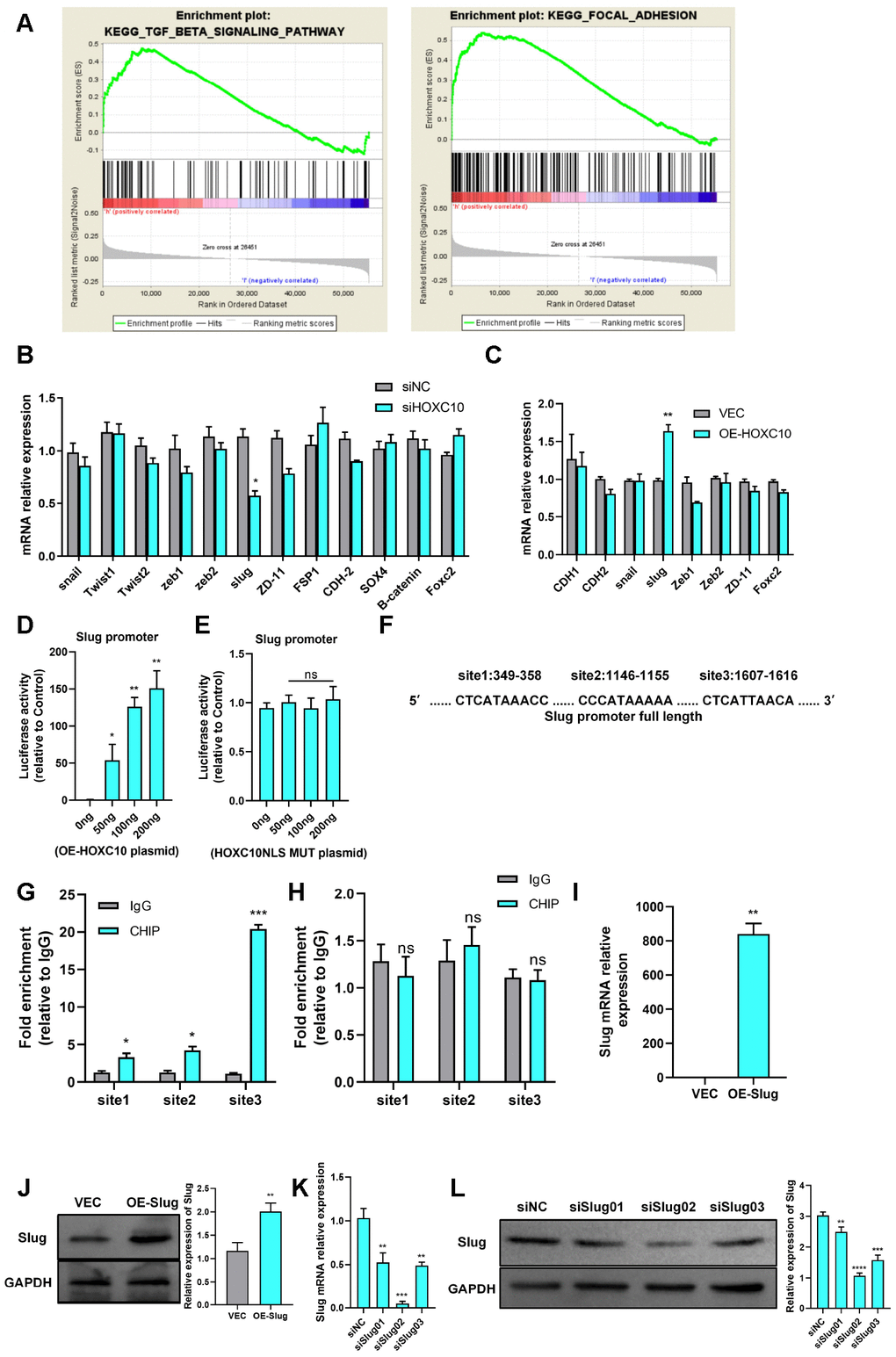
Figure 3.HOXC10 promotes OC cell migration by regulating Slug transcription. (A) The GSEA plot indicates that HOXC10 expression is positively correlated with TGF-β and FAK pathway signatures (OC sample data were downloaded from TCGA, n=379). (B, C) mRNA expression of EMT-related genes in 8910 cells transfected with HOXC10 siRNA, negative control siRNA, the HOXC10 overexpression plasmid, and empty vector. P=0.0257 and P=0.0093. (D) 8910 cells were cotransfected with a plasmid containing the full-length Slug promoter and increasing concentrations of the HOXC10 overexpression plasmid. P=0.0499, P=0.0032 and P=0.0079. (E) 8910 cells were cotransfected with the plasmid containing the full-length Slug promoter and increasing concentrations of the HOXC10 NLS mutation plasmid. P=0.1005, P=0.9559 and P=0.4059. (F) Schematic diagram of three predicted HOXC10 binding sites in the Slug promoter region. (G) Relative fold enrichment for IgG at the three predicted binding sites, as evaluated by ChIP-qPCR. P=0.0162, P=0.0158 and P=0.0003. (H) Relative fold enrichment for IgG at the three predicted binding sites after deletion of the HOXC10 DNA binding sites. P=0.5259, P=0.3579 and P=0.8293. (I, J) Relative mRNA and protein expression levels of Slug in 8910 cells transfected with the Slug overexpression plasmid and empty vector. P=0.0018, P=0.0044. (K–L) Transfection efficiencies of the Slug siRNA products. P=0.0048, P=0.0001, and P=0.0013; P=0.0083, P<0.0001, and P=0.0002.
(M, N) Rescue experiment using 8910 cells cotransfected with the HOXC10 overexpression plasmid, siRNA targeting Slug, empty vector or negative control. P<0.0001, P<0.0001, and P=0.0832; P<0.0001, P=0.0107, and P=0.5562. Scale bars, 200 μm and 100 μm, respectively. (O, P) Rescue experiment using 8910 cells cotransfected with siRNA targeting HOXC10, the Slug overexpression plasmid, negative control or empty vector. P<0.0001, P<0.0001, and P=0.1647; P<0.0001, P=0.0107, and P=0.5562. Scale bars, 200 μm and 100 μm, respectively.