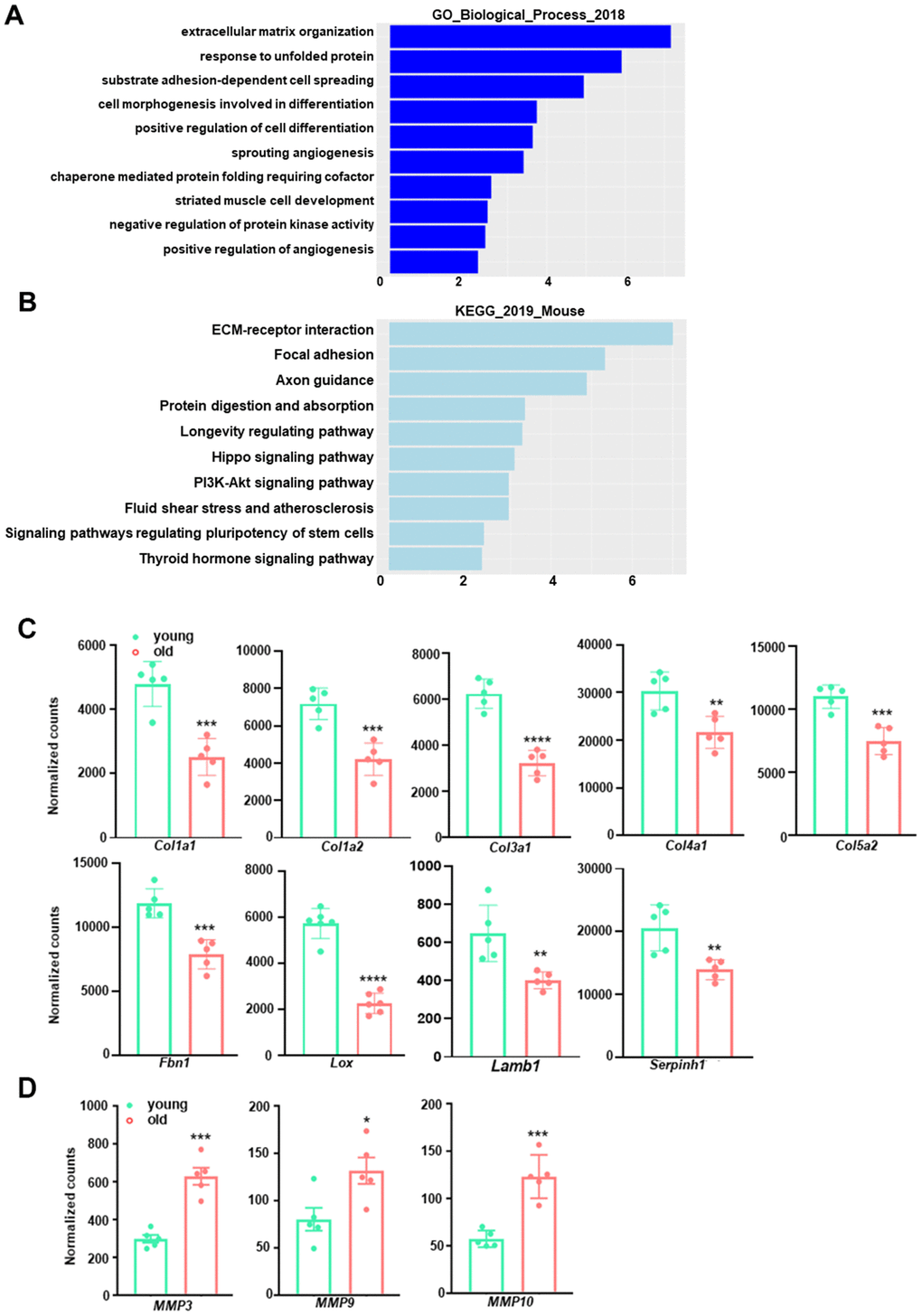
Figure 3.GO term enrichment and KEGG pathway analysis for downregulated transcripts in young and old mouse aortae. (A) The top 10 enriched GO biological process terms in the transcripts downregulated in old mouse aortae (adjusted p<0.05 and abs (log2FoldChange) > 0). Individual GO terms were sorted by adjusted p values. (B) The top 10 enriched KEGG pathways in the transcripts downregulated in old mouse aortae (adjusted p<0.05 and abs (log2FoldChange) > 0). Individual pathways were sorted by adjusted p values. (C, D) Normalized counts of significantly reduced genes in old aortae compared with young counterparts, including important extracellular matrix (ECM) components or regulators (C) and matrix metalloproteinases (MMPs) (D) Data are presented as a scatterplot of individual points with mean±SD, n=5. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 compared to young aortae, unpaired two-tailed Student's t-test.