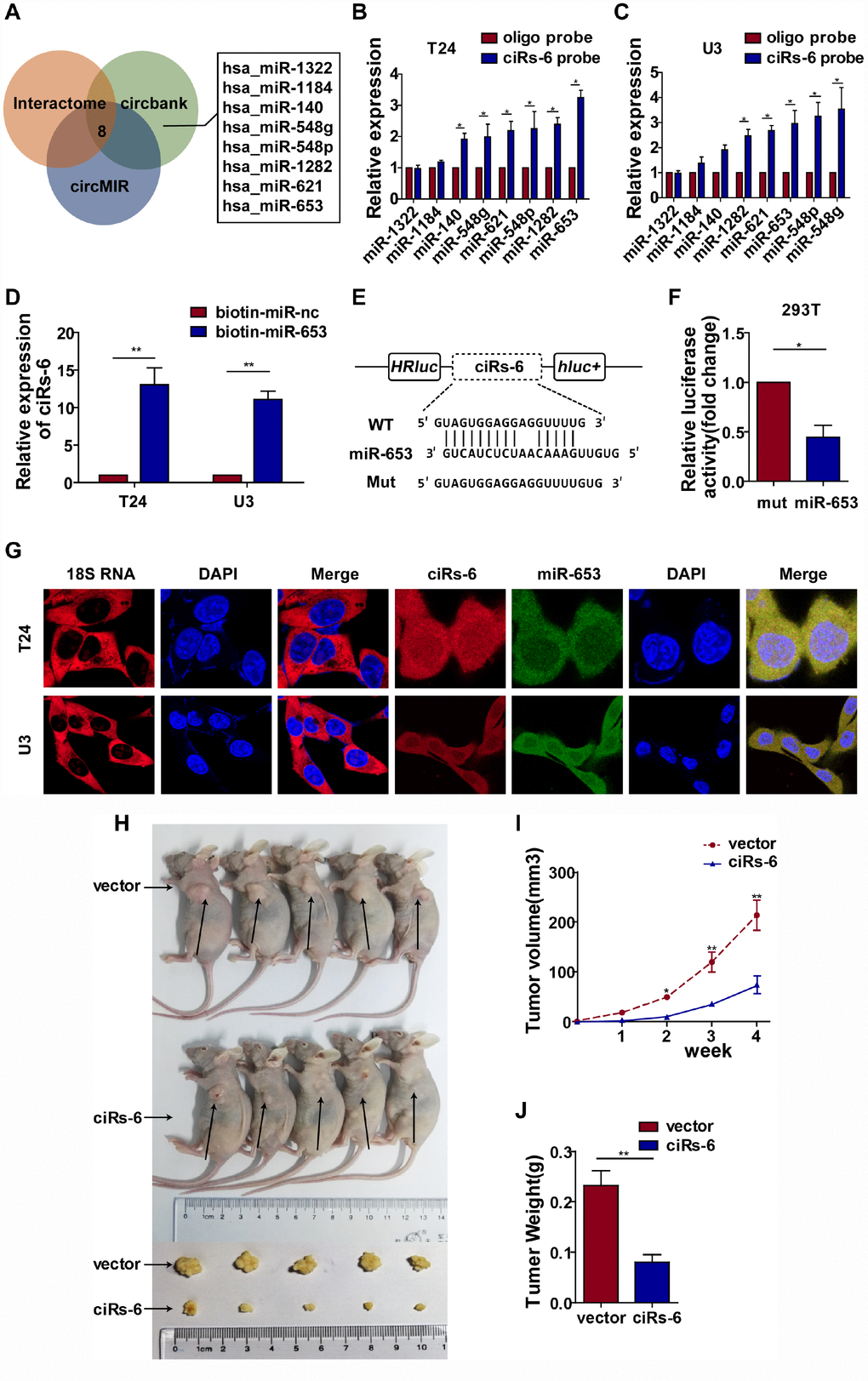
Figure 3.ciRs-6 effectively sponges miR-653. (A) miRNAs that could bind to ciRs-6 were predicted by Interactome, circBANK, and circMIR. Finally, 8 miRNAs were found commonly in each database; (B, C) RNA pulldown assays were performed to examine the affinity of ciRs-6 for the 8 above miRNAs in bladder cancer cells; (D) RNA pulldown assays were performed to evaluate the affinity of ciRs-6 for miR-653 in bladder cancer cells; (E, F) dual luciferase reporter assays showed binding properties of ciRs-6 and miR-653; (G) RNA FISH showed the cellular location of ciRs-6 and miR-653. Nuclei were stained with DAPI. Pictures were photographed at 400× under a light microscope. The results are displayed as the mean±SEM, *p<0.05, **p<0.01.