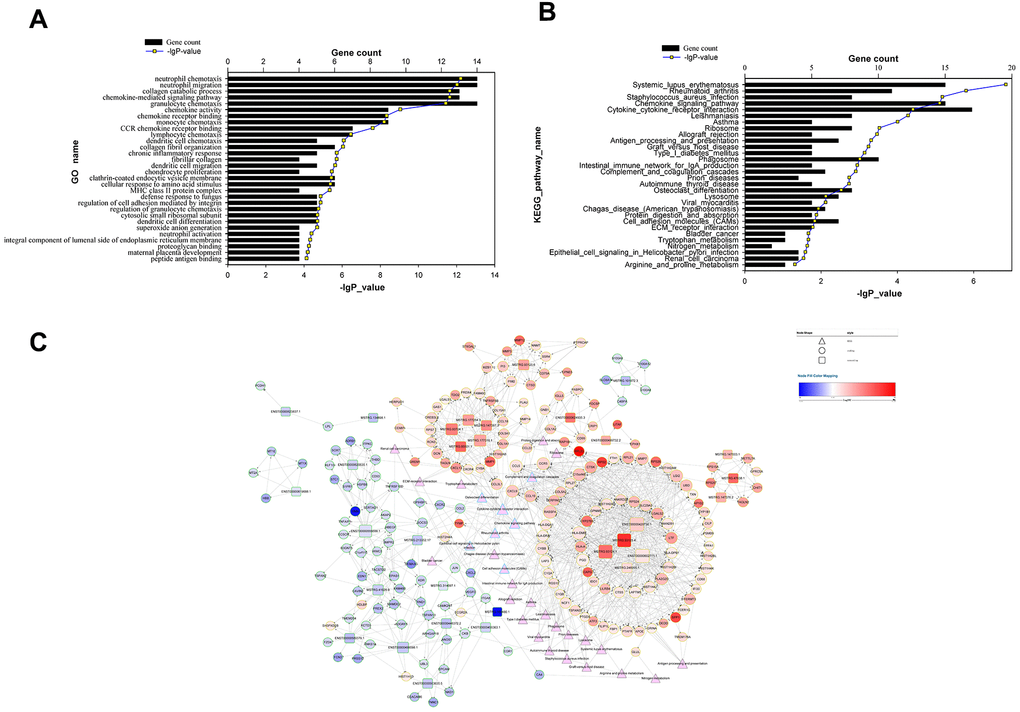
Figure 3.Functional analysis of target genes predicted for differentially expressed lncRNAs and the lncRNA-target-pathway network. (A) significant GO terms for differentially expressed target genes predicted for differentially expressed lncRNAs; (B) significant KEGG pathways enriched for differentially expressed target genes predicted for differentially expressed lncRNAs; (C) lncRNA-target-pathway network constructed by regulatory correlations of differentially expressed lncRNAs with predicted cis- or trans-target genes, as well as the top 30 KEGG pathways.